5ACR
 
 | | W228Y-Investigation of the impact from residues W228 and Y233 in the metallo-beta-lactamase GIM-1 | | Descriptor: | CALCIUM ION, GIM-1 PROTEIN, ZINC ION | | Authors: | Skagseth, S, Carlsen, T.J, Bjerga, G.E.K, Spencer, J, Samuelsen, O, Leiros, H.-K.S. | | Deposit date: | 2015-08-17 | | Release date: | 2015-12-23 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Role of Residues W228 and Y233 in the Structure and Activity of Metallo-Beta-Lactamase Gim-1.
Antimicrob.Agents Chemother., 60, 2015
|
|
2UXQ
 
 | | Isocitrate dehydrogenase from the psychrophilic bacterium Desulfotalea psychrophila: biochemical properties and crystal structure analysis | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, ISOCITRATE DEHYDROGENASE NATIVE, ... | | Authors: | Fedoy, A.-E, Yang, N, Martinez, A, Leiros, H.-K.S, Steen, I.H. | | Deposit date: | 2007-03-29 | | Release date: | 2008-02-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural and Functional Properties of Isocitrate Dehydrogenase from the Psychrophilic Bacterium Desulfotalea Psychrophila Reveal a Cold -Active Enzyme with an Unusual High Thermal
J.Mol.Biol., 372, 2007
|
|
2UXR
 
 | | Complex with isocitrate and the protein isocitrate dehydrogenase from the psychrophilic bacterium Desulfotalea psychrophila | | Descriptor: | GLYCEROL, ISOCITRATE DEHYDROGENASE, ISOCITRIC ACID, ... | | Authors: | Fedoy, A.-E, Yang, N, Martinez, A, Leiros, H.-K.S, Steen, I.H. | | Deposit date: | 2007-03-29 | | Release date: | 2008-02-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and Functional Properties of Isocitrate Dehydrogenase from the Psychrophilic Bacterium Desulfotalea Psychrophila Reveal a Cold -Active Enzyme with an Unusual High Thermal
J.Mol.Biol., 372, 2007
|
|
3SIU
 
 | | Structure of a hPrp31-15.5K-U4atac 5' stem loop complex, monomeric form | | Descriptor: | NHP2-like protein 1, SULFATE ION, U4/U6 small nuclear ribonucleoprotein Prp31, ... | | Authors: | Liu, S, Ghalei, H, Luhrmann, R, Wahl, M.C. | | Deposit date: | 2011-06-20 | | Release date: | 2011-08-10 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.626 Å) | | Cite: | Structural basis for the dual U4 and U4atac snRNA-binding specificity of spliceosomal protein hPrp31.
Rna, 17, 2011
|
|
2IUC
 
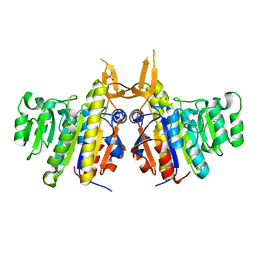 | | Structure of alkaline phosphatase from the Antarctic bacterium TAB5 | | Descriptor: | ALKALINE PHOSPHATASE, CACODYLATE ION, MAGNESIUM ION, ... | | Authors: | Wang, E, Koutsioulis, D, Leiros, H.K.S, Andersen, O.A, Bouriotis, V, Hough, E, Heikinheimo, P. | | Deposit date: | 2006-06-01 | | Release date: | 2006-11-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure of Alkaline Phosphatase from the Antarctic Bacterium Tab5.
J.Mol.Biol., 366, 2007
|
|
2Y1W
 
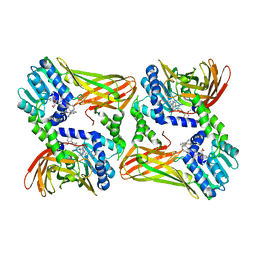 | | CRYSTAL STRUCTURE OF COACTIVATOR ASSOCIATED ARGININE METHYLTRANSFERASE 1 (CARM1) IN COMPLEX WITH SINEFUNGIN AND INDOLE INHIBITOR | | Descriptor: | 2-{4-[3-FLUORO-2-(2-METHOXYPHENYL)-1H-INDOL-5-YL] PIPERIDIN-1-YL}-N-METHYLETHANAMINE, HISTONE-ARGININE METHYLTRANSFERASE CARM1, SINEFUNGIN | | Authors: | Sack, J.S, Thieffine, S, Bandiera, T, Fasolini, M, Duke, G.J, Jayaraman, L, Kish, K.F, Klei, H.E, Purandare, A.V, Rosettani, P, Troiani, S, Xie, D, Bertrand, J.A. | | Deposit date: | 2010-12-10 | | Release date: | 2011-03-23 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Basis for Carm1 Inhibition by Indole and Pyrazole Inhibitors
Biochem.J., 436, 2011
|
|
2YNT
 
