4V62
 
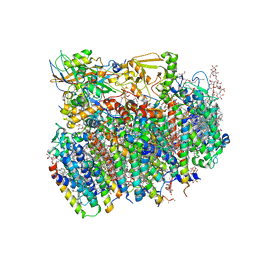 | | Crystal Structure of cyanobacterial Photosystem II | | Descriptor: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, ... | | Authors: | Guskov, A, Gabdulkhakov, A, Kern, J, Broser, M, Zouni, A, Saenger, W. | | Deposit date: | 2008-01-17 | | Release date: | 2014-07-09 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Cyanobacterial photosystem II at 2.9-A resolution and the role of quinones, lipids, channels and chloride
Nat.Struct.Mol.Biol., 16, 2009
|
|
4EGW
 
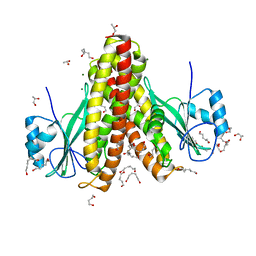 | | The structure of the soluble domain of CorA from Methanocaldococcus jannaschii | | Descriptor: | 1,4-BUTANEDIOL, HEXANE-1,6-DIOL, MAGNESIUM ION, ... | | Authors: | Guskov, A, Nordin, N, Reynaud, A, Engman, H, Lundback, A.-K, Jong, A.J.O, Cornvik, T, Phua, T, Eshaghi, S. | | Deposit date: | 2012-04-02 | | Release date: | 2012-10-31 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insights into the mechanisms of Mg2+ uptake, transport, and gating by CorA
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4EV6
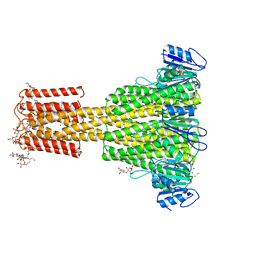 
 | | The complete structure of CorA magnesium transporter from Methanocaldococcus jannaschii | | Descriptor: | MAGNESIUM ION, Magnesium transport protein CorA, UNDECYL-MALTOSIDE | | Authors: | Guskov, A, Nordin, N, Reynaud, A, Engman, H, Lundback, A.-K, Jong, A.J.O, Cornvik, T, Phua, T, Eshaghi, S. | | Deposit date: | 2012-04-25 | | Release date: | 2012-10-31 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural insights into the mechanisms of Mg2+ uptake, transport, and gating by CorA
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
5DWY
 
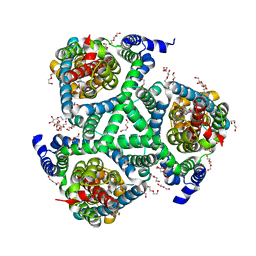 | |
5E9S
 
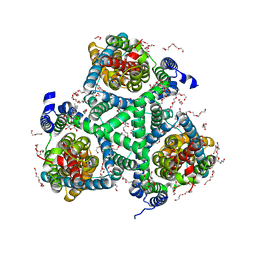 | |
5D3M
 
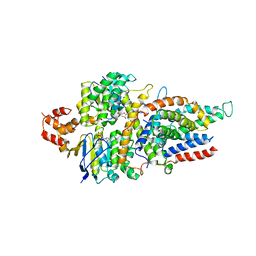 | | Folate ECF transporter: AMPPNP bound state | | Descriptor: | Energy-coupling factor transporter ATP-binding protein EcfA1, Energy-coupling factor transporter ATP-binding protein EcfA2, Energy-coupling factor transporter transmembrane protein EcfT, ... | | Authors: | Guskov, A, Swier, L.J.Y.M, Slotboom, D.J. | | Deposit date: | 2015-08-06 | | Release date: | 2016-04-06 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.303 Å) | | Cite: | Structural insight in the toppling mechanism of an energy-coupling factor transporter.
Nat Commun, 7, 2016
|
|
4QTN
 
