2LBS
 
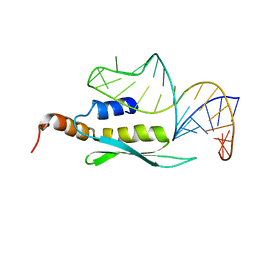 | | Solution structure of double-stranded RNA binding domain of S. cerevisiae RNase III (Rnt1p) in complex with AAGU tetraloop hairpin | | Descriptor: | RNA (32-MER), Ribonuclease 3 | | Authors: | Wang, Z, Hartman, E, Roy, K, Chanfreau, G, Feigon, J. | | Deposit date: | 2011-04-06 | | Release date: | 2011-08-31 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structure of a Yeast RNase III dsRBD Complex with a Noncanonical RNA Substrate Provides New Insights into Binding Specificity of dsRBDs.
Structure, 19, 2011
|
|
1F4I
 
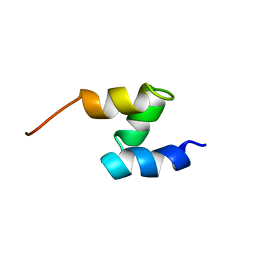 | | SOLUTION STRUCTURE OF THE HHR23A UBA(2) MUTANT P333E, DEFICIENT IN BINDING THE HIV-1 ACCESSORY PROTEIN VPR | | Descriptor: | UV EXCISION REPAIR PROTEIN PROTEIN RAD23 HOMOLOG A | | Authors: | Withers-Ward, E.S, Mueller, T.D, Chen, I.S, Feigon, J. | | Deposit date: | 2000-06-07 | | Release date: | 2000-12-20 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Biochemical and structural analysis of the interaction between the UBA(2) domain of the DNA repair protein HHR23A and HIV-1 Vpr.
Biochemistry, 39, 2000
|
|
1RKJ
 | | Solution structure of the complex formed by the two N-terminal RNA-binding domains of nucleolin and a pre-rRNA target | | Descriptor: | 5'-R(*GP*GP*AP*UP*GP*CP*CP*UP*CP*CP*CP*GP*AP*GP*UP*GP*CP*AP*UP*CP*C)-3', Nucleolin | | Authors: | Johansson, C, Finger, L.D, Trantirek, L, Mueller, T.D, Kim, S, Laird-Offringa, I.A, Feigon, J. | | Deposit date: | 2003-11-21 | | Release date: | 2004-04-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the complex formed by the two N-terminal RNA-binding domains of nucleolin and a pre-rRNA target.
J.Mol.Biol., 337, 2004
|
|
2K95
 
 | | Solution structure of the wild-type P2B-P3 pseudoknot of human telomerase RNA | | Descriptor: | Telomerase RNA P2b-P3 pseudoknot | | Authors: | Kim, N.-K, Zhang, Q, Zhou, J, Theimer, C.A, Peterson, R.D, Feigon, J. | | Deposit date: | 2008-09-29 | | Release date: | 2008-11-25 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure and Dynamics of the Wild-type Pseudoknot of Human Telomerase RNA.
J.Mol.Biol., 384, 2008
|
|
2LUQ
 
 | |
1K4X
 
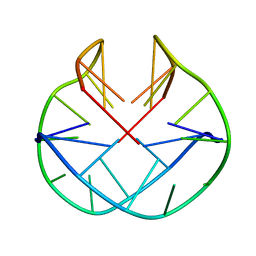 | | POTASSIUM FORM OF OXY-1.5 QUADRUPLEX DNA | | Descriptor: | DNA (5'-D(*GP*GP*GP*GP*TP*TP*TP*TP*GP*GP*GP*G)-3') | | Authors: | Schultze, P, Hud, N.V, Smith, F.W, Feigon, J. | | Deposit date: | 1999-06-08 | | Release date: | 1999-06-23 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | The effect of sodium, potassium and ammonium ions on the conformation of the dimeric quadruplex formed by the Oxytricha nova telomere repeat oligonucleotide d(G(4)T(4)G(4)).
Nucleic Acids Res., 27, 1999
|
|
1LWM
 
