3GVT
 
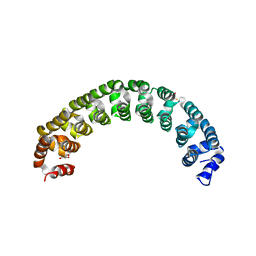 | |
3GVO
 
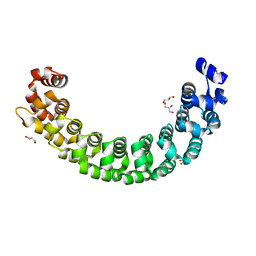 | |
2XS2
 
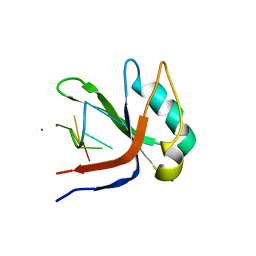 | |
2XS7
 
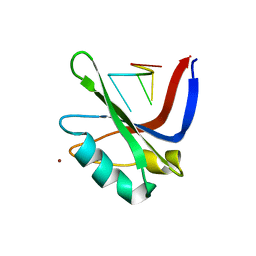 | |
2XSF
 
 | |
2XS5
 
 | |
5MFX
 
 | | Zika NS3 helicase:RNA complex | | Descriptor: | ACETATE ION, CITRATE ANION, Genome polyprotein, ... | | Authors: | Jenkins, H.T, Ge, M, Chechik, M, Greive, S.J, Antson, A.A. | | Deposit date: | 2016-11-18 | | Release date: | 2017-01-11 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.598 Å) | | Cite: | How Zika virus NS3 helicase grips the first
nucleotide of the leading RNA strand
To Be Published
|
|
2A55
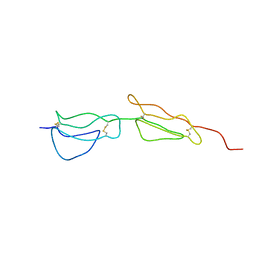 
 | | Solution structure of the two N-terminal CCP modules of C4b-binding protein (C4BP) alpha-chain. | | Descriptor: | C4b-binding protein | | Authors: | Jenkins, H.T, Mark, L, Ball, G, Lindahl, G, Uhrin, D, Blom, A.M, Barlow, P.N. | | Deposit date: | 2005-06-30 | | Release date: | 2005-12-13 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Human C4b-binding Protein, Structural Basis for Interaction with Streptococcal M Protein, a Major Bacterial Virulence Factor
J.Biol.Chem., 281, 2006
|
|
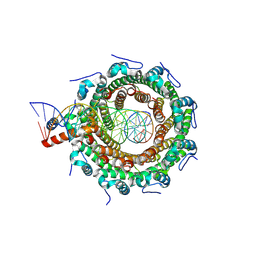
8POP
 | |
6EJQ
 
 | |
5M1O
 
 | | Crystal structure of the large terminase nuclease from thermophilic phage G20c with bound Cobalt | | Descriptor: | COBALT (II) ION, Phage terminase large subunit | | Authors: | Xu, R.G, Jenkins, H.T, Chechik, M, Blagova, E.V, Greive, S.J, Antson, A.A. | | Deposit date: | 2016-10-09 | | Release date: | 2016-10-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Viral genome packaging terminase cleaves DNA using the canonical RuvC-like two-metal catalysis mechanism.
Nucleic Acids Res., 45, 2017
|
|
5M1F
 
 | | Crystal structure of the large terminase nuclease from thermophilic phage G20c | | Descriptor: | Phage terminase large subunit | | Authors: | Xu, R.G, Jenkins, H.T, Chechik, M, Blagova, E.V, Greive, S.J, Antson, A.A. | | Deposit date: | 2016-10-07 | | Release date: | 2016-10-26 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Viral genome packaging terminase cleaves DNA using the canonical RuvC-like two-metal catalysis mechanism.
Nucleic Acids Res., 45, 2017
|
|
5M1Q
 
