4UAM
 
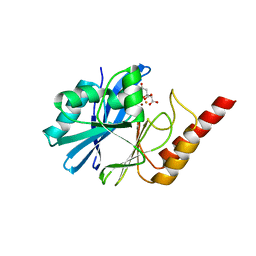 | | 1.8 Angstrom crystal structure of IMP-1 metallo-beta-lactamase with a mixed iron-zinc center in the active site | | Descriptor: | CITRATE ANION, FE (III) ION, IMP-1 metallo-beta-lactamase, ... | | Authors: | Carruthers, T.J, Carr, P.D, Jackson, C.J, Otting, G. | | Deposit date: | 2014-08-11 | | Release date: | 2014-09-17 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Iron(III) Located in the Dinuclear Metallo-beta-Lactamase IMP-1 by Pseudocontact Shifts.
Angew.Chem.Int.Ed.Engl., 53, 2014
|
|
7LL3
 
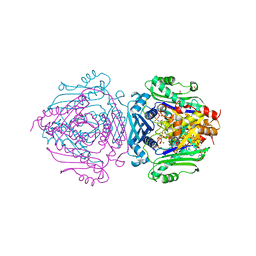 | | S-adenosylmethionine synthetase co-crystallized with UppNHp | | Descriptor: | (DIPHOSPHONO)AMINOPHOSPHONIC ACID, 1,2-ETHANEDIOL, 5'-O-[(R)-hydroxy{[(S)-hydroxy(phosphonoamino)phosphoryl]oxy}phosphoryl]uridine, ... | | Authors: | Tan, L.L, Jackson, C.J. | | Deposit date: | 2021-02-03 | | Release date: | 2021-03-31 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Substrate Dynamics Contribute to Enzymatic Specificity in Human and Bacterial Methionine Adenosyltransferases.
Jacs Au, 1, 2021
|
|
7LOW
 
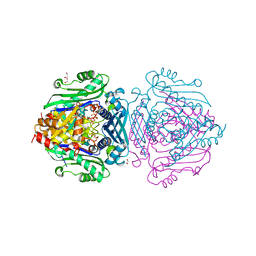 | | S-adenosylmethionine synthetase | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Tan, L.L, Jackson, C.J. | | Deposit date: | 2021-02-10 | | Release date: | 2021-03-31 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Substrate Dynamics Contribute to Enzymatic Specificity in Human and Bacterial Methionine Adenosyltransferases.
Jacs Au, 1, 2021
|
|
7LOZ
 
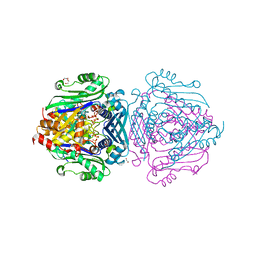 | | S-adenosylmethionine synthetase | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, PHOSPHATE ION, ... | | Authors: | Tan, L.L, Jackson, C.J. | | Deposit date: | 2021-02-11 | | Release date: | 2021-03-31 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Substrate Dynamics Contribute to Enzymatic Specificity in Human and Bacterial Methionine Adenosyltransferases.
Jacs Au, 1, 2021
|
|
7LNN
 
 | |
5BNC
 
 | | Structure of heme binding protein MSMEG_6519 from Mycobacterium smegmatis | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, MAGNESIUM ION, NICKEL (II) ION, ... | | Authors: | Ahmed, F.H, Carr, P.D, Jackson, C.J. | | Deposit date: | 2015-05-26 | | Release date: | 2015-10-21 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Sequence-Structure-Function Classification of a Catalytically Diverse Oxidoreductase Superfamily in Mycobacteria.
J.Mol.Biol., 427, 2015
|
|
5IVK
 
 | |
4WGX
 
 | | Crystal Structure of Molinate Hydrolase | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, COBALT (II) ION, Molinate hydrolase | | Authors: | Sugrue, E, Carr, P.D, Fraser, N.J, Hopkins, D.H, Jackson, C.J. | | Deposit date: | 2014-09-19 | | Release date: | 2015-02-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Evolutionary Expansion of the Amidohydrolase Superfamily in Bacteria in Response to the Synthetic Compounds Molinate and Diuron.
Appl.Environ.Microbiol., 81, 2015
|
|
6C2C
 
 | | The molecular basis for the functional evolution of an organophosphate hydrolysing enzyme | | Descriptor: | DI(HYDROXYETHYL)ETHER, MAGNESIUM ION, ZINC ION, ... | | Authors: | Hong, N.-S, Jackson, C.J, Carr, P.D, Tokuriki, N, Baier, F, Yang, G. | | Deposit date: | 2018-01-08 | | Release date: | 2019-01-16 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.597 Å) | | Cite: | Higher-order epistasis shapes the fitness landscape of a xenobiotic-degrading enzyme.
Nat.Chem.Biol., 15, 2019
|
|
5HMF
 
