2JSD
 
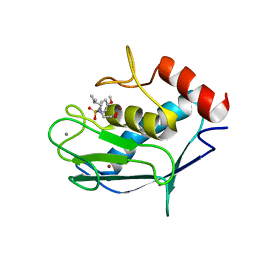 | | Solution structure of MMP20 complexed with NNGH | | Descriptor: | CALCIUM ION, Matrix metalloproteinase-20, N-ISOBUTYL-N-[4-METHOXYPHENYLSULFONYL]GLYCYL HYDROXAMIC ACID, ... | | Authors: | Arendt, Y, Banci, L, Bertini, I, Cantini, F, Cozzi, R, Del Conte, R, Gonnelli, L, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2007-07-03 | | Release date: | 2007-11-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Catalytic domain of MMP20 (Enamelysin) - the NMR structure of a new matrix metalloproteinase.
Febs Lett., 581, 2007
|
|
2JT6
 
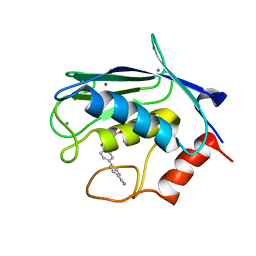 | | Solution structure of matrix metalloproteinase 3 (MMP-3) in the presence of 3-4'-cyanobyphenyl-4-yloxy)-n-hdydroxypropionamide (MMP-3 inhibitor VII) | | Descriptor: | 3-[(4'-cyanobiphenyl-4-yl)oxy]-N-hydroxypropanamide, CALCIUM ION, Stromelysin-1, ... | | Authors: | Alcaraz, L.A, Banci, L, Bertini, I, Cantini, F, Donaire, A, Gonnelli, L. | | Deposit date: | 2007-07-23 | | Release date: | 2008-02-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Matrix metalloproteinase-inhibitor interaction: the solution structure of the catalytic domain of human matrix metalloproteinase-3 with different inhibitors
J.Biol.Inorg.Chem., 12, 2007
|
|
2JTA
 
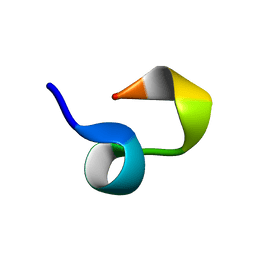 | |
5CJ0
 
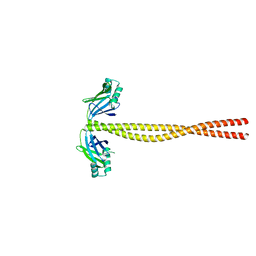 | | Crystal Structure of Amino Acids 1631-1692 of MYH7 | | Descriptor: | Xrcc4-MYH7-(1631-1692) chimera protein | | Authors: | Korkmaz, N.E, Taylor, K.C, Andreas, M.P, Ajay, G, Heinze, N.T, Cui, Q, Rayment, I. | | Deposit date: | 2015-07-13 | | Release date: | 2015-12-02 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A composite approach towards a complete model of the myosin rod.
Proteins, 84, 2016
|
|
8RDD
 
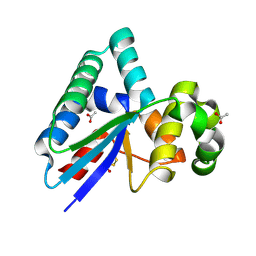 | |
2BXA
 
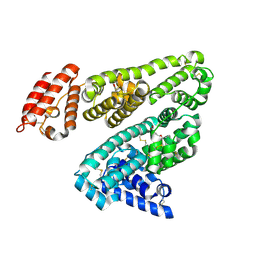 | | Human serum albumin complexed with 3-carboxy-4-methyl-5-propyl-2- furanpropanoic acid (CMPF) | | Descriptor: | 3-CARBOXY-4-METHYL-5-PROPYL-2-FURANPROPIONIC, SERUM ALBUMIN | | Authors: | Ghuman, J, Zunszain, P.A, Petitpas, I, Bhattacharya, A.A, Otagiri, M, Curry, S. | | Deposit date: | 2005-07-26 | | Release date: | 2005-09-22 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural Basis of the Drug-Binding Specificity of Human Serum Albumin.
J.Mol.Biol., 353, 2005
|
|
3OA7
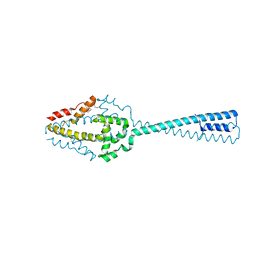 
 | | Structure of the C-terminal domain of Cnm67, a core component of the spindle pole body of Saccharomyces cerevisiae | | Descriptor: | Head morphogenesis protein, Chaotic nuclear migration protein 67 fusion protein | | Authors: | Klenchin, V.A, Frye, J.J, Rayment, I. | | Deposit date: | 2010-08-04 | | Release date: | 2011-03-23 | | Last modified: | 2017-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure-function analysis of the C-terminal domain of CNM67, a core component of the Saccharomyces cerevisiae spindle pole body.
J.Biol.Chem., 286, 2011
|
|
1AUL
 