 | | Crystal structure of the large terminase nuclease from thermophilic phage G20c with bound Zinc | | Descriptor: | Phage terminase large subunit, ZINC ION | | Authors: | Xu, R.G, Jenkins, H.T, Chechik, M, Blagova, E.V, Greive, S.J, Antson, A.A. | | Deposit date: | 2016-10-09 | | Release date: | 2016-10-26 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Viral genome packaging terminase cleaves DNA using the canonical RuvC-like two-metal catalysis mechanism.
Nucleic Acids Res., 45, 2017
|
|
7AVJ
 | |
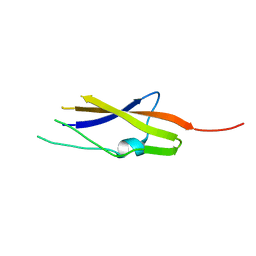
9FFH
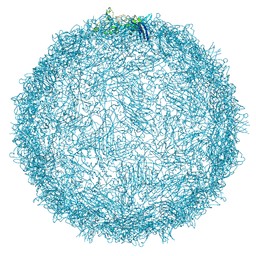 
 | | Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry | | Descriptor: | Major capsid protein | | Authors: | Bardy, P, MacDonald, C.I.W, Jenkins, H.T, Chechik, M, Hart, S.J, Turkenburg, J.P, Blaza, J.N, Fogg, P.C.M, Antson, A.A. | | Deposit date: | 2024-05-23 | | Release date: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | A stargate mechanism of Microviridae genome delivery unveiled by cryogenic electron tomography
To be published
|
|
9FFG
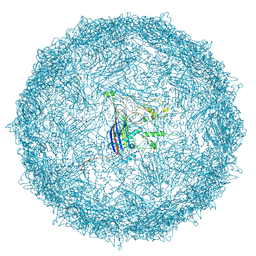 
 | | Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry | | Descriptor: | Major capsid protein | | Authors: | Bardy, P, MacDonald, C.I.W, Jenkins, H.T, Byrom, L, Chechik, M, Hart, S.J, Turkenburg, J.P, Blaza, J.N, Fogg, P.C.M, Antson, A.A. | | Deposit date: | 2024-05-23 | | Release date: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | A stargate mechanism of Microviridae genome delivery unveiled by cryogenic electron tomography
To be published
|
|
6RWZ
 
 | | Structure of Zika virus NS3 helicase in complex with ADP-BeF3 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, BERYLLIUM TRIFLUORIDE ION, MANGANESE (II) ION, ... | | Authors: | Ge, M, Molt Jr, R.W, Jenkins, H.T, Blackburn, G.M, Jin, Y, Antson, A.A. | | Deposit date: | 2019-06-07 | | Release date: | 2020-07-15 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Octahedral Trifluoromagnesate, an Anomalous Metal Fluoride Species, Stabilizes the Transition State in a Biological Motor.
Acs Catalysis, 11, 2021
|
|
6SE1
 
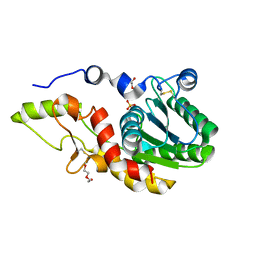 | | Structure of Salmonella ser. Paratyphi A lipopolysaccharide acetyltransferase periplasmic domain | | Descriptor: | DI(HYDROXYETHYL)ETHER, Putative lipopolysaccharide modification acyltransferase, SULFATE ION | | Authors: | Tindall, S.N, Pearson, C, Herman, R, Jenkins, H.T, Thomas, G.H, Potts, J.R, Van der Woude, M. | | Deposit date: | 2019-07-29 | | Release date: | 2020-08-26 | | Last modified: | 2020-09-16 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | Acetylation of Surface Carbohydrates in Bacterial Pathogens Requires Coordinated Action of a Two-Domain Membrane-Bound Acyltransferase.
Mbio, 11, 2020
|
|
6S0J
 
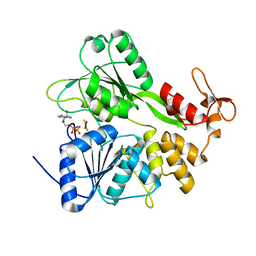 | | Structure of Zika virus NS3 helicase in complex with ADP-MgF3(H2O)- | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Genome polyprotein, MAGNESIUM ION, ... | | Authors: | Ge, M, Molt Jr, R.W, Jenkins, H.T, Blackburn, G.M, Jin, Y, Antson, A.A. | | Deposit date: | 2019-06-16 | | Release date: | 2020-07-15 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Octahedral Trifluoromagnesate, an Anomalous Metal Fluoride Species, Stabilizes the Transition State in a Biological Motor.
Acs Catalysis, 11, 2021
|
|
7AVH
 
