7S7W
 
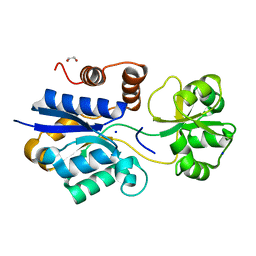 | | Crystal structure of iNicSnFR3a Nicotine Sensor precursor binding protein | | Descriptor: | 1,2-ETHANEDIOL, SODIUM ION, iNicSnFR 3.0 Fluorescent Nicotine Sensor precursor binding protein | | Authors: | Fan, C, Shivange, A.V, Looger, L.L, Lester, H.A, Rees, D.C. | | Deposit date: | 2021-09-17 | | Release date: | 2021-10-13 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Structure, Function, and Application of Bacterial ABC Transporters
Ph.D.Thesis,California Institute of Technology, 2020
|
|
7S7T
 
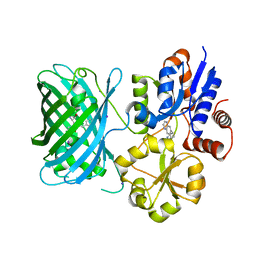 | | iNicSnFR3a Nicotine Sensor comprising Periplasmic Binding sequence plus Fluorescent Sequence with varenicline bound | | Descriptor: | IODIDE ION, VARENICLINE, iNicSnFR 3.0 Fluorescent Nicotine Sensor | | Authors: | Fan, C, Shivange, A.V, Looger, L.L, Lester, H.A, Rees, D.C. | | Deposit date: | 2021-09-17 | | Release date: | 2021-10-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Correction: Fluorescence activation mechanism and imaging of drug permeation with new sensors for smoking-cessation ligands.
Elife, 11, 2022
|
|
7S80
 
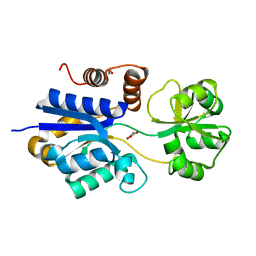 | | Crystal structure of iAChSnFR Acetylcholine Sensor precursor binding protein | | Descriptor: | 1,2-ETHANEDIOL, MALONATE ION, iAchSnFR Fluorescent Acetylcholine Sensor precursor binding protein | | Authors: | Fan, C, Borden, P.M, Looger, L.L, Lester, H.A, Rees, D.C. | | Deposit date: | 2021-09-17 | | Release date: | 2021-10-13 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structure, Function, and Application of Bacterial ABC Transporters
Ph.D.Thesis,California Institute of Technology, 2020
|
|
7S7Z
 
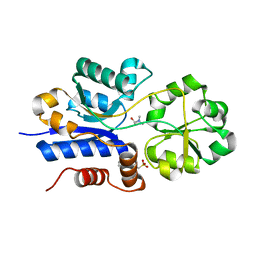 | | Crystal structure of iAChSnFR Acetylcholine Sensor precursor binding protein with choline bound | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, CHOLINE ION, iAchSnFR Fluorescent Acetylcholine Sensor precursor binding protein | | Authors: | Fan, C, Borden, P.M, Looger, L.L, Lester, H.A, Rees, D.C. | | Deposit date: | 2021-09-17 | | Release date: | 2021-10-13 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure, Function, and Application of Bacterial ABC Transporters
Ph.D.Thesis,California Institute of Technology, 2020
|
|
7S7X
 
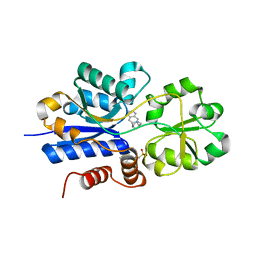 | | Crystal structure of iCytSnFR Cytisine Sensor precursor binding protein with varenicline bound | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, DI(HYDROXYETHYL)ETHER, VARENICLINE, ... | | Authors: | Fan, C, Nichols, N.L, Luebbert, L, Looger, L.L, Lester, H.A, Rees, D.C. | | Deposit date: | 2021-09-17 | | Release date: | 2021-10-13 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure, Function, and Application of Bacterial ABC Transporters
Ph.D.Thesis,California Institute of Technology, 2020
|
|
7S7U
 
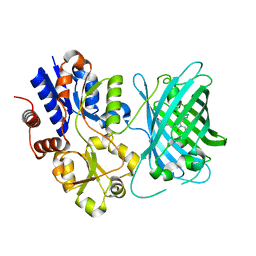 | | Crystal structure of iNicSnFR3a Fluorescent Nicotine Sensor with nicotine bound | | Descriptor: | iNicSnFR 3.0 Fluorescent Nicotine Sensor | | Authors: | Fan, C, Shivange, A.V, Looger, L.L, Lester, H.A, Rees, D.C. | | Deposit date: | 2021-09-17 | | Release date: | 2021-10-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Correction: Fluorescence activation mechanism and imaging of drug permeation with new sensors for smoking-cessation ligands.
Elife, 11, 2022
|
|
1EYB
 
