3S9K
 
 | | Crystal structure of the Itk SH2 domain. | | Descriptor: | CITRIC ACID, Tyrosine-protein kinase ITK/TSK | | Authors: | Joseph, R.E, Ginder, N.D, Hoy, J.A, Nix, J.C, Fulton, B.D, Honzatko, R.B, Andreotti, A.H. | | Deposit date: | 2011-06-01 | | Release date: | 2012-02-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.354 Å) | | Cite: | Structure of the interleukin-2 tyrosine kinase Src homology 2 domain; comparison between X-ray and NMR-derived structures.
Acta Crystallogr.,Sect.F, 68, 2012
|
|
6TW5
 
 | | Plasmodium vivax N-myristoyltransferase with bound indazole inhibitor IMP-917 | | Descriptor: | 1-[5-[4-fluoranyl-2-[2-(1,3,5-trimethylpyrazol-4-yl)ethoxy]phenyl]-2~{H}-indazol-3-yl]-~{N},~{N}-dimethyl-methanamine, CHLORIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Brannigan, J.A. | | Deposit date: | 2020-01-12 | | Release date: | 2021-07-21 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Plasmodium vivax N-myristoyltransferase with bound indazole inhibitor IMP-917
To Be Published
|
|
6TW6
 
 | |
6TW7
 
 | |
6TW8
 
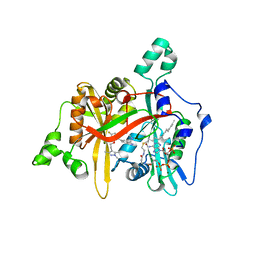 | | Leishmania major N-myristoyltransferase in complex with indazole inhibitor IMP-917 | | Descriptor: | 1-[5-[4-fluoranyl-2-[2-(1,3,5-trimethylpyrazol-4-yl)ethoxy]phenyl]-2~{H}-indazol-3-yl]-~{N},~{N}-dimethyl-methanamine, Glycylpeptide N-tetradecanoyltransferase, MAGNESIUM ION, ... | | Authors: | Brannigan, J.A. | | Deposit date: | 2020-01-12 | | Release date: | 2021-07-21 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Leishmania major N-myristoyltransferase in complex with indazole inhibitor IMP-917
To Be Published
|
|
1JHK
 
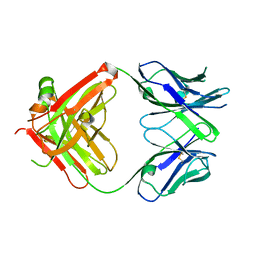 | |
6UY5
 
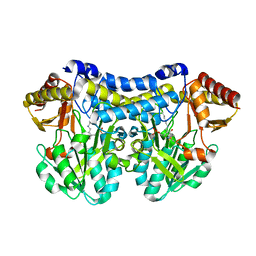 | |
3RW9
 
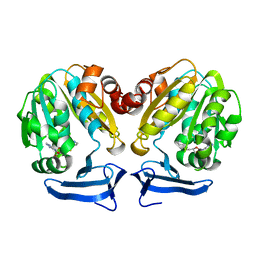 | | Crystal Structure of human Spermidine Synthase in Complex with decarboxylated S-adenosylhomocysteine | | Descriptor: | 5'-S-(3-aminopropyl)-5'-thioadenosine, Spermidine synthase | | Authors: | Seckute, J, McCloskey, D.E, Thomas, H.J, Secrist III, J.A, Pegg, A.E, Ealick, S.E. | | Deposit date: | 2011-05-08 | | Release date: | 2011-09-21 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Binding and inhibition of human spermidine synthase by decarboxylated S-adenosylhomocysteine.
Protein Sci., 20, 2011
|
|
6VBH
 
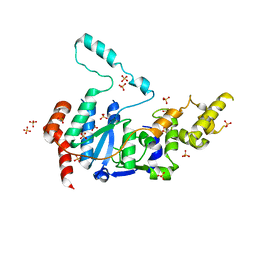 | | Human XPG endonuclease catalytic domain | | Descriptor: | DNA repair protein complementing XP-G cells,Flap endonuclease 1, SULFATE ION | | Authors: | Tsutakawa, S.E, Arvai, A.S, Tainer, J.A. | | Deposit date: | 2019-12-18 | | Release date: | 2020-06-17 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.995 Å) | | Cite: | Human XPG nuclease structure, assembly, and activities with insights for neurodegeneration and cancer from pathogenic mutations.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
6UOS
 