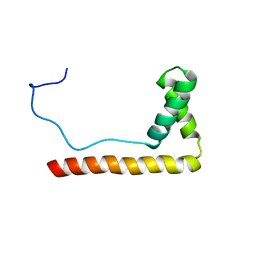 | | Solution Structure of the Sequence-Non-Specific HMGB protein NHP6A | | Descriptor: | NONHISTONE CHROMOSOMAL PROTEIN 6A | | Authors: | Masse, J.E, Wong, B, Yen, Y.-M, Allain, F.H.-T, Johnson, R.C, Feigon, J. | | Deposit date: | 2002-05-31 | | Release date: | 2002-10-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The S. cerevisiae architectural HMGB protein NHP6A complexed with DNA: DNA and protein conformational changes upon binding
J.Mol.Biol., 323, 2002
|
|
1J5N
 | | Solution Structure of the Non-Sequence-Specific HMGB protein NHP6A in complex with SRY DNA | | Descriptor: | 5'-D(*CP*TP*GP*AP*AP*CP*AP*AP*TP*CP*AP*CP*CP*CP*C)-3', 5'-D(*GP*GP*GP*GP*TP*GP*AP*TP*TP*GP*TP*TP*CP*AP*G)-3', Nonhistone chromosomal protein 6A | | Authors: | Masse, J.E, Wong, B, Yen, Y.-M, Allain, F.H.-T, Johnson, R.C, Feigon, J. | | Deposit date: | 2002-05-15 | | Release date: | 2002-10-16 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | The S. cerevisiae architectural HMGB protein NHP6A complexed with DNA: DNA and protein conformational changes upon binding
J.Mol.Biol., 323, 2002
|
|
1LWA
 
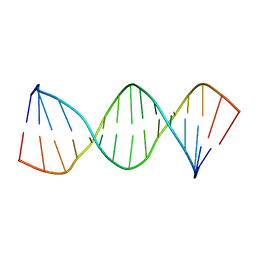 | | Solution Structure of SRY_DNA | | Descriptor: | 5'-D(*CP*TP*GP*AP*AP*CP*AP*AP*TP*CP*AP*CP*CP*CP*C)-3', 5'-D(*GP*GP*GP*GP*TP*GP*AP*TP*TP*GP*TP*TP*CP*AP*G)-3' | | Authors: | Masse, J.E, Wong, B, Yen, Y.-M, Allain, F.H.-T, Johnson, R.C, Feigon, J. | | Deposit date: | 2002-05-30 | | Release date: | 2002-10-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The S. cerevisiae architectural HMGB protein NHP6A complexed with DNA: DNA and protein conformational changes upon binding
J.Mol.Biol., 323, 2002
|
|
2LBW
 
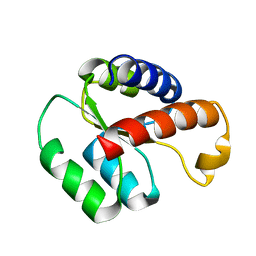 | | Solution structure of the S. cerevisiae H/ACA RNP protein Nhp2p-S82W mutant | | Descriptor: | H/ACA ribonucleoprotein complex subunit 2 | | Authors: | Koo, B, Park, C, Fernandez, C.F, Chim, N, Ding, Y, Chanfreau, G, Feigon, J. | | Deposit date: | 2011-04-07 | | Release date: | 2011-07-06 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structure of H/ACA RNP Protein Nhp2p Reveals Cis/Trans Isomerization of a Conserved Proline at the RNA and Nop10 Binding Interface.
J.Mol.Biol., 411, 2011
|
|
1P9D
 
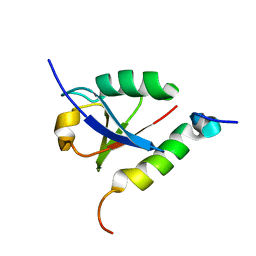 | |
1R3X
 
 | | INTRAMOLECULAR DNA TRIPLEX WITH RNA THIRD STRAND, NMR, 10 STRUCTURES | | Descriptor: | DNA (5'-D(*AP*GP*AP*GP*AP*GP*AP*A)-3'), DNA (5'-D(*TP*TP*CP*TP*CP*TP*CP*T)-3'), RNA (5'-R(*UP*CP*UP*CP*UP*CP*UP*U)-3') | | Authors: | Gotfredsen, C.H, Schultze, P, Feigon, J. | | Deposit date: | 1998-02-06 | | Release date: | 1998-05-20 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of an Intramolecular Pyrimidine-Purine-Pyrimidine Triplex Containing an RNA Third Strand
J.Am.Chem.Soc., 120, 1998
|
|
1P9C
 