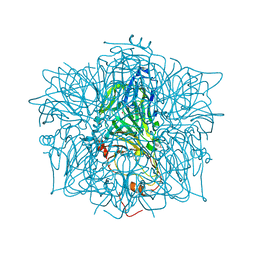 | | CRYSTAL STRUCTURE OF APO HUMAN HOMOGENTISATE DIOXYGENASE | | Descriptor: | HOMOGENTISATE 1,2-DIOXYGENASE | | Authors: | Timm, D.E, Titus, G.P, Penalva, M.A, Mueller, H.A, de Cordoba, S.M. | | Deposit date: | 2000-05-05 | | Release date: | 2000-11-05 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of human homogentisate dioxygenase.
Nat.Struct.Biol., 7, 2000
|
|
1E95
 
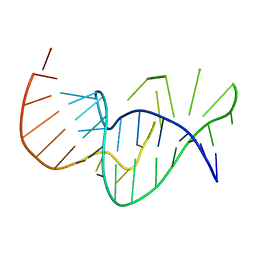 | | Solution structure of the pseudoknot of SRV-1 RNA, involved in ribosomal frameshifting | | Descriptor: | RNA (5'-(*GP*CP*GP*GP*CP*CP*AP*GP*CP*UP*CP* CP*AP*GP*GP*CP*CP*GP*CP*CP*AP*AP*AP*CP* AP*AP*UP*AP*UP*GP*GP*AP*GP*CP*AP*C)-3') | | Authors: | Michiels, P.J.A, Versleyen, A, Pleij, C.W.A, Hilbers, C.W, Heus, H.A. | | Deposit date: | 2000-10-09 | | Release date: | 2001-08-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Pseudoknot of Srv-1 RNA, Involved in Ribosomal Frameshifting
J.Mol.Biol., 310, 2001
|
|
1EGF
 
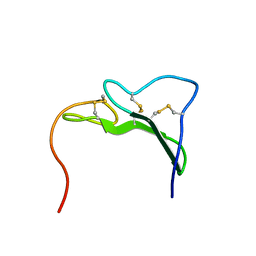 | |
1E0E
 
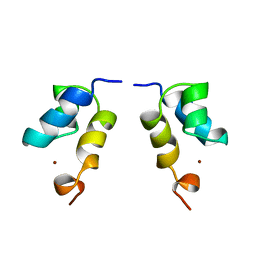 | | N-terminal zinc-binding HHCC domain of HIV-2 integrase | | Descriptor: | HUMAN IMMUNODEFICIENCY VIRUS TYPE 2 INTEGRASE, ZINC ION | | Authors: | Eijkelenboom, A.P.A.M, Van Den ent, F.M.I, Plasterk, R.H.A, Kaptein, R, Boelens, R. | | Deposit date: | 2000-03-25 | | Release date: | 2001-03-19 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Refined Solution Structure of the Dimeric N-Terminal Hhcc Domain of HIV-2 Integrase
J.Biomol.NMR, 18, 2000
|
|
1DTJ
 
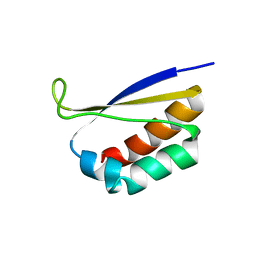 | | CRYSTAL STRUCTURE OF NOVA-2 KH3 K-HOMOLOGY RNA-BINDING DOMAIN | | Descriptor: | RNA-BINDING NEUROONCOLOGICAL VENTRAL ANTIGEN 2 | | Authors: | Lewis, H.A, Chen, H, Edo, C, Buckanovich, R.J, Yang, Y.Y.L, Musunuru, K, Zhong, R, Darnell, R.B, Burley, S.K. | | Deposit date: | 2000-01-12 | | Release date: | 2000-02-18 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of Nova-1 and Nova-2 K-homology RNA-binding domains.
Structure Fold.Des., 7, 1999
|
|
3IXY
 
