2ZU1
 
 | |
2ZTY
 
 | |
2ZTZ
 
 | |
5CXH
 
 | |
1ZLT
 
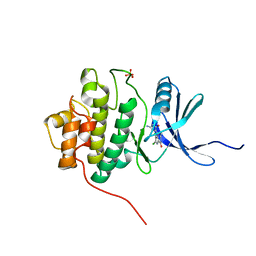 | | Crystal Structure of Chk1 Complexed with a Hymenaldisine Analog | | Descriptor: | (4Z)-4-(2-AMINO-5-OXO-3,5-DIHYDRO-4H-IMIDAZOL-4-YLIDENE)-2,3-DICHLORO-4,5,6,7-TETRAHYDROPYRROLO[2,3-C]AZEPIN-8(1H)-ONE, SULFATE ION, Serine/threonine-protein kinase Chk1 | | Authors: | Lee, C.C, Ng, K, Wan, Y, Gray, N, Spraggon, G. | | Deposit date: | 2005-05-09 | | Release date: | 2006-06-27 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Crystal Structure of Chk1 Complexed with a Hymenaldisine Analog
To be Published
|
|
1ZVW
 
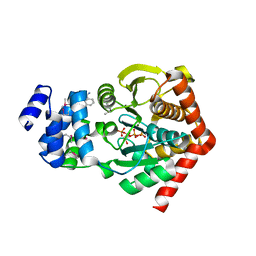 | | The Crystal Structure of TrpD (Rv2192c) from Mycobacterium tuberculosis in Complex with PRPP and Magnesium | | Descriptor: | 1-O-pyrophosphono-5-O-phosphono-alpha-D-ribofuranose, Anthranilate phosphoribosyltransferase, BENZAMIDINE, ... | | Authors: | Lee, C.E, Lott, J.S, Baker, E.N, Arcus, V.L, Javid-Majd, F, Goodfellow, C, Hung, L.-W, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2005-06-02 | | Release date: | 2006-01-17 | | Last modified: | 2021-08-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Crystal Structure of TrpD, a Metabolic Enzyme Essential for Lung Colonization by Mycobacterium tuberculosis, in Complex with its Substrate Phosphoribosylpyrophosphate.
J.Mol.Biol., 355, 2006
|
|
5Z1X
 
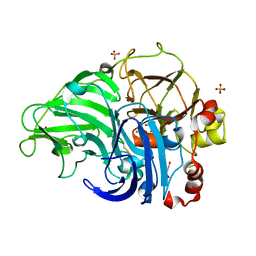 | | Crystal Structure of Laccase from Cerrena sp. RSD1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, COPPER (II) ION, ... | | Authors: | Lee, C.C, Wu, M.H, Ho, T.H, Wang, A.H.J. | | Deposit date: | 2017-12-28 | | Release date: | 2018-09-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Kinetic analysis and structural studies of a high-efficiency laccase fromCerrenasp. RSD1.
FEBS Open Bio, 8, 2018
|
|
5HXQ
 
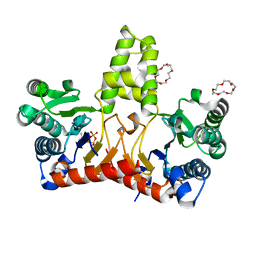 | | Crystal Structure of Z,Z-Farnesyl Diphosphate Synthase (D71M, E75A and H103Y Mutants) Complexed with DMSPP | | Descriptor: | (2Z,6Z)-farnesyl diphosphate synthase, chloroplastic, 1,4,7,10,13,16-HEXAOXACYCLOOCTADECANE, ... | | Authors: | Lee, C.C, Chan, Y.T, Wang, A.H.J. | | Deposit date: | 2016-01-31 | | Release date: | 2017-04-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure and Potential Head-to-Middle Condensation Function of a Z,Z-Farnesyl Diphosphate Synthase
Acs Omega, 2, 2017
|
|
2ZU2
 
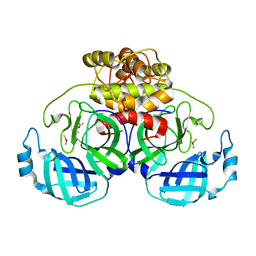 | | complex structure of CoV 229E 3CL protease with EPDTC | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 3C-like proteinase, zinc(II)hydrogensulfide | | Authors: | Lee, C.C, Wang, A.H.-J. | | Deposit date: | 2008-10-12 | | Release date: | 2009-01-13 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural Basis of Inhibition Specificities of 3C and 3C-like Proteases by Zinc-coordinating and Peptidomimetic Compounds
J.Biol.Chem., 284, 2009
|
|
5HXP
 
