4BRV
 
 | | Superoxide reductase (Neelaredoxin) from Ignicoccus hospitalis E23A | | Descriptor: | DESULFOFERRODOXIN, FERROUS IRON-BINDING REGION, FE (III) ION | | Authors: | Romao, C.V, Matias, P.M, Pinho, F.G, Sousa, C.M, Barradas, A.R, Pinto, A.F, Teixeira, M, Bandeiras, T.M. | | Deposit date: | 2013-06-05 | | Release date: | 2014-06-18 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure of a Natural Sor Mutant
To be Published
|
|
4BOS
 
 | | Structure of OTUD2 OTU domain in complex with Ubiquitin K11-linked peptide | | Descriptor: | MAGNESIUM ION, NITRATE ION, OTUD2, ... | | Authors: | Mevissen, T.E.T, Hospenthal, M.K, Geurink, P.P, Elliott, P.R, Akutsu, M, Arnaudo, N, Ekkebus, R, Kulathu, Y, Wauer, T, El Oualid, F, Freund, S.M.V, Ovaa, H, Komander, D. | | Deposit date: | 2013-05-22 | | Release date: | 2013-07-24 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Otu Deubiquitinases Reveal Mechanisms of Linkage Specificity and Enable Ubiquitin Chain Restriction Analysis.
Cell(Cambridge,Mass.), 154, 2013
|
|
4BLV
 
 | | Crystal structure of Escherichia coli 23S rRNA (A2030-N6)- methyltransferase RlmJ in complex with S-adenosylmethionine (AdoMet) | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Punekar, A.S, Liljeruhm, J, Shepherd, T.R, Forster, A.C, Selmer, M. | | Deposit date: | 2013-05-04 | | Release date: | 2013-08-21 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Functional Insights Into the Molecular Mechanism of Rrna M6A Methyltransferase Rlmj.
Nucleic Acids Res., 41, 2013
|
|
4BM8
 
 | | Galectin-3c in complex with Bisamido-thiogalactoside derivate 3 | | Descriptor: | (3-Deoxy-3-(2,3,5,6-tetra-fluoro-4-methoxy-benzamido)-b-D-galactopyranosyl)-(3-deoxy-3-(2,3,5,6-tetra-fluoro-4-methoxy-benzamido)-2-O-sulfo-b-D-galactopyranosyl)-sulfide, GALECTIN-3 | | Authors: | Noresson, A.L, Oberg, C.T, Engstrom, O, Hakansson, M, Logan, D.T, Leffler, H, Nilsson, U.J. | | Deposit date: | 2013-05-07 | | Release date: | 2014-05-21 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (0.96 Å) | | Cite: | Controlling Protein Conformation Through Electronic Fine-Tuning of Arginine-Arene Interactions: Synthetic, Structural, and Biological Studies
To be Published
|
|
4BQE
 
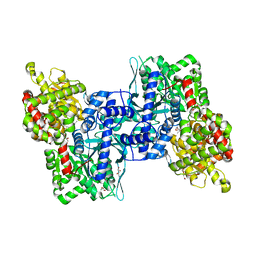 | | Arabidopsis thaliana Cytosolic Alpha-1,4-glucan Phosphorylase (PHS2) | | Descriptor: | ALPHA-GLUCAN PHOSPHORYLASE 2,4-GLUCAN PHOSPHORYLASE, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | O'Neill, E.C, Rashid, A.M, Stevenson, C.E.M, Hetru, A.C, Gunning, A.P, Rejzek, M, Nepogodiev, S.A, Bornemann, S, Lawson, D.M, Field, R.A. | | Deposit date: | 2013-05-30 | | Release date: | 2014-02-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Sugar-Coated Sensor Chip and Nanoparticle Surfaces for the in Vitro Enzymatic Synthesis of Starch-Like Materials
Chem.Sci., 5, 2014
|
|
4BT4
 