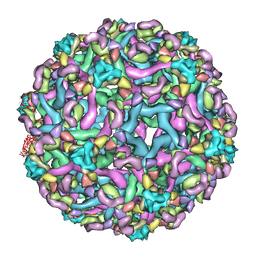 | | The pseudo-atomic structure of dengue immature virus in complex with Fab fragments of the anti-fusion loop antibody E53 | | Descriptor: | E53 Fab Fragment (chain H), E53 Fab Fragment (chain L), Envelope protein E, ... | | Authors: | Cherrier, M.V, Kaufmann, B, Nybakken, G.E, Lok, S.M, Warren, J.T, Nelson, C.A, Kostyuchenko, V.A, Holdaway, H.A, Chipman, P.R, Kuhn, R.J, Diamond, M.S, Rossmann, M.G, Fremont, D.H. | | Deposit date: | 2009-02-26 | | Release date: | 2009-10-27 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (23 Å) | | Cite: | Structural basis for the preferential recognition of immature flaviviruses by a fusion-loop antibody
Embo J., 28, 2009
|
|
2N3M
 
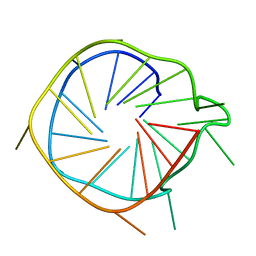 | |
2N4T
 | |
3ISO
 
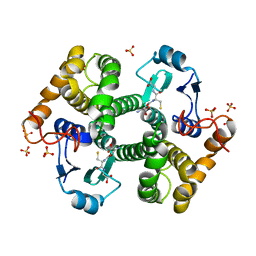 | | Crystal structure of 26 kDa GST of Clonorchis sinensis in P3221 symmetry | | Descriptor: | GLUTATHIONE, Putative glutathione transferase, SULFATE ION, ... | | Authors: | Han, Y.H, Seo, H.A, Kim, G.H, Chung, Y.J. | | Deposit date: | 2009-08-27 | | Release date: | 2010-09-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A histidine substitution confers metal binding affinity to a Schistosoma japonicum Glutathione S-transferase.
Protein Expr.Purif., 73, 2010
|
|
3IXX
 
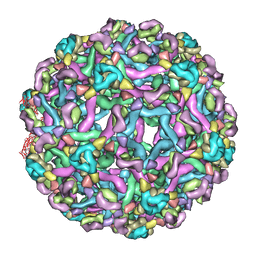 | | The pseudo-atomic structure of West Nile immature virus in complex with Fab fragments of the anti-fusion loop antibody E53 | | Descriptor: | E53 Fab Fragment (chain H), E53 Fab Fragment (chain L), Envelope protein E, ... | | Authors: | Cherrier, M.V, Kaufmann, B, Nybakken, G.E, Lok, S.M, Warren, J.T, Nelson, C.A, Kostyuchenko, V.A, Holdaway, H.A, Chipman, P.R, Kuhn, R.J, Diamond, M.S, Rossmann, M.G, Fremont, D.H. | | Deposit date: | 2009-02-26 | | Release date: | 2009-10-27 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (15 Å) | | Cite: | Structural basis for the preferential recognition of immature flaviviruses by a fusion-loop antibody
Embo J., 28, 2009
|
|
3IYA
 
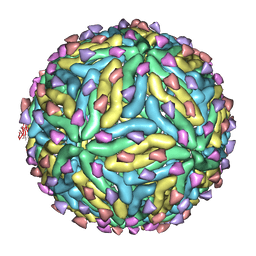 | | Association of the pr peptides with dengue virus blocks membrane fusion at acidic pH | | Descriptor: | Envelope protein, prM protein | | Authors: | Yu, I, Holdaway, H.A, Chipman, P.R, Kuhn, R.J, Rossmann, M.G, Chen, J. | | Deposit date: | 2009-06-01 | | Release date: | 2010-04-14 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (22 Å) | | Cite: | Association of the pr peptides with dengue virus at acidic pH blocks membrane fusion.
J.Virol., 83, 2009
|
|
3JXO
 
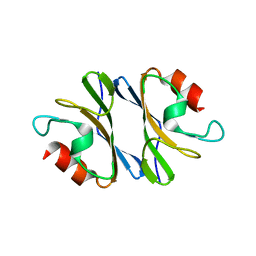 | | Crystal Structure of an Octomeric Two-Subunit TrkA K+ Channel Ring Gating Assembly, TM1088A:TM1088B, from Thermotoga maritima | | Descriptor: | TrkA-N domain protein | | Authors: | Deller, M.C, Johnson, H.A, Miller, M, Spraggon, G, Wilson, I.A, Lesley, S.A, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2009-09-20 | | Release date: | 2009-10-06 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Crystal Structure of an Octomeric Two-Subunit TrkA K+ Channel Ring Gating Assembly, TM1088A:TM1088B, from Thermotoga maritima
To be Published
|
|
2NV7
 