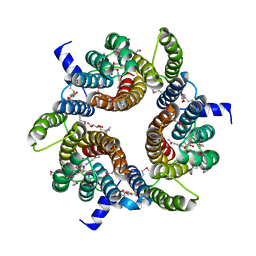 | |
6FXG
 
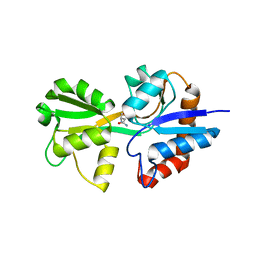 | |
4KY0
 
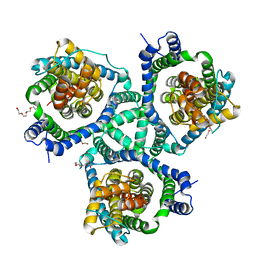 | | Crystal structure of a substrate-free glutamate transporter homologue from Thermococcus kodakarensis | | Descriptor: | Proton/glutamate symporter, SDF family, TETRAETHYLENE GLYCOL | | Authors: | Guskov, A, Jensen, S, Rempel, S, Hanelt, I, Slotboom, D.J. | | Deposit date: | 2013-05-28 | | Release date: | 2013-09-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of a substrate-free aspartate transporter.
Nat.Struct.Mol.Biol., 20, 2013
|
|
4LA9
 
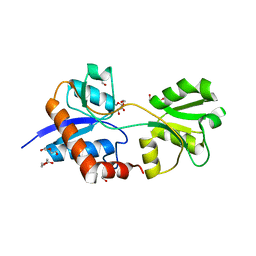 | | Crystal structure of an empty substrate binding domain 1 (SBD1) OF ABC TRANSPORTER GLNPQ from lactococcus lactis | | Descriptor: | DI(HYDROXYETHYL)ETHER, Glutamine ABC transporter permease and substrate binding protein protein, PHOSPHATE ION | | Authors: | Guskov, A, Schuurman-Wolters, G.K, Slotboom, D.J, Poolman, B. | | Deposit date: | 2013-06-19 | | Release date: | 2013-09-11 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Functional Diversity of Tandem Substrate-Binding Domains in ABC Transporters from Pathogenic Bacteria.
Structure, 21, 2013
|
|
4V82
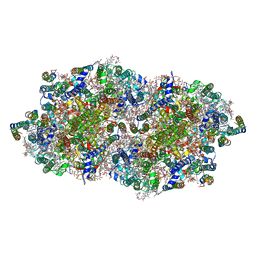 
 | | Crystal structure of cyanobacterial Photosystem II in complex with terbutryn | | Descriptor: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, ... | | Authors: | Gabdulkhakov, A, Broser, M, Guskov, A, Kern, J, Glockner, C, Muh, F, Saenger, W, Zouni, A. | | Deposit date: | 2010-11-30 | | Release date: | 2014-07-09 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural basis of cyanobacterial photosystem II Inhibition by the herbicide terbutryn
J.Biol.Chem., 286, 2011
|
|
3KZI
 
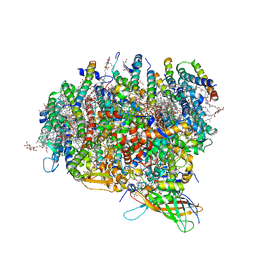 | | Crystal Structure of Monomeric Form of Cyanobacterial Photosystem II | | Descriptor: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, ... | | Authors: | Gabdulkhakov, A, Guskov, A, Broser, M, Kern, J, Zouni, A, Saenger, W. | | Deposit date: | 2009-12-08 | | Release date: | 2010-06-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Crystal Structure of Monomeric Photosystem II from Thermosynechococcus elongatus at 3.6-A Resolution
J.Biol.Chem., 285, 2010
|
|
7QZ1
 
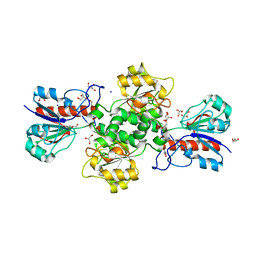 | | Formate dehydrogenase from Starkeya novella | | Descriptor: | CHLORIDE ION, DI(HYDROXYETHYL)ETHER, Formate dehydrogenase, ... | | Authors: | Pontillo, N, Slotboom, D.J, Guskov, A. | | Deposit date: | 2022-01-30 | | Release date: | 2023-02-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Biochemical and structural insight into the chemical resistance and cofactor specificity of the formate dehydrogenase from Starkeya novella.
Febs J., 290, 2023
|
|
4G4P
 