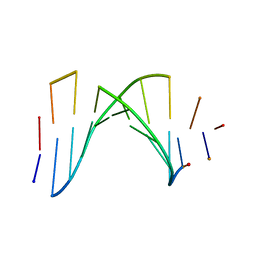 | | SOLUTION STRUCTURE OF A HIGHLY STABLE DNA DUPLEX CONJUGATED TO A MINOR GROOVE BINDER, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | DNA (5'-D(*CP*AP*GP*AP*TP*AP*AP*TP*CP*A)-3'), DNA (5'-D(P*THXP*GP*AP*TP*TP*AP*TP*CP*TP*G)-3') | | Authors: | Kumar, S, Reed, M.W, Gamper Junior, H.B, Gorn, V.V, Lukhtanov, E.A, Foti, M, West, J, Meyer Junior, R.B, Schweitzer, B.I. | | Deposit date: | 1997-08-29 | | Release date: | 1997-12-24 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a highly stable DNA duplex conjugated to a minor groove binder.
Nucleic Acids Res., 26, 1998
|
|
2BMB
 
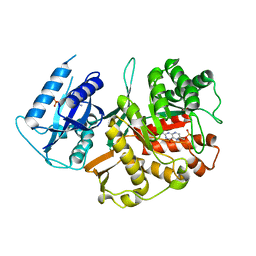 | | X-ray structure of the bifunctional 6-hydroxymethyl-7,8- dihydroxypterin pyrophosphokinase dihydropteroate synthase from Saccharomyces cerevisiae | | Descriptor: | FOLIC ACID SYNTHESIS PROTEIN FOL1, PTERIN-6-YL-METHYL-MONOPHOSPHATE | | Authors: | Lawrence, M.C, Iliades, P, Fernley, R.T, Berglez, J, Pilling, P.A, Macreadie, I.G. | | Deposit date: | 2005-03-11 | | Release date: | 2006-01-17 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Three-Dimensional Structure of the Bifunctional 6-Hydroxymethyl-7,8-Dihydropterin Pyrophosphokinase/Dihydropteroate Synthase of Saccharomyces Cerevisiae
J.Mol.Biol., 348, 2005
|
|
2K3J
 
 | | The solution structure of human Mia40 | | Descriptor: | Mitochondrial intermembrane space import and assembly protein 40 | | Authors: | Ciofi Baffoni, S, Bertini, I, Gallo, A. | | Deposit date: | 2008-05-08 | | Release date: | 2009-02-10 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | MIA40 is an oxidoreductase that catalyzes oxidative protein folding in mitochondria.
Nat.Struct.Mol.Biol., 16, 2009
|
|
2JP9
 
 | | Structure of the Wilms Tumor Suppressor Protein Zinc Finger Domain Bound to DNA | | Descriptor: | DNA (5'-D(P*DCP*DGP*DCP*DGP*DGP*DGP*DGP*DGP*DCP*DGP*DTP*DCP*DTP*DGP*DCP*DGP*DC)-3'), DNA (5'-D(P*DGP*DCP*DGP*DCP*DAP*DGP*DAP*DCP*DGP*DCP*DCP*DCP*DCP*DCP*DGP*DCP*DG)-3'), Wilms tumor 1, ... | | Authors: | Stoll, R, Lee, B.M, Debler, E.W, Laity, J.H, Wilson, I.A, Dyson, H.J, Wright, P.E. | | Deposit date: | 2007-04-30 | | Release date: | 2007-10-30 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Structure of the wilms tumor suppressor protein zinc finger domain bound to DNA
J.Mol.Biol., 372, 2007
|
|
3NYF
 