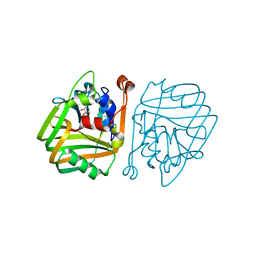 | | acetolactate decarboxylase with a bound (2S,3S)-2,3-Dihydroxy-2- methylbutanoic acid | | Descriptor: | (2S,3S)-2,3-dihydroxy-2-methylbutanoic acid, ALPHA-ACETOLACTATE DECARBOXYLASE, ZINC ION | | Authors: | A Marlow, V, Rea, D, Najmudin, S, Wills, M, Fulop, V. | | Deposit date: | 2013-06-12 | | Release date: | 2013-09-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure and Mechanism of Acetolactate Decarboxylase.
Acs Chem.Biol., 8, 2013
|
|
4BMQ
 
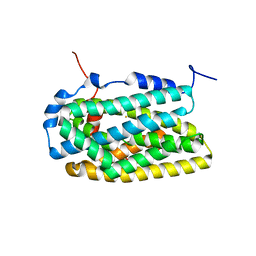 | | Crystal Structure of Ribonucleotide Reductase apo-NrdF from Bacillus cereus (space group C2) | | Descriptor: | FE (II) ION, RIBONUCLEOSIDE-DIPHOSPHATE REDUCTASE SUBUNIT BETA | | Authors: | Tomter, A.B, Hersleth, H.-P, Hammerstad, M, Rohr, A.K, Andersson, K.K. | | Deposit date: | 2013-05-10 | | Release date: | 2014-03-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of Bacillus Cereus Class Ib Ribonucleotide Reductase Di-Iron Nrdf in Complex with Nrdi.
Acs Chem.Biol., 9, 2014
|
|
2N4L
 
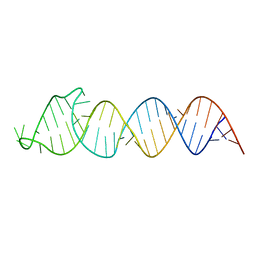 | | Solution Structure of the HIV-1 Intron Splicing Silencer and its Interactions with the UP1 Domain of hnRNP A1 | | Descriptor: | RNA (53-MER) | | Authors: | Tolbert, B.S, Jain, N, Morgan, C.E, Rife, B.D, Salemi, M. | | Deposit date: | 2015-06-23 | | Release date: | 2015-12-02 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the HIV-1 Intron Splicing Silencer and Its Interactions with the UP1 Domain of Heterogeneous Nuclear Ribonucleoprotein (hnRNP) A1.
J.Biol.Chem., 291, 2016
|
|
2MVL
 
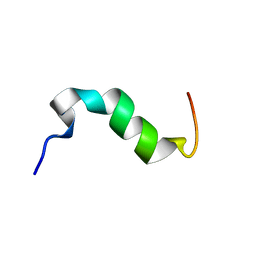 | | Solution structure of cytosolic part of Trop2 | | Descriptor: | Tumor-associated calcium signal transducer 2 | | Authors: | Ilc, G, Plavec, J, Vidmar, T, Pavsic, M, Lenarcic, B. | | Deposit date: | 2014-10-08 | | Release date: | 2015-05-06 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The cytosolic tail of the tumor marker protein Trop2--a structural switch triggered by phosphorylation.
Sci Rep, 5, 2015
|
|
4BRJ
 
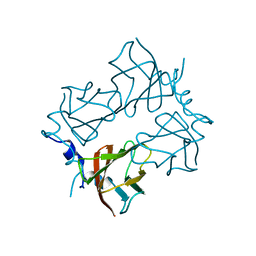 | | Superoxide reductase (Neelaredoxin) from Ignicoccus hospitalis T24K | | Descriptor: | DESULFOFERRODOXIN, FERROUS IRON-BINDING REGION, FE (III) ION | | Authors: | Romao, C.V, Matias, P.M, Pinho, F.G, Sousa, C.M, Barradas, A.R, Pinto, A.F, Teixeira, M, Bandeiras, T.M. | | Deposit date: | 2013-06-04 | | Release date: | 2014-06-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of a Natural Sor Mutant
To be Published
|
|
2MRA
 
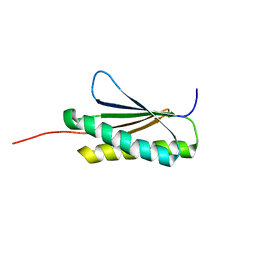 | | Solution NMR Structure of De novo designed protein, Northeast Structural Genomics Consortium (NESG) Target OR459 | | Descriptor: | De novo designed protein OR459 | | Authors: | Pulavarti, S.V.S.R.K, Kipnis, Y, Sukumaran, D, Maglaqui, M, Janjua, H, Mao, L, Xiao, R, Kornhaber, G, Baker, D, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2014-07-02 | | Release date: | 2014-09-17 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of De novo designed protein, Northeast Structural Genomics Consortium (NESG) Target OR459
To be Published
|
|
4BUV
 