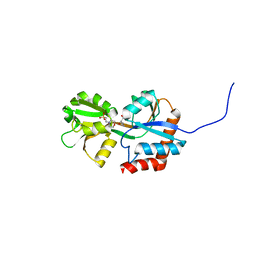 | | Crystal structure of glutamine-binding protein from Enterococcus faecalis at 1.5 A | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Amino acid ABC transporter, amino acid-binding/permease protein, ... | | Authors: | Fulyani, F, Guskov, A, Zagar, A.V, Slotboom, D.-J, Poolman, B. | | Deposit date: | 2012-07-16 | | Release date: | 2013-07-17 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Functional Diversity of Tandem Substrate-Binding Domains in ABC Transporters from Pathogenic Bacteria.
Structure, 21, 2013
|
|
4ZEF
 
 | |
8ODS
 
 | |
5JSZ
 
 | | Folate ECF transporter: apo state | | Descriptor: | Conserved hypothetical membrane protein, Energy-coupling factor transporter ATP-binding protein EcfA1, Energy-coupling factor transporter ATP-binding protein EcfA2, ... | | Authors: | Swier, L.J.Y.M, Guskov, A, Slotboom, D.J. | | Deposit date: | 2016-05-09 | | Release date: | 2016-05-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.004 Å) | | Cite: | Structural insight in the toppling mechanism of an energy-coupling factor transporter.
Nat Commun, 7, 2016
|
|
4I0U
 
 | | Improved structure of Thermotoga maritima CorA at 2.7 A resolution | | Descriptor: | 2-(2-METHOXYETHOXY)ETHANOL, CHLORIDE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Nordin, N, Guskov, A, Phua, T, Sahaf, N, Xia, Y, Lu, S.Y, Eshaghi, H, Eshaghi, S. | | Deposit date: | 2012-11-19 | | Release date: | 2013-03-06 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Exploring the structure and function of Thermotoga maritima CorA reveals the mechanism of gating and ion selectivity in Co2+/Mg2+ transport.
Biochem.J., 451, 2013
|
|
2XZ5
 
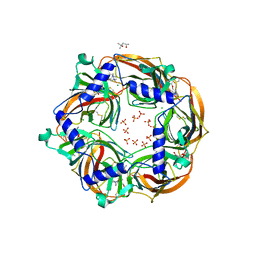 | | MMTS-modified Y53C mutant of Aplysia AChBP in complex with acetylcholine | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ACETYLCHOLINE, ... | | Authors: | Brams, M, Gay, E.A, Colon Saez, J, Guskov, A, van Elk, R, van der Schors, R.C, Peigneur, S, Tytgat, J, Strelkov, S.V, Smit, A.B, Yakel, J.L, Ulens, C. | | Deposit date: | 2010-11-23 | | Release date: | 2010-12-08 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structures of a Cysteine-Modified Mutant in Loop D of Acetylcholine Binding Protein
J.Biol.Chem., 286, 2011
|
|
8AFA
 
 | |
5N9Y
 
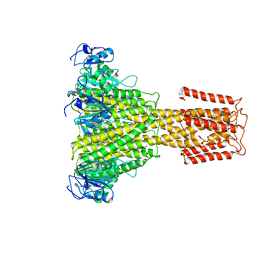 | | The full-length structure of ZntB | | Descriptor: | Zinc transport protein ZntB | | Authors: | Cornelius, G, Stetsenko, A, Scheres, S.H.W, Slotboom, D.J, Guskov, A. | | Deposit date: | 2017-02-27 | | Release date: | 2017-11-15 | | Last modified: | 2019-10-23 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | The structural basis of proton driven zinc transport by ZntB.
Nat Commun, 8, 2017
|
|
7P4I
 
 | |
7NH9
 
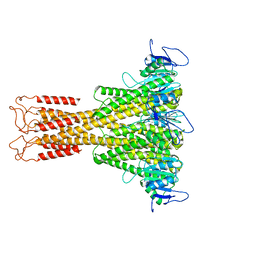 | |
8BK6
 
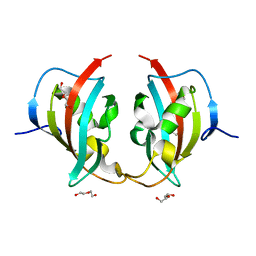 | | A truncated structure of LpMIP with bound inhibitor JK095. | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, DI(HYDROXYETHYL)ETHER, Peptidyl-prolyl cis-trans isomerase | | Authors: | Whittaker, J.J, Guskov, A, Hellmich, A.U, Goretzki, B. | | Deposit date: | 2022-11-08 | | Release date: | 2023-09-06 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.263 Å) | | Cite: | Legionella pneumophila macrophage infectivity potentiator protein appendage domains modulate protein dynamics and inhibitor binding.
Int.J.Biol.Macromol., 252, 2023
|
|
5D0Y
 
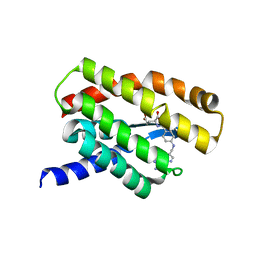 | |