 | |
2K2M
 
 | | Structural Basis of PxxDY Motif Recognition in SH3 Binding | | Descriptor: | Eps8-like protein 1 | | Authors: | Aitio, O, Hellman, M, Kesti, T, Kleino, I, Samuilova, O, Tossavainen, H, Saksela, K, Permi, P. | | Deposit date: | 2008-04-02 | | Release date: | 2009-02-17 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Structural basis of PxxDY motif recognition in SH3 binding.
J.Mol.Biol., 382, 2008
|
|
2K3T
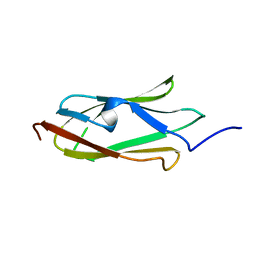 
 | |
5CG6
 
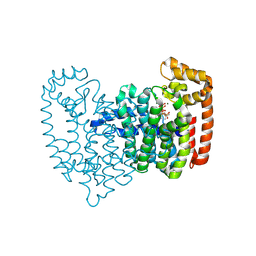 | | Neutron crystal structure of human farnesyl pyrophosphate synthase in complex with risedronate and isopentenyl pyrophosphate | | Descriptor: | 1-HYDROXY-2-(3-PYRIDINYL)ETHYLIDENE BIS-PHOSPHONIC ACID, 3-METHYLBUT-3-ENYL TRIHYDROGEN DIPHOSPHATE, Farnesyl pyrophosphate synthase, ... | | Authors: | Yokoyama, T, Mizuguchi, M, Ostermann, A, Kusaka, K, Niimura, N, Schrader, T.E, Tanaka, I. | | Deposit date: | 2015-07-09 | | Release date: | 2015-10-14 | | Last modified: | 2024-04-03 | | Method: | NEUTRON DIFFRACTION (1.7 Å), X-RAY DIFFRACTION | | Cite: | Protonation State and Hydration of Bisphosphonate Bound to Farnesyl Pyrophosphate Synthase
J.Med.Chem., 58, 2015
|
|
3O3T
 
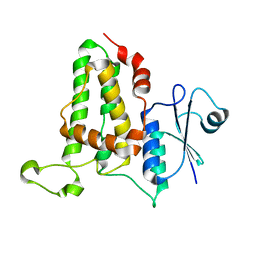 | | Crystal Structure Analysis of M32A mutant of human CLIC1 | | Descriptor: | Chloride intracellular channel protein 1 | | Authors: | Fanucchi, S, Achilonu, I.A, Adamson, R.J, Fernandes, M.A, Stoychev, S, Dirr, H.W. | | Deposit date: | 2010-07-26 | | Release date: | 2010-08-11 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structure Analysis of M32A mutant of human CLIC1
To be Published
|
|
2I5D
 
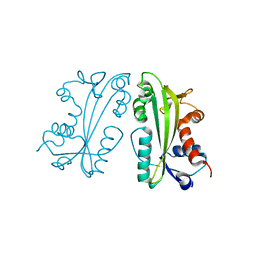 | | Crystal Structure of Human Inosine Triphosphate Pyrophosphatase | | Descriptor: | inosine triphosphate pyrophosphohydrolase | | Authors: | Porta, J.C, Kozmin, S.G, Pavlov, Y.I, Borgstahl, G.E.O. | | Deposit date: | 2006-08-24 | | Release date: | 2006-09-05 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Structure of the orthorhombic form of human inosine triphosphate pyrophosphatase.
Acta Crystallogr.,Sect.F, 62, 2006
|
|
1BEE
 
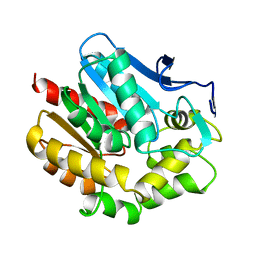 | | HALOALKANE DEHALOGENASE MUTANT WITH TRP 175 REPLACED BY TYR | | Descriptor: | HALOALKANE DEHALOGENASE | | Authors: | Ridder, I.S, Vos, G.J, Rozeboom, H.J, Kalk, K.H, Dijkstra, B.W. | | Deposit date: | 1998-05-13 | | Release date: | 1998-11-11 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Kinetic analysis and X-ray structure of haloalkane dehalogenase with a modified halide-binding site.
Biochemistry, 37, 1998
|
|
5CIW
 