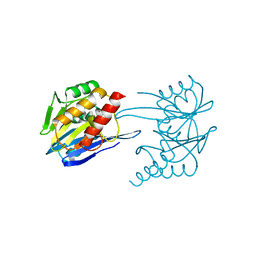 | | GIM-1-3Mol native. Crystal structures of Pseudomonas aeruginosa GIM- 1: active site plasticity in metallo-beta-lactamases | | Descriptor: | GIM-1 PROTEIN, GLYCEROL, ZINC ION | | Authors: | Borra, P.S, Samuelsen, O, Spencer, J, Lorentzen, M.S, Leiros, H.-K.S. | | Deposit date: | 2012-10-18 | | Release date: | 2013-07-24 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.598 Å) | | Cite: | Crystal Structures of Pseudomonas Aeruginosa Gim-1: Active-Site Plasticity in Metallo-Beta-Lactamases.
Antimicrob.Agents Chemother., 57, 2013
|
|
2YNW
 
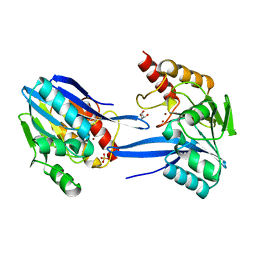 | | GIM-1-2Mol native. Crystal structures of Pseudomonas aeruginosa GIM- 1: active site plasticity in metallo-beta-lactamases | | Descriptor: | GIM-1 PROTEIN, GLYCEROL, SULFATE ION, ... | | Authors: | Borra, P.S, Samuelsen, O, Spencer, J, Lorentzen, M.S, Leiros, H.-K.S. | | Deposit date: | 2012-10-18 | | Release date: | 2013-07-24 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structures of Pseudomonas Aeruginosa Gim-1: Active-Site Plasticity in Metallo-Beta-Lactamases.
Antimicrob.Agents Chemother., 57, 2013
|
|
2Y1X
 
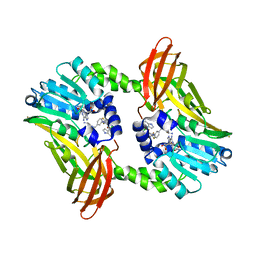 | | CRYSTAL STRUCTURE OF COACTIVATOR ASSOCIATED ARGININE METHYLTRANSFERASE 1 (CARM1) IN COMPLEX WITH SINEFUNGIN AND INDOLE INHIBITOR | | Descriptor: | CHLORIDE ION, HISTONE-ARGININE METHYLTRANSFERASE CARM1, N-(3-{5-[5-(1H-INDOL-4-YL)-1,3,4-OXADIAZOL-2-YL]-3-(TRIFLUOROMETHYL)-1H-PYRAZOL-1-YL}BENZYL)-L-ALANINAMIDE, ... | | Authors: | Sack, J.S, Thieffine, S, Bandiera, T, Fasolini, M, Duke, G.J, Jayaraman, L, Kish, K.F, Klei, H.E, Purandare, A.V, Rosettani, P, Troiani, S, Xie, D, Bertrand, J.A. | | Deposit date: | 2010-12-10 | | Release date: | 2011-03-23 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural Basis for Carm1 Inhibition by Indole and Pyrazole Inhibitors
Biochem.J., 436, 2011
|
|
1N6X
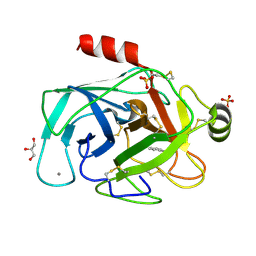 
 | | RIP-phasing on Bovine Trypsin | | Descriptor: | BENZYLAMINE, CALCIUM ION, GLYCEROL, ... | | Authors: | Ravelli, R.B.G, Leiros, H.-K.S, Pan, B, Caffrey, M, McSweeney, S. | | Deposit date: | 2002-11-12 | | Release date: | 2003-03-04 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Specific Radiation-Damage Can Be Used To Solve Macromolecular Crystal Structures
Structure, 11, 2003
|
|
1N6Y
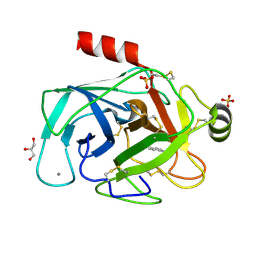 
 | | RIP-phasing on Bovine Trypsin | | Descriptor: | BENZYLAMINE, CALCIUM ION, GLYCEROL, ... | | Authors: | Ravelli, R.B.G, Leiros, H.-K.S, Pan, B, Caffrey, M, McSweeney, S. | | Deposit date: | 2002-11-12 | | Release date: | 2003-03-04 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Specific Radiation-Damage Can Be Used To Solve Macromolecular Crystal Structures
Structure, 11, 2003
|
|
2YNU
 
