7C11
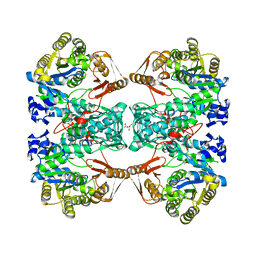 
 | | Formate--tetrahydrofolate ligase from Methylobacterium extorquens CM4 strain | | Descriptor: | ACETATE ION, CITRATE ANION, Formate-tetrahydrofolate ligase, ... | | Authors: | Kim, K.-J, Kim, S, Seo, H, Lee, S. | | Deposit date: | 2020-05-02 | | Release date: | 2020-10-28 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.815 Å) | | Cite: | Biochemical properties and crystal structure of formate-tetrahydrofolate ligase from Methylobacterium extorquens CM4.
Biochem.Biophys.Res.Commun., 528, 2020
|
|
2W8R
 
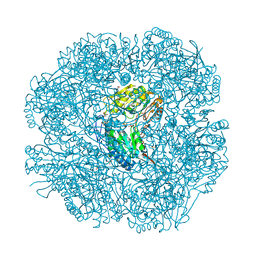 | |
2W8P
 
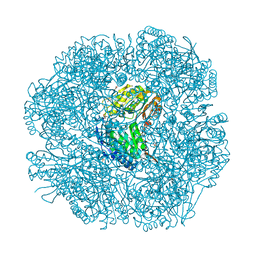 | | The crystal structure of human C340A SSADH | | Descriptor: | GLYCEROL, SUCCINIC SEMIALDEHYDE DEHYDROGENASE MITOCHONDRIAL, SULFATE ION | | Authors: | Kim, Y.-G, Kim, K.-J. | | Deposit date: | 2009-01-19 | | Release date: | 2009-06-09 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Redox-Switch Modulation of Human Ssadh by Dynamic Catalytic Loop.
Embo J., 28, 2009
|
|
8IDR
 
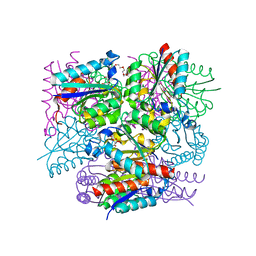 | |
8IDU
 
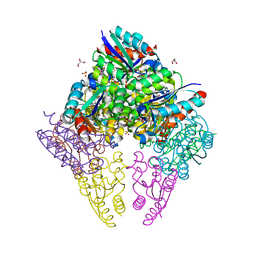 | | Crystal structure of substrate bound-form dehydroquinate dehydratase from Corynebacterium glutamicum | | Descriptor: | 1,3,4-TRIHYDROXY-5-OXO-CYCLOHEXANECARBOXYLIC ACID, 3-dehydroquinate dehydratase, GLYCEROL, ... | | Authors: | Lee, C.H, Kim, S, Kim, K.-J. | | Deposit date: | 2023-02-14 | | Release date: | 2023-12-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and Biochemical Analysis of 3-Dehydroquinate Dehydratase from Corynebacterium glutamicum .
J Microbiol Biotechnol., 33, 2023
|
|
8I4N
 
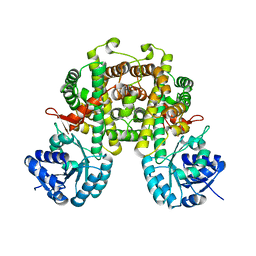 | |
8I4Q
 
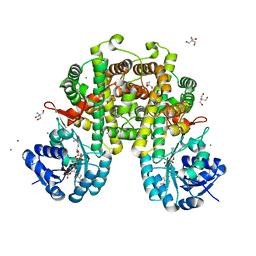 | |
5M47
 
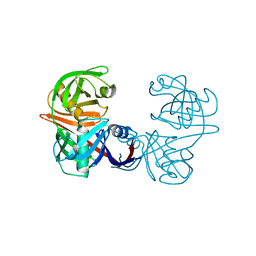 | |
8I4P
 
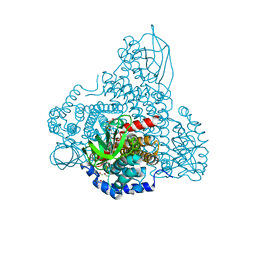 | |
4REP
 
 | | Crystal Structure of gamma-carotenoid desaturase | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, Gamma-carotene desaturase | | Authors: | Ahn, J.-W, Kim, E.-J, Kim, S, Kim, K.-J. | | Deposit date: | 2014-09-23 | | Release date: | 2015-07-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Crystal structure of 1'-OH-carotenoid 3,4-desaturase from Nonlabens dokdonensis DSW-6.
Enzyme.Microb.Technol., 77, 2015
|
|
6IOI
 
 | |
6ISS
 
 | | Lignin peroxidase H8 triple mutant S49C/A67C/H239 | | Descriptor: | CALCIUM ION, Ligninase H8, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Seo, H, Son, H, Kim, K.-J. | | Deposit date: | 2018-11-19 | | Release date: | 2019-11-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Extra disulfide and ionic salt bridge improves the thermostability of lignin peroxidase H8 under acidic condition
Enzyme.Microb.Technol., 148, 2021
|
|
6ITL
 
 | |
6ITK
 
 | |
8JCT
 
 | |
2WNW
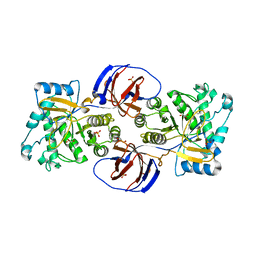 
 | | The crystal structure of SrfJ from salmonella typhimurium | | Descriptor: | ACTIVATED BY TRANSCRIPTION FACTOR SSRB, GLYCEROL, PHOSPHATE ION | | Authors: | Kim, Y.-G, Kim, J.-H, Kim, K.-J. | | Deposit date: | 2009-07-20 | | Release date: | 2010-03-02 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structure of the Salmonella Enterica Serovar Typhimurium Virulence Factor Srfj, a Glycoside Hydrolase Family Enzyme.
J.Bacteriol., 191, 2009
|
|
8JG3
 
 | |
8JG2
 
 | |
5B68
 
 | | Crystal structure of apo amylomaltase from Corynebacterium glutamicum | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 4-alpha-glucanotransferase, GLYCEROL, ... | | Authors: | Joo, S, Kim, S, Kim, K.-J. | | Deposit date: | 2016-05-25 | | Release date: | 2016-08-03 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structure of Amylomaltase from Corynebacterium glutamicum.
J.Agric.Food Chem., 64, 2016
|
|
2W8O
 
 | |
2W8Q
 
 | |
2W8N
 
 | |
4JDY
 
 | | Crystal structure of Rv2606c | | Descriptor: | GLYCEROL, Pyridoxal biosynthesis lyase PdxS | | Authors: | Kim, S, Kim, K.-J. | | Deposit date: | 2013-02-25 | | Release date: | 2013-05-22 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Mycobacterium tuberculosis Rv2606c: a pyridoxal biosynthesis lyase.
Biochem.Biophys.Res.Commun., 435, 2013
|
|
7ETX
 
 | |
7ETY
 
 | |