 | |
5HMD
 
 | |
4YBN
 
 | | Structure of the FAD and Heme binding protein msmeg_4975 from Mycobacterium smegmatis | | Descriptor: | ACETATE ION, FLAVIN-ADENINE DINUCLEOTIDE, Flavin-nucleotide-binding protein, ... | | Authors: | Ahmed, F.H, Carr, P.D, Jackson, C.J. | | Deposit date: | 2015-02-18 | | Release date: | 2015-10-21 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Sequence-Structure-Function Classification of a Catalytically Diverse Oxidoreductase Superfamily in Mycobacteria.
J.Mol.Biol., 427, 2015
|
|
5HME
 
 | |
7K7U
 
 | | BetaB2-crystallin | | Descriptor: | Beta-crystallin B2 | | Authors: | Tan, L.L, Jackson, C.J. | | Deposit date: | 2020-09-24 | | Release date: | 2022-02-09 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.03 Å) | | Cite: | BetaB2-crystallin
To Be Published
|
|
5W1U
 
 | | Culex quinquefasciatus carboxylesterase B2 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Carboxylic ester hydrolase, MALONATE ION | | Authors: | Hopkins, D.H, Jackson, C.J. | | Deposit date: | 2017-06-04 | | Release date: | 2017-08-16 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of an Insecticide Sequestering Carboxylesterase from the Disease Vector Culex quinquefasciatus: What Makes an Enzyme a Good Insecticide Sponge?
Biochemistry, 56, 2017
|
|
5W7H
 
 | | Supercharged arPTE variant R5 | | Descriptor: | Phosphotriesterase, ZINC ION | | Authors: | Campbell, E, Grant, J, Wang, Y, Sandhu, M, Williams, R.J, Nisbet, D.R, Perriman, A, Lupton, D, Jackson, C.J. | | Deposit date: | 2017-06-19 | | Release date: | 2019-01-23 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Hydrogel-Immobilized Supercharged Proteins
Adv Biosyst, 2018
|
|
5C8V
 
 | |
5CH3
 
 | | E3 alpha-esterase-7 carboxylesterase | | Descriptor: | Carboxylic ester hydrolase | | Authors: | Correy, G, Mabbitt, P, Jackson, C.J. | | Deposit date: | 2015-07-10 | | Release date: | 2016-06-15 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Mapping the Accessible Conformational Landscape of an Insect Carboxylesterase Using Conformational Ensemble Analysis and Kinetic Crystallography.
Structure, 24, 2016
|
|
7KW3
 
 | | Non Ribosomal PCP domain | | Descriptor: | PCP domain, SULFATE ION | | Authors: | Izore, T, Ho, Y.T.C, Kaczmarski, J.A, Gavriilidou, A, Chow, K.H, Steer, D, Goode, R.J.A, Schittenhelm, R.B, Tailhades, J, Tosin, M, Challis, G.L, Krenske, E.H, Ziemert, N, Jackson, C.J, Cryle, M.J. | | Deposit date: | 2020-11-29 | | Release date: | 2021-03-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of a non-ribosomal peptide synthetase condensation domain suggest the basis of substrate selectivity.
Nat Commun, 12, 2021
|
|
5THM
 
 | | Esterase-6 from Drosophila melanogaster | | Descriptor: | Esterase-6, N,N-dimethylboranamine | | Authors: | Fraser, N.J, Jackson, C.J. | | Deposit date: | 2016-09-29 | | Release date: | 2017-03-08 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Molecular basis for the behavioral effects of the odorant degrading enzyme Esterase 6 in Drosophila.
Sci Rep, 7, 2017
|
|
5EH9
 
 | | Indirect contributions of mutations underlie optimization of new enzyme function | | Descriptor: | 2-HYDROXYETHYL DISULFIDE, GLYCEROL, N-acyl homoserine lactonase AiiA, ... | | Authors: | Hong, N.-S, Jackson, C.J, Tokuriki, N, Yang, G, Baier, F. | | Deposit date: | 2015-10-28 | | Release date: | 2016-09-07 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.29 Å) | | Cite: | Conformational Tinkering Drives Evolution of a Promiscuous Activity through Indirect Mutational Effects.
Biochemistry, 55, 2016
|
|
5DQ6
 
 | | Mus musculus A20 OTU domain | | Descriptor: | Tumor necrosis factor alpha-induced protein 3 | | Authors: | Mabbitt, P.D, Jackson, C.J. | | Deposit date: | 2015-09-14 | | Release date: | 2015-12-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Mus musculus A20 OTU domain
To Be Published
|
|
5TYP
 
 | |
5TYK
 
 | |
5TUJ
 
 | |