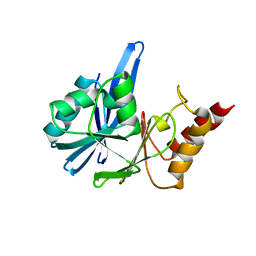 | | Apo GIM-1 with 2Mol. Crystal structures of Pseudomonas aeruginosa GIM-1: active site plasticity in metallo-beta-lactamases | | Descriptor: | GIM-1 PROTEIN | | Authors: | Borra, P.S, Samuelsen, O, Spencer, J, Lorentzen, M.S, Leiros, H.-K.S. | | Deposit date: | 2012-10-18 | | Release date: | 2013-07-24 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Crystal Structures of Pseudomonas Aeruginosa Gim-1: Active-Site Plasticity in Metallo-Beta-Lactamases.
Antimicrob.Agents Chemother., 57, 2013
|
|
2YNV
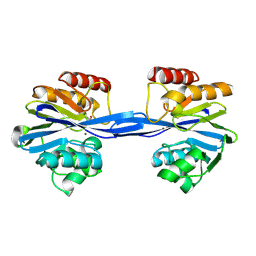 
 | | Cys221 oxidized, Mono zinc GIM-1 - GIM-1-Ox. Crystal structures of Pseudomonas aeruginosa GIM-1: active site plasticity in metallo-beta- lactamases | | Descriptor: | GIM-1 PROTEIN, MAGNESIUM ION, ZINC ION | | Authors: | Borra, P.S, Samuelsen, O, Spencer, J, Lorentzen, M.S, Leiros, H.-K.S. | | Deposit date: | 2012-10-18 | | Release date: | 2013-07-24 | | Last modified: | 2025-04-09 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Crystal Structures of Pseudomonas Aeruginosa Gim-1: Active-Site Plasticity in Metallo-Beta-Lactamases.
Antimicrob.Agents Chemother., 57, 2013
|
|
2VND
 
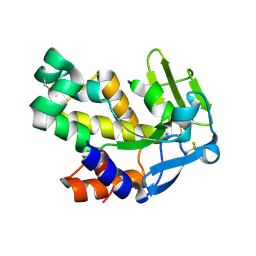 | | The N69Q mutant of Vibrio cholerae endonuclease I | | Descriptor: | CHLORIDE ION, ENDONUCLEASE I, MAGNESIUM ION | | Authors: | Niiranen, L, Altermark, B, Brandsdal, B.O, Leiros, H.-K.S, Helland, R, Smalas, A.O, Willassen, N.P. | | Deposit date: | 2008-02-04 | | Release date: | 2008-02-12 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Effects of Salt on the Kinetics and Thermodynamic Stability of Endonuclease I from Vibrio Salmonicida and Vibrio Cholerae.
FEBS J., 275, 2008
|
|
7ASS
 
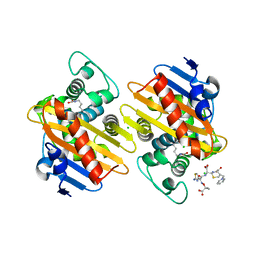 | |
5QAG
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 9a | | Descriptor: | 1,2-ETHANEDIOL, 3-(2-methoxycarbonylphenyl)benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAW
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 27 | | Descriptor: | 3-naphthalen-2-ylbenzoic acid, Beta-lactamase, CHLORIDE ION | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAI
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 12a | | Descriptor: | 1,2-ETHANEDIOL, 3-(3-methylsulfonylphenyl)benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAX
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 28 | | Descriptor: | 1,2-ETHANEDIOL, 3-quinolin-6-ylbenzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAC
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 6c | | Descriptor: | 1,2-ETHANEDIOL, 3-(4-methoxyphenyl)benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAM
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 17 | | Descriptor: | 1,2-ETHANEDIOL, 3-[4-(methylsulfonylamino)phenyl]benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QB3
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 40 | | Descriptor: | 1,2-ETHANEDIOL, 3-(3-acetamidophenyl)-5-quinolin-7-yl-benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAA
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 6a | | Descriptor: | 1,2-ETHANEDIOL, 3-(2-methoxyphenyl)benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QAU
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 26a | | Descriptor: | 1,2-ETHANEDIOL, 3-[3-(1~{H}-1,2,3,4-tetrazol-5-yl)phenyl]benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
5QA9
 
 | | OXA-48 IN COMPLEX WITH COMPOUND 5 | | Descriptor: | 1,2-ETHANEDIOL, 3-[3-(hydroxymethyl)phenyl]benzoic acid, Beta-lactamase, ... | | Authors: | Lund, B.A, Leiros, H.K.S. | | Deposit date: | 2017-07-11 | | Release date: | 2018-01-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A focused fragment library targeting the antibiotic resistance enzyme - Oxacillinase-48: Synthesis, structural evaluation and inhibitor design.
Eur J Med Chem, 145, 2018
|
|