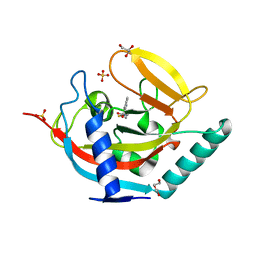 | | Crystal structure of human tankyrase 2 in complex with N-(4-(4-oxo-3, 4-dihydroquinazolin-2-yl)phenyl)thiophene-2-carboxamide | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, N-[4-(4-OXO-3,4-DIHYDROQUINAZOLIN-2-YL)PHENYL]THIOPHENE-2-CARBOXAMIDE, ... | | Authors: | Haikarainen, T, Narwal, M, Lehtio, L. | | Deposit date: | 2013-06-24 | | Release date: | 2013-10-30 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Para-Substituted 2-Phenyl-3,4-Dihydroquinazolin-4-Ones as Potent and Selective Tankyrase Inhibitors.
Chemmedchem, 8, 2013
|
|
4BTH
 
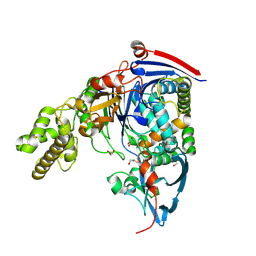 | | The LeuA146Trp,PheB24Tyr Double Mutant of the Quorum Quenching N-acyl Homoserine Lactone Acylase PvdQ Has an Altered Substrate Specificity Towards Small Acyl Chains | | Descriptor: | ACYL-HOMOSERINE LACTONE ACYLASE PVDQ SUBUNIT ALPHA, ACYL-HOMOSERINE LACTONE ACYLASE PVDQ SUBUNIT BETA, GLYCEROL | | Authors: | Koch, G, Nadal-Jimenez, P, Reis, C.R, Muntendam, R, Bokhove, M, Melillo, E, Dijkstra, B.W, Cool, R.H, Quax, W.J. | | Deposit date: | 2013-06-18 | | Release date: | 2014-01-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Reducing Virulence of the Human Pathogen Burkholderia by Altering the Substrate Specificity of the Quorum-Quenching Acylase Pvdq
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
2N4E
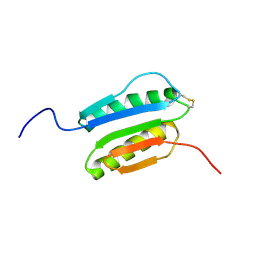 
 | | Solution NMR Structure of DE NOVO DESIGNED PROTEIN Top7NNSTYCC, Northeast Structural Genomics Consortium (NESG) Target OR34 | | Descriptor: | OR34 | | Authors: | Liu, G, Chan, K, Basanta, B, Xiao, R, Janjua, H, Kogan, S, Maglaqui, M, Ciccosanti, C, Acton, T.B, Kornhaber, G, Everett, J.K, Baker, D, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2015-06-17 | | Release date: | 2015-12-09 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of DE NOVO DESIGNED PROTEIN Top7NNSTYCC, Northeast Structural Genomics Consortium (NESG) Target OR34
To be Published
|
|
4BW9
 
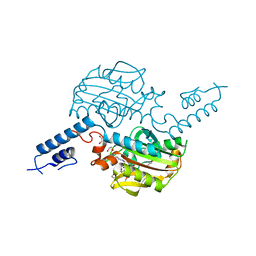 | | PylRS Y306G, Y384F, I405R mutant in complex with AMP-PNP | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, PENTAETHYLENE GLYCOL, ... | | Authors: | Schneider, S, Vrabel, M, Gattner, M.J, Fluegel, V, Lopez-Carillo, V, Carell, T. | | Deposit date: | 2013-07-01 | | Release date: | 2013-07-31 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural Insights Into Incorporation of Norbornene Amino Acids for Click Modification of Proteins
Chem.Bio.Chem., 14, 2013
|
|
2MQM
 