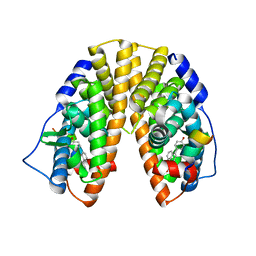 | | Crystal Structure of Estrogen Receptor Beta Complexed with WAY-555 | | Descriptor: | 4-(4-HYDROXYPHENYL)-1-NAPHTHALDEHYDE OXIME, Estrogen receptor beta, Nuclear receptor coactivator 1 | | Authors: | Mewshaw, R.E, Bowen, M.S, Harris, H.A, Xu, Z.B, Manas, E.S, Cohn, S.T, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-11-10 | | Release date: | 2007-08-21 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | ERbeta ligands. Part 5: synthesis and structure-activity relationships of a series of 4'-hydroxyphenyl-aryl-carbaldehyde oxime derivatives.
Bioorg.Med.Chem.Lett., 17, 2007
|
|
1CTL
 
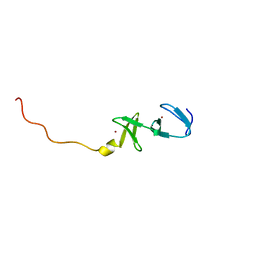 | | STRUCTURE OF THE CARBOXY-TERMINAL LIM DOMAIN FROM THE CYSTEINE RICH PROTEIN CRP | | Descriptor: | AVIAN CYSTEINE RICH PROTEIN, ZINC ION | | Authors: | Perez-Alvarado, G.C, Miles, C, Michelsen, J.W, Louis, H.A, Winge, D.R, Beckerle, M.C, Summers, M.F. | | Deposit date: | 1995-01-06 | | Release date: | 1995-06-03 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the carboxy-terminal LIM domain from the cysteine rich protein CRP.
Nat.Struct.Biol., 1, 1994
|
|
3I5U
 
 | |
7TEF
 
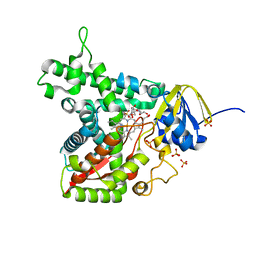 | | Cytochrome P450 14 alpha-sterol demethylase CYP51 from Mycobacterium marinum | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Cytochrome P450 51B1 Cyp51B1, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Mohamed, H.A, Bruning, J.B, Bell, S.G. | | Deposit date: | 2022-01-04 | | Release date: | 2022-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | A comparison of the bacterial CYP51 cytochrome P450 enzymes from Mycobacterium marinum and Mycobacterium tuberculosis.
J.Steroid Biochem.Mol.Biol., 221, 2022
|
|
1DU2
 
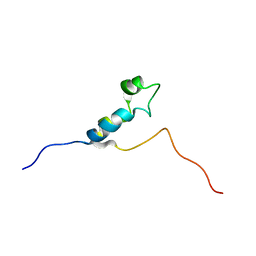 | | SOLUTION STRUCTURE OF THE THETA SUBUNIT OF DNA POLYMERASE III | | Descriptor: | DNA POLYMERASE III | | Authors: | Keniry, M.A, Berthon, H.A, Yang, J.-Y, Miles, C.S, Dixon, N.E. | | Deposit date: | 2000-01-13 | | Release date: | 2000-05-31 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of the theta subunit of DNA polymerase III from Escherichia coli.
Protein Sci., 9, 2000
|
|
1DT4
 
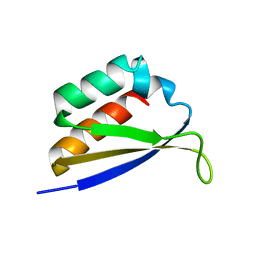 | | CRYSTAL STRUCTURE OF NOVA-1 KH3 K-HOMOLOGY RNA-BINDING DOMAIN | | Descriptor: | NEURO-ONCOLOGICAL VENTRAL ANTIGEN 1 | | Authors: | Lewis, H.A, Chen, H, Edo, C, Buckanovich, R.J, Yang, Y.Y.L. | | Deposit date: | 2000-01-11 | | Release date: | 2000-02-11 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structures of Nova-1 and Nova-2 K-homology RNA-binding domains.
Structure Fold.Des., 7, 1999
|
|
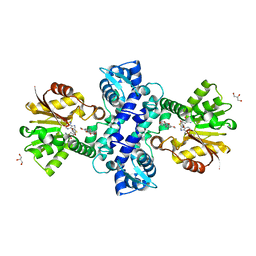
3I58
 
 | |