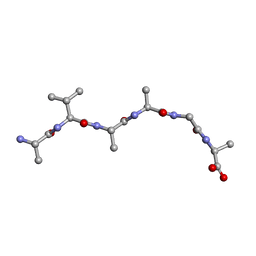 | |
6UOW
 
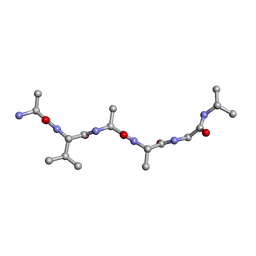 | |
1JPZ
 
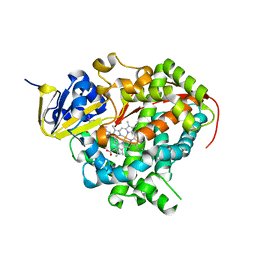 | | Crystal structure of a complex of the heme domain of P450BM-3 with N-Palmitoylglycine | | Descriptor: | BIFUNCTIONAL P-450:NADPH-P450 REDUCTASE, N-PALMITOYLGLYCINE, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Haines, D.C, Tomchick, D.R, Machius, M, Peterson, J.A. | | Deposit date: | 2001-08-03 | | Release date: | 2001-11-09 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Pivotal role of water in the mechanism of P450BM-3.
Biochemistry, 40, 2001
|
|
6UOP
 
 | |
6UOR
 
 | |
6UOU
 
 | |
6UUR
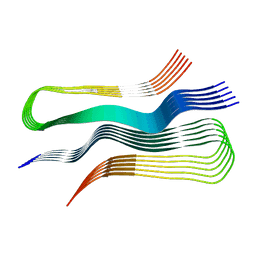 
 | | Human prion protein fibril, M129 variant | | Descriptor: | Major prion protein | | Authors: | Glynn, C, Sawaya, M.R, Ge, P, Zhou, Z.H, Rodriguez, J.A. | | Deposit date: | 2019-10-31 | | Release date: | 2020-04-15 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Cryo-EM structure of a human prion fibril with a hydrophobic, protease-resistant core.
Nat.Struct.Mol.Biol., 27, 2020
|
|
6UVJ
 
 | |
6UOQ
 
 | |
6UPP
 
 | | Radiation Damage Test of PixJ Pb state crystals | | Descriptor: | 1,2-ETHANEDIOL, MAGNESIUM ION, Methyl-accepting chemotaxis protein, ... | | Authors: | Clinger, J.A, Miller, M.D, Burgie, E.S, Vierstra, R.D, Phillips Jr, G.N. | | Deposit date: | 2019-10-18 | | Release date: | 2019-12-18 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Photoreversible interconversion of a phytochrome photosensory module in the crystalline state.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
1I6P
 
 | | CRYSTAL STRUCTURE OF E. COLI BETA CARBONIC ANHYDRASE (ECCA) | | Descriptor: | CARBONIC ANHYDRASE, ZINC ION | | Authors: | Cronk, J.D, Endrizzi, J.A, Cronk, M.R, O'Neill, J.W, Zhang, K.Y.J. | | Deposit date: | 2001-03-02 | | Release date: | 2001-05-09 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of E. coli beta-carbonic anhydrase, an enzyme with an unusual pH-dependent activity.
Protein Sci., 10, 2001
|
|
6UWX
 
 | |
6UPU
 
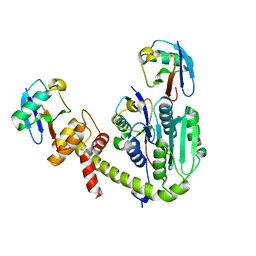 | |
6UPS
 
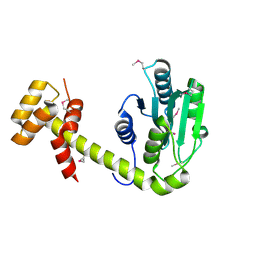 | |
1JQG
 
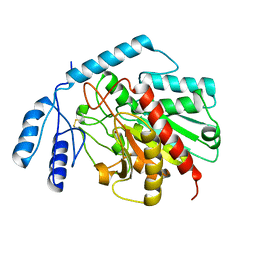 | | Crystal Structure of the Carboxypeptidase A from Helicoverpa Armigera | | Descriptor: | ZINC ION, carboxypeptidase A | | Authors: | Estebanez-Perpina, E, Bayes, A, Vendrell, J, Jongsma, M.A, Bown, D.P, Gatehouse, J.A, Huber, R, Bode, W, Aviles, F.X, Reverter, D. | | Deposit date: | 2001-08-07 | | Release date: | 2002-08-07 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of a novel mid-gut procarboxypeptidase from the cotton pest Helicoverpa armigera.
J.Mol.Biol., 313, 2001
|
|
1JGL
 
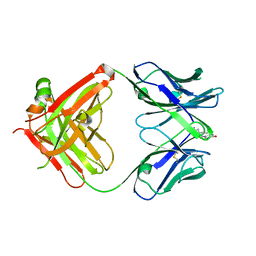 | |