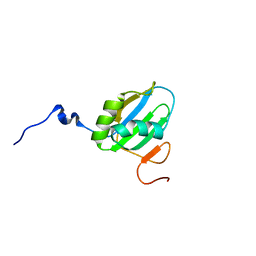 | | Structural Investigation of hnRNP L | | Descriptor: | Protein Hnrnpl | | Authors: | Blatter, M, Allain, F. | | Deposit date: | 2014-06-24 | | Release date: | 2015-05-06 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The Signature of the Five-Stranded vRRM Fold Defined by Functional, Structural and Computational Analysis of the hnRNP L Protein.
J.Mol.Biol., 427, 2015
|
|
4ER6
 
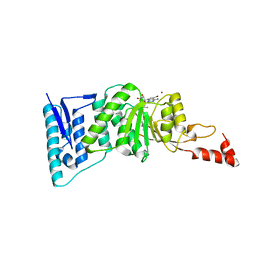 | | Crystal structure of human DOT1L in complex with inhibitor SGC0946 | | Descriptor: | 5-bromo-7-{5-[(3-{[(4-tert-butylphenyl)carbamoyl]amino}propyl)(propan-2-yl)amino]-5-deoxy-beta-D-ribofuranosyl}-7H-pyrrolo[2,3-d]pyrimidin-4-amine, BROMIDE ION, Histone-lysine N-methyltransferase, ... | | Authors: | Wernimont, A.K, Tempel, W, Yu, W, Scopton, A, Li, Y, Nguyen, K.T, Vedadi, M, Bradner, J.E, Schapira, M, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Brown, P.J, Structural Genomics Consortium (SGC) | | Deposit date: | 2012-04-19 | | Release date: | 2012-05-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Catalytic site remodelling of the DOT1L methyltransferase by selective inhibitors.
Nat Commun, 3, 2012
|
|
4BR3
 
 | | Determination of potential scaffolds for human choline kinase alpha 1 by chemical deconvolution studies | | Descriptor: | 1-((4'-((6-amino-3H-purin-3-yl)methyl)biphenyl-4-yl)methyl)-4-(dimethylamino)pyridinium, 3-benzyladenine, CHOLINE KINASE ALPHA, ... | | Authors: | Sahun-Roncero, M, Rubio-Ruiz, B, Conejo-Garcia, A, Velazquez-Campoy, A, Entrena, A, Hurtado-Guerrero, R. | | Deposit date: | 2013-06-03 | | Release date: | 2013-06-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Determination of Potential Scaffolds for Human Choline Kinase Alpha 1 by Chemical Deconvolution Studies
Chembiochem, 14, 2013
|
|
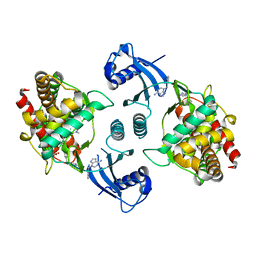
4BWS
 
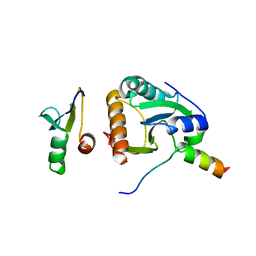 | | Crystal structure of the heterotrimer of PQBP1, U5-15kD and U5-52kD. | | Descriptor: | CD2 ANTIGEN CYTOPLASMIC TAIL-BINDING PROTEIN 2, POLYGLUTAMINE-BINDING PROTEIN 1, THIOREDOXIN-LIKE PROTEIN 4A | | Authors: | Mizuguchi, M, Obita, T, Serita, T, Kojima, R, Morimoto, T, Nabeshima, Y, Okazawa, H. | | Deposit date: | 2013-07-04 | | Release date: | 2014-04-30 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Mutations in the Pqbp1 Gene Prevent its Interaction with the Spliceosomal Protein U5-15Kd.
Nat.Commun., 5, 2014
|
|
4ZV5
 
 | | Crystal structure of N-myristoylated mouse mammary tumor virus matrix protein | | Descriptor: | MYRISTIC ACID, Matrix protein p10 | | Authors: | Zabransky, A, Dolezal, M, Dostal, J, Vanek, O, Hadravova, R, Stokrova, J, Brynda, J, Pichova, I. | | Deposit date: | 2015-05-18 | | Release date: | 2016-01-27 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | Myristoylation drives dimerization of matrix protein from mouse mammary tumor virus.
Retrovirology, 13, 2016
|
|
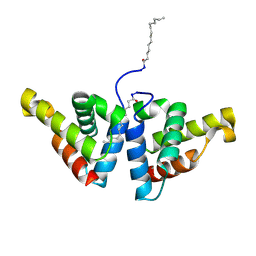
2PJ1
 