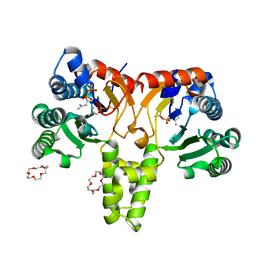 | | Crystal Structure of Z,Z-Farnesyl Diphosphate Synthase (D71M, E75A and H103Y Mutants) Complexed with IPP | | Descriptor: | (2Z,6Z)-farnesyl diphosphate synthase, chloroplastic, 1,4,7,10,13,16-HEXAOXACYCLOOCTADECANE, ... | | Authors: | Lee, C.C, Chan, Y.T, Wang, A.H.J. | | Deposit date: | 2016-01-31 | | Release date: | 2017-04-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal Structure and Potential Head-to-Middle Condensation Function of aZ,Z-Farnesyl Diphosphate Synthase.
Acs Omega, 2, 2017
|
|
7KHM
 
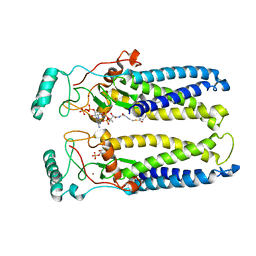 | | Crystal structure of hDHHS20 bound to palmitoyl CoA | | Descriptor: | Isoform 4 of Palmitoyltransferase ZDHHC20, PHOSPHATE ION, Palmitoyl-CoA, ... | | Authors: | Lee, C.-J, Banerjee, A. | | Deposit date: | 2020-10-21 | | Release date: | 2022-02-02 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.88 Å) | | Cite: | Bivalent recognition of fatty acyl-CoA by a human integral membrane palmitoyltransferase.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
3Q6N
 
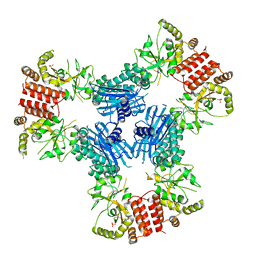 | | Crystal Structure of Human MC-HSP90 in P21 space group | | Descriptor: | Heat shock protein HSP 90-alpha, SULFATE ION | | Authors: | Lee, C.C, Lin, T.W, Ko, T.P, Wang, A.H.-J. | | Deposit date: | 2011-01-03 | | Release date: | 2011-06-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | The hexameric structures of human heat shock protein 90
Plos One, 6, 2011
|
|
3W9E
 
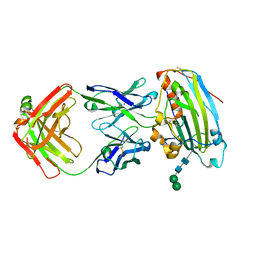 | | Structure of Human Monoclonal Antibody E317 Fab Complex with HSV-2 gD | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Antibody Fab heavy chain, Antibody Fab light chain, ... | | Authors: | Lee, C.C, Lin, L.L, Wang, A.H.J. | | Deposit date: | 2013-04-03 | | Release date: | 2013-10-02 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis for the antibody neutralization of herpes simplex virus
Acta Crystallogr.,Sect.D, 69, 2013
|
|
3Q6M
 
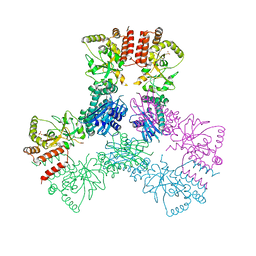 | | Crystal Structure of Human MC-HSP90 in C2221 Space Group | | Descriptor: | Heat shock protein HSP 90-alpha, SULFATE ION | | Authors: | Lee, C.C, Lin, T.W, Ko, T.P, Wang, A.H.-J. | | Deposit date: | 2011-01-03 | | Release date: | 2011-06-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The hexameric structures of human heat shock protein 90
Plos One, 6, 2011
|
|
5Z22
 
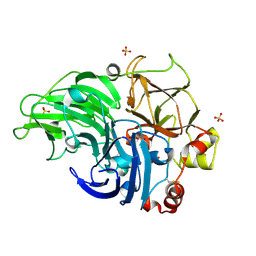 | | Crystal Structure of Laccase from Cerrena sp. RSD1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, COPPER (II) ION, ... | | Authors: | Lee, C.C, Wu, M.H, Ho, T.H, Wang, A.H.J. | | Deposit date: | 2017-12-28 | | Release date: | 2019-01-30 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Enhancement of laccase activity by pre-incubation with organic solvents.
Sci Rep, 9, 2019
|
|
6A4K
 
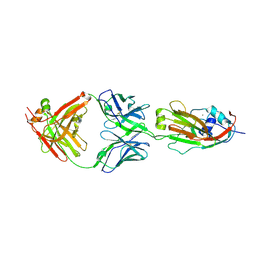 | | Human antibody 32D6 Fab in complex with H1N1 influenza A virus HA1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETATE ION, CALCIUM ION, ... | | Authors: | Lee, C.C, Ko, T.P, Lin, L.L, Wang, A.H.J. | | Deposit date: | 2018-06-20 | | Release date: | 2019-03-27 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | An Effective Neutralizing Antibody Against Influenza Virus H1N1 from Human B Cells.
Sci Rep, 9, 2019
|
|
3W9D
 
