1MWB
 
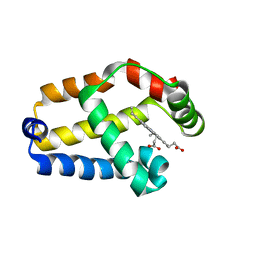 | | Solution structure of the recombinant hemoglobin from the cyanobacterium Synechocystis sp. PCC 6803 in its hemichrome state | | Descriptor: | Cyanoglobin, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Falzone, C.J, Vu, B.C, Scott, N.L, Lecomte, J.T. | | Deposit date: | 2002-09-27 | | Release date: | 2002-12-04 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The Solution Structure of the Recombinant Hemoglobin from the Cyanobacterium Synechocystis sp. PCC 6803 in its Hemichrome State
J.MOL.BIOL., 324, 2002
|
|
1I87
 
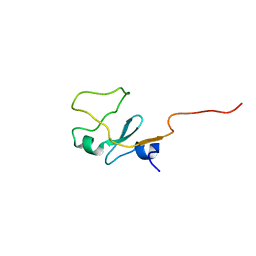 | | SOLUTION STRUCTURE OF THE WATER-SOLUBLE FRAGMENT OF RAT HEPATIC APOCYTOCHROME B5 | | Descriptor: | CYTOCHROME B5 | | Authors: | Falzone, C.J, Wang, Y, Vu, B.C, Scott, N.L, Bhattacharya, S, Lecomte, J.T. | | Deposit date: | 2001-03-12 | | Release date: | 2001-05-16 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structural and dynamic perturbations induced by heme binding in cytochrome b5.
Biochemistry, 40, 2001
|
|
1I8C
 
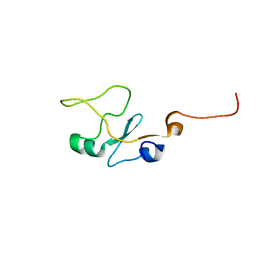 | | SOLUTION STRUCTURE OF THE WATER-SOLUBLE FRAGMENT OF RAT HEPATIC APOCYTOCHROME B5 | | Descriptor: | CYTOCHROME B5 | | Authors: | Falzone, C.J, Wang, Y, Vu, B.C, Scott, N.L, Bhattacharya, S, Lecomte, J.T. | | Deposit date: | 2001-03-13 | | Release date: | 2001-05-16 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structural and dynamic perturbations induced by heme binding in cytochrome b5.
Biochemistry, 40, 2001
|
|
1PSF
 
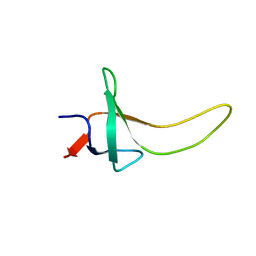 | | THE THREE-DIMENSIONAL SOLUTION STRUCTURE OF PSAE FROM THE CYANOBACTERIUM SYNECHOCOCCUS SP. STRAIN PCC 7002: A PHOTOSYSTEM I PROTEIN THAT SHOWS STRUCTURAL HOMOLOGY WITH SH3 DOMAINS | | Descriptor: | PHOTOSYSTEM I ACCESSORY PROTEIN E | | Authors: | Falzone, C.J, Kao, Y.-H, Zhao, J, Bryant, D.A, Lecomte, J.T.J. | | Deposit date: | 1994-04-13 | | Release date: | 1995-04-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional solution structure of PsaE from the cyanobacterium Synechococcus sp. strain PCC 7002, a photosystem I protein that shows structural homology with SH3 domains.
Biochemistry, 33, 1994
|
|
1PSE
 
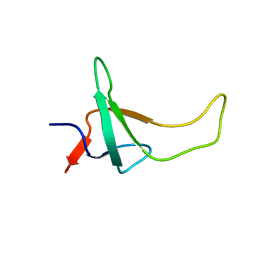 | | THE THREE-DIMENSIONAL SOLUTION STRUCTURE OF PSAE FROM THE CYANOBACTERIUM SYNECHOCOCCUS SP. STRAIN PCC 7002: A PHOTOSYSTEM I PROTEIN THAT SHOWS STRUCTURAL HOMOLOGY WITH SH3 DOMAINS | | Descriptor: | PHOTOSYSTEM I ACCESSORY PROTEIN E | | Authors: | Falzone, C.J, Kao, Y.-H, Zhao, J, Bryant, D.A, Lecomte, J.T.J. | | Deposit date: | 1994-04-13 | | Release date: | 1995-04-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional solution structure of PsaE from the cyanobacterium Synechococcus sp. strain PCC 7002, a photosystem I protein that shows structural homology with SH3 domains.
Biochemistry, 33, 1994
|
|
1IET
 