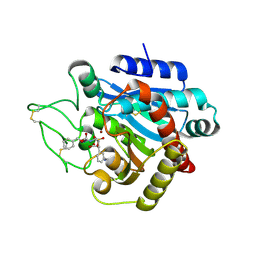 | | CRYSTAL STRUCTURE OF ACTIVATED PORCINE PANCREATIC CARBOXYPEPTIDASE B (3-Aminomethyl-phenyl)-[((R)-1-benzyloxycarbonylamino-2-methyl-propyl)-hydroxy-phosphinoyloxy]-acetic acid COMPLEX | | Descriptor: | (5R,6S,8S)-8-[3-(AMINOMETHYL)PHENYL]-6-HYDROXY-5-ISOPROPYL-3-OXO-1-PHENYL-2,7-DIOXA-4-AZA-6-PHOSPHANONAN-9-OIC ACID 6-OXIDE, Carboxypeptidase B, ZINC ION | | Authors: | Adler, M, Whitlow, M. | | Deposit date: | 2007-04-15 | | Release date: | 2008-01-22 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Structures of potent selective peptide mimetics bound to carboxypeptidase B.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
3H9N
 
 | | Crystal structure of the ribosome maturation factor rimm (hi0203) from h.influenzae. northeast structural genomics consortium target IR66. | | Descriptor: | Ribosome maturation factor rimM, SULFATE ION | | Authors: | Kuzin, A, Su, M, Seetharaman, J, Mao, M, Xiao, R, Maglaqui, M, Zhao, L, Everett, J.K, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-04-30 | | Release date: | 2009-05-19 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Northeast Structural Genomics Consortium Target IR66
To be Published
|
|
8CD9
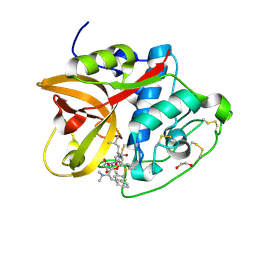 
 | | Cathepsin B1 from Schistosoma mansoni in complex with gallinamide analog 6 | | Descriptor: | 1,2-ETHANEDIOL, Cathepsin B-like peptidase (C01 family), SODIUM ION, ... | | Authors: | Rubesova, P, Brynda, J, Fanfrlik, J, Gerwick, W.H, Mares, M. | | Deposit date: | 2023-01-30 | | Release date: | 2024-02-21 | | Last modified: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Nature-Inspired Gallinamides Are Potent Antischistosomal Agents: Inhibition of the Cathepsin B1 Protease Target and Binding Mode Analysis.
Acs Infect Dis., 10, 2024
|
|
4C0A
 
 | | Arf1(Delta1-17)in complex with BRAG2 Sec7-PH domain | | Descriptor: | ADP-RIBOSYLATION FACTOR 1, GUANOSINE-3'-MONOPHOSPHATE-5'-DIPHOSPHATE, IQ MOTIF AND SEC7 DOMAIN-CONTAINING PROTEIN 1 | | Authors: | Aizel, K, Biou, V, Navaza, J, Duarte, L, Campanacci, V, Cherfils, J, Zeghouf, M. | | Deposit date: | 2013-08-01 | | Release date: | 2013-09-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Integrated conformational and lipid-sensing regulation of endosomal ArfGEF BRAG2.
PLoS Biol., 11, 2013
|
|
2MU9
 
 | | Changing ABRA protein peptide to fit the HLA-DR B1*0301 molecule renders it protection-inducing | | Descriptor: | P101/acidic basic repeat antigen | | Authors: | Salazar, L, Alba, M, Curtidor, H, Bermudez, A, Vargas, L, Rivera, Z, Patarroyo, M. | | Deposit date: | 2014-09-04 | | Release date: | 2015-09-16 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Changing ABRA protein peptide to fit into the HLA-DRbeta1*0301 molecule renders it protection-inducing.
Biochem.Biophys.Res.Commun., 322, 2004
|
|