 | |
6JU0
 
 | | Mouse antibody 3.3 Fab in complex with PEG | | Descriptor: | 2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXYL, 3,6,9,12,15,18,21,24,27,30,33,36,39-TRIDECAOXAHENTETRACONTANE-1,41-DIOL, Fab heavy chain, ... | | Authors: | Lee, C.C, Ko, T.P, Lin, L.L, Wang, A.H.J. | | Deposit date: | 2019-04-12 | | Release date: | 2020-02-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.601 Å) | | Cite: | Structural basis of polyethylene glycol recognition by antibody.
J.Biomed.Sci., 27, 2020
|
|
5ZW8
 
 | | PigA with FAD and proline | | Descriptor: | 1,4,7,10,13,16-HEXAOXACYCLOOCTADECANE, CHLORIDE ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Lee, C.-C, Ko, T.-P, Wang, A.H.J. | | Deposit date: | 2018-05-14 | | Release date: | 2018-09-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.689 Å) | | Cite: | Crystal Structure of PigA: A Prolyl Thioester-Oxidizing Enzyme in Prodigiosin Biosynthesis.
Chembiochem, 20, 2019
|
|
5ZW2
 
 | | FAD complex of PigA | | Descriptor: | 1,2-ETHANEDIOL, 1,4,7,10,13,16-HEXAOXACYCLOOCTADECANE, ACETATE ION, ... | | Authors: | Lee, C.-C, Ko, T.-P, Wang, A.H.J. | | Deposit date: | 2018-05-14 | | Release date: | 2018-09-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.803 Å) | | Cite: | Crystal Structure of PigA: A Prolyl Thioester-Oxidizing Enzyme in Prodigiosin Biosynthesis.
Chembiochem, 20, 2019
|
|
2IJM
 
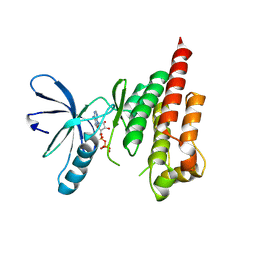 | |
5ZW0
 
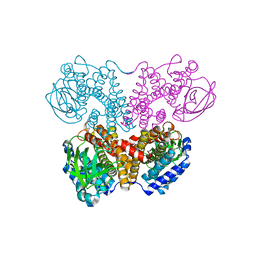 | | Apo-form PigA | | Descriptor: | L-prolyl-[peptidyl-carrier protein] dehydrogenase | | Authors: | Lee, C.-C, Ko, T.-P, Wang, A.H.J. | | Deposit date: | 2018-05-14 | | Release date: | 2018-09-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.54 Å) | | Cite: | Crystal Structure of PigA: A Prolyl Thioester-Oxidizing Enzyme in Prodigiosin Biosynthesis.
Chembiochem, 20, 2019
|
|
6AF6
 
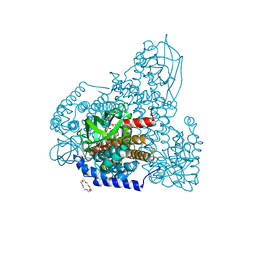 | | PigA with FAD and proline | | Descriptor: | 1,4,7,10,13,16-HEXAOXACYCLOOCTADECANE, CHLORIDE ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Lee, C.-C, Ko, T.-P, Wang, A.H.J. | | Deposit date: | 2018-08-08 | | Release date: | 2018-09-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Crystal Structure of PigA: A Prolyl Thioester-Oxidizing Enzyme in Prodigiosin Biosynthesis.
Chembiochem, 20, 2019
|
|
5ZW7
 
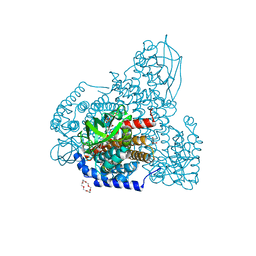 | | FAD-PigA complex at 1.3 A | | Descriptor: | 1,4,7,10,13,16-HEXAOXACYCLOOCTADECANE, CHLORIDE ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Lee, C.-C, Ko, T.-P, Wang, A.H.J. | | Deposit date: | 2018-05-14 | | Release date: | 2018-09-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal Structure of PigA: A Prolyl Thioester-Oxidizing Enzyme in Prodigiosin Biosynthesis.
Chembiochem, 20, 2019
|
|
6JWC
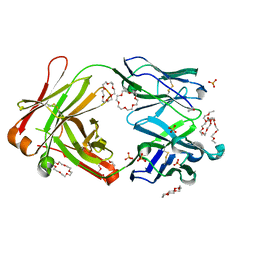 
 | | Mouse antibody 2B5 Fab in complex with PEG | | Descriptor: | 1,4,7,10,13,16-HEXAOXACYCLOOCTADECANE, DI(HYDROXYETHYL)ETHER, DODECAETHYLENE GLYCOL, ... | | Authors: | Lee, C.C, Ko, T.P, Su, Y.C, Lin, L.L, Roffler, S.R, Wang, A.H.J. | | Deposit date: | 2019-04-19 | | Release date: | 2020-02-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.297 Å) | | Cite: | Structural basis of polyethylene glycol recognition by antibody.
J.Biomed.Sci., 27, 2020
|
|