 | | Ran GDP Y39A mutant monoclinic crystal form | | Descriptor: | GTP-binding nuclear protein Ran, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION | | Authors: | Vetter, I.R, Brucker, S. | | Deposit date: | 2015-07-13 | | Release date: | 2015-09-09 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Catalysis of GTP Hydrolysis by Small GTPases at Atomic Detail by Integration of X-ray Crystallography, Experimental, and Theoretical IR Spectroscopy.
J.Biol.Chem., 290, 2015
|
|
2BSC
 
 | | E. coli F17a-G lectin domain complex with N-acetylglucosamine, high- resolution structure | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, F17A-G ADHESIN | | Authors: | Buts, L, Wellens, A, Van Molle, I, Wyns, L, Loris, R, Lahmann, M, Oscarson, S, De Greve, H, Bouckaert, J. | | Deposit date: | 2005-05-20 | | Release date: | 2006-05-24 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Impact of Natural Variation in Bacterial F17G Adhesins on Crystallization Behaviour.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
3NQ9
 
 | | Bovine beta-lactoglobulin complex with caprylic acid | | Descriptor: | Beta-lactoglobulin, CHLORIDE ION, OCTANOIC ACID (CAPRYLIC ACID) | | Authors: | Loch, J.I, Lewinski, K. | | Deposit date: | 2010-06-29 | | Release date: | 2010-07-14 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Two modes of fatty acid binding to bovine beta-lactoglobulin-crystallographic and spectroscopic studies
J.Mol.Recognit., 24, 2011
|
|
5CIQ
 
 | | Ran GDP wild type tetragonal crystal form | | Descriptor: | GTP-binding nuclear protein Ran, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION | | Authors: | Vetter, I.R, Brucker, S. | | Deposit date: | 2015-07-13 | | Release date: | 2015-09-09 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Catalysis of GTP Hydrolysis by Small GTPases at Atomic Detail by Integration of X-ray Crystallography, Experimental, and Theoretical IR Spectroscopy.
J.Biol.Chem., 290, 2015
|
|
5CIT
 
 | | Ran GDP wild type monoclinic crystal form | | Descriptor: | GTP-binding nuclear protein Ran, GUANOSINE-5'-DIPHOSPHATE, MAGNESIUM ION | | Authors: | Vetter, I.R, Brucker, S. | | Deposit date: | 2015-07-13 | | Release date: | 2015-09-09 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Catalysis of GTP Hydrolysis by Small GTPases at Atomic Detail by Integration of X-ray Crystallography, Experimental, and Theoretical IR Spectroscopy.
J.Biol.Chem., 290, 2015
|
|
5CJ4
 
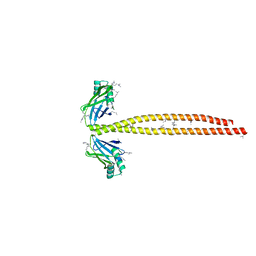 | | Crystal Structure of Amino Acids 1562-1622 of MYH7 | | Descriptor: | Xrcc4-MYH7-(1562-1622) chimera protein | | Authors: | Korkmaz, N.E, Taylor, K.C, Andreas, M.P, Ajay, G, Heinze, N.T, Cui, Q, Rayment, I. | | Deposit date: | 2015-07-13 | | Release date: | 2015-12-02 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.102 Å) | | Cite: | A composite approach towards a complete model of the myosin rod.
Proteins, 84, 2016
|
|
1B45
 
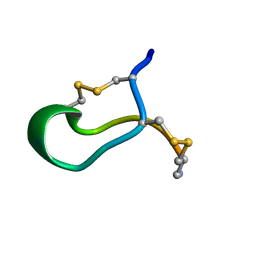 | | ALPHA-CNIA CONOTOXIN FROM CONUS CONSORS, NMR, 43 STRUCTURES | | Descriptor: | ALPHA-CNIA | | Authors: | Favreau, P, Krimm, I, Le Gall, F, Bobenrieth, M.J, Lamthanh, H, Bouet, F, Servent, D, Molgo, J, Menez, A, Letourneux, Y, Lancelin, J.M. | | Deposit date: | 1999-01-05 | | Release date: | 1999-07-09 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Biochemical characterization and nuclear magnetic resonance structure of novel alpha-conotoxins isolated from the venom of Conus consors.
Biochemistry, 38, 1999
|
|