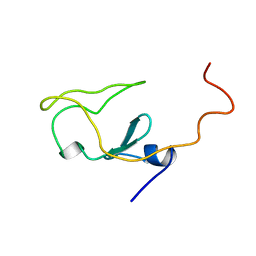 | |
1IEU
 
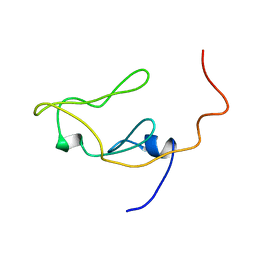 | |
2KSC
 
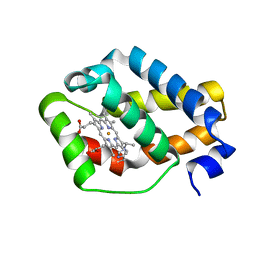 | |
1QP2
 
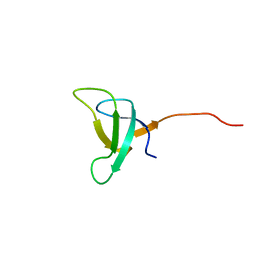 | | SOLUTION STRUCTURE OF PHOTOSYSTEM I ACCESSORY PROTEIN E FROM THE CYANOBACTERIUM NOSTOC SP. STRAIN PCC 8009 | | Descriptor: | PROTEIN (PSAE PROTEIN) | | Authors: | Mayer, K.L, Shen, G, Bryant, D.A, Lecomte, J.T.J, Falzone, C.J. | | Deposit date: | 1999-05-29 | | Release date: | 1999-10-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The solution structure of photosystem I accessory protein E from the cyanobacterium Nostoc sp. strain PCC 8009.
Biochemistry, 38, 1999
|
|
1QP3
 
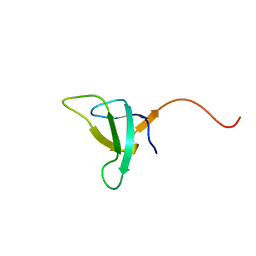 | | SOLUTION STRUCTURE OF PHOTOSYSTEM I ACCESSORY PROTEIN E FROM THE CYANOBACTERIUM NOSTOC SP. STRAIN PCC 8009 | | Descriptor: | PROTEIN (PSAE PROTEIN) | | Authors: | Mayer, K.L, Shen, G, Bryant, D.A, Lecomte, J.T.J, Falzone, C.J. | | Deposit date: | 1999-05-29 | | Release date: | 1999-10-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The solution structure of photosystem I accessory protein E from the cyanobacterium Nostoc sp. strain PCC 8009.
Biochemistry, 38, 1999
|
|
2WGI
 
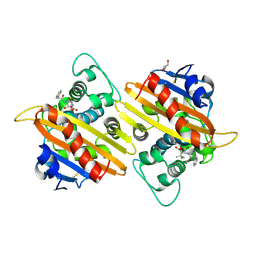 | | Crystal structure of the acyl-enzyme OXA-10 W154A-benzylpenicillin at pH 6 | | Descriptor: | BETA-LACTAMASE OXA-10, GLYCEROL, OPEN FORM - PENICILLIN G | | Authors: | Vercheval, L, Falzone, C, Sauvage, E, Herman, R, Charlier, P, Galleni, M, Kerff, F. | | Deposit date: | 2009-04-20 | | Release date: | 2009-11-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Critical Role of Tryptophan 154 for the Activity and Stability of Class D Beta-Lactamases.
Biochemistry, 48, 2009
|
|
2HP5
 
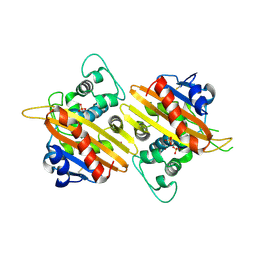 | | Crystal Structure of the OXA-10 W154G mutant at pH 7.0 | | Descriptor: | Beta-lactamase PSE-2, COBALT (II) ION, SULFATE ION | | Authors: | Kerff, F, Falzone, C, Herman, R, Sauvage, E, Charlier, P. | | Deposit date: | 2006-07-17 | | Release date: | 2007-07-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Biochemistry, 48, 2009
|
|
2HPB
 
 | | Crystal structure of the OXA-10 W154A mutant at pH 9.0 | | Descriptor: | Beta-lactamase PSE-2, SULFATE ION | | Authors: | Kerff, F, Falzone, C, Herman, R, Sauvage, E, Charlier, P. | | Deposit date: | 2006-07-17 | | Release date: | 2007-07-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Biochemistry, 48, 2009
|
|
2HP6
 
 | | Crystal structure of the OXA-10 W154A mutant at pH 7.5 | | Descriptor: | Beta-lactamase PSE-2, SULFATE ION | | Authors: | Kerff, F, Falzone, C, Herman, R, Sauvage, E, Charlier, P. | | Deposit date: | 2006-07-17 | | Release date: | 2007-07-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Biochemistry, 48, 2009
|
|
2HP9
 
 | | Crystal Structure of the OXA-10 W154A mutant at pH 6.0 | | Descriptor: | Beta-lactamase PSE-2, SULFATE ION | | Authors: | Kerff, F, Falzone, C, Herman, R, Sauvage, E, Charlier, P. | | Deposit date: | 2006-07-17 | | Release date: | 2007-07-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Biochemistry, 48, 2009
|
|