 | | Streptococcal High Identity Repeats in Tandem (SHIRT) domains 3-4 from cell surface protein SGO_0707 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CHLORIDE ION, LPXTG cell wall surface protein | | Authors: | Whelan, F, Jenkins, H.T, Potts, J.R. | | Deposit date: | 2020-11-05 | | Release date: | 2021-06-09 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Periscope Proteins are variable-length regulators of bacterial cell surface interactions.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7AVK
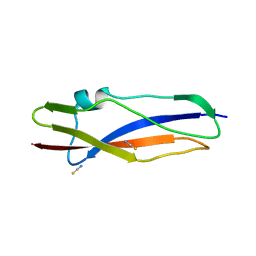 
 | | Streptococcal High Identity Repeats in Tandem (SHIRT) domain 10 from cell surface protein SGO_0707 | | Descriptor: | LPXTG cell wall surface protein, isothiocyanate | | Authors: | Degut, C, Gilburt, J, Whelan, F, Jenkins, H.T, Potts, J.R. | | Deposit date: | 2020-11-05 | | Release date: | 2021-06-09 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (0.82 Å) | | Cite: | Periscope Proteins are variable-length regulators of bacterial cell surface interactions.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7NT6
 
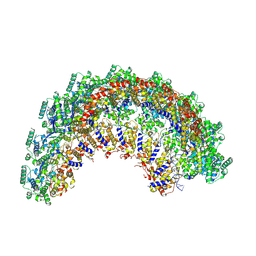 | | CryoEM structure of the Nipah virus nucleocapsid spiral clam-shaped assembly | | Descriptor: | Nucleoprotein, RNA (42-MER), RNA (48-MER) | | Authors: | Ker, D.S, Jenkins, H.T, Greive, S.J, Antson, A.A. | | Deposit date: | 2021-03-09 | | Release date: | 2021-07-07 | | Last modified: | 2021-07-28 | | Method: | ELECTRON MICROSCOPY (4.3 Å) | | Cite: | CryoEM structure of the Nipah virus nucleocapsid assembly.
Plos Pathog., 17, 2021
|
|
7NT5
 
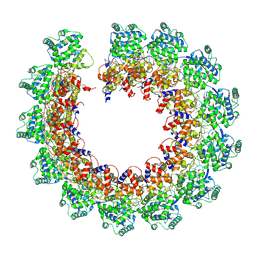 | | CryoEM structure of the Nipah virus nucleocapsid single helical turn assembly | | Descriptor: | Nucleoprotein, RNA (78-MER) | | Authors: | Ker, D.S, Jenkins, H.T, Greive, S.J, Antson, A.A. | | Deposit date: | 2021-03-09 | | Release date: | 2021-07-07 | | Last modified: | 2021-07-28 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | CryoEM structure of the Nipah virus nucleocapsid assembly.
Plos Pathog., 17, 2021
|
|
4XP7
 
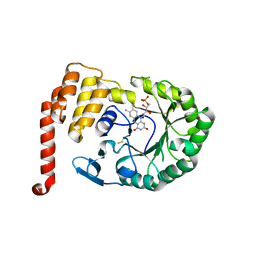 | | Crystal structure of Human tRNA dihydrouridine synthase 2 | | Descriptor: | 1-DEOXY-1-(7,8-DIMETHYL-2,4-DIOXO-3,4-DIHYDRO-2H-BENZO[G]PTERIDIN-1-ID-10(5H)-YL)-5-O-PHOSPHONATO-D-RIBITOL, tRNA-dihydrouridine(20) synthase [NAD(P)+]-like | | Authors: | Whelan, F, Jenkins, H.T, Griffiths, S, Byrne, R.T, Dodson, E.J, Antson, A.A. | | Deposit date: | 2015-01-16 | | Release date: | 2015-01-28 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | From bacterial to human dihydrouridine synthase: automated structure determination.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
4XVN
 
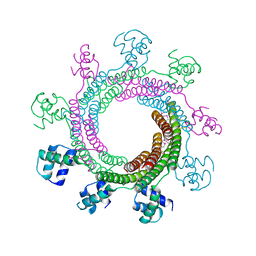 | |