 | |
2KYE
 
 | | Solution structure of the pseudouridine modified P6.1 hairpin of human telomerase RNA | | Descriptor: | RNA (5'-R(*GP*AP*GP*AP*GP*(PSU)P*(PSU)P*GP*GP*GP*CP*(PSU)P*CP*(PSU)P*C)-3') | | Authors: | Kim, N.-K, Theimer, C.A, Mitchell, J.R, Collins, K, Feigon, J. | | Deposit date: | 2010-05-25 | | Release date: | 2010-06-30 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Effect of pseudouridylation on the structure and activity of the catalytically essential P6.1 hairpin in human telomerase RNA.
Nucleic Acids Res., 38, 2010
|
|
1TLR
 
 | |
1RAW
 
 | |
1DV0
 | |
2LUP
 
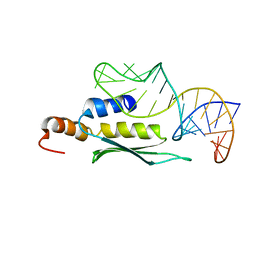 | |
1K4B
 
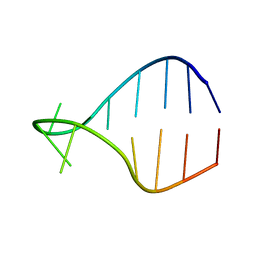 | | STRUCTURE OF AGUU RNA TETRALOOP, NMR, 20 STRUCTURES | | Descriptor: | 5'-R(*GP*GP*UP*UP*CP*AP*GP*UP*UP*GP*AP*AP*CP*C)-3' | | Authors: | Wu, H, Yang, P.K, Butcher, S.E, Kang, S, Chanfreau, G, Feigon, J. | | Deposit date: | 2001-10-07 | | Release date: | 2001-12-19 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | A novel family of RNA tetraloop structure forms the recognition site for Saccharomyces cerevisiae RNase III.
EMBO J., 20, 2001
|
|
1K4A
 
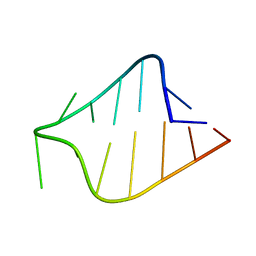 | | STRUCTURE OF AGAA RNA TETRALOOP, NMR, 20 STRUCTURES | | Descriptor: | 5'-R(*GP*GP*UP*UP*CP*AP*GP*AP*AP*GP*AP*AP*CP*C)-3' | | Authors: | Wu, H, Yang, P.K, Butcher, S.E, Kang, S, Chanfreau, G, Feigon, J. | | Deposit date: | 2001-10-07 | | Release date: | 2001-12-19 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | A novel family of RNA tetraloop structure forms the recognition site for Saccharomyces cerevisiae RNase III.
EMBO J., 20, 2001
|
|
1QWB
 
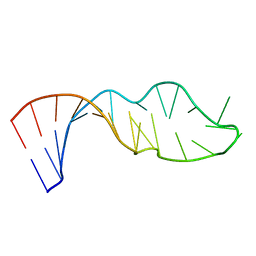 | |
1QWA
 
 | | NMR structure of 5'-r(GGAUGCCUCCCGAGUGCAUCC): an RNA hairpin derived from the mouse 5'ETS that binds nucleolin RBD12. | | Descriptor: | 18S ribosomal RNA, 5'ETS | | Authors: | Finger, L.D, Trantirek, L, Johansson, C, Feigon, J. | | Deposit date: | 2003-09-01 | | Release date: | 2003-11-25 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution Strucutres of Stem-loop RNAs that Bind to the Two N-terminal RNA Binding Domains of Nucleolin
Nucleic Acids Res., 31, 2003
|
|
1EBR
 
 | |
1EBS
 
 | |
1EBQ
 
 | |
