1TOZ
 
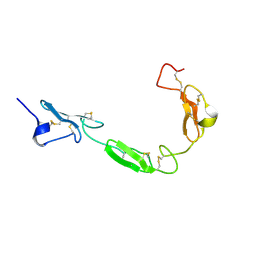 | | NMR structure of the human NOTCH-1 ligand binding region | | Descriptor: | Neurogenic locus notch homolog protein 1 | | Authors: | Hambleton, S, Valeyev, N.Y, Muranyi, A, Knott, V, Werner, J.M, Mcmichael, A.J, Handford, P.A, Downing, A.K. | | Deposit date: | 2004-06-15 | | Release date: | 2004-10-12 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Structural and functional properties of the human notch-1 ligand binding region
STRUCTURE, 12, 2004
|
|
2Z48
 
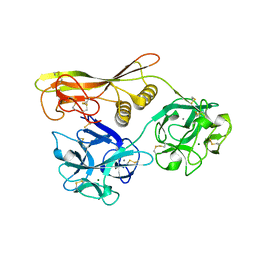 | | Crystal Structure of Hemolytic Lectin CEL-III Complexed with GalNac | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-galactopyranose, 2-acetamido-2-deoxy-beta-D-galactopyranose, CALCIUM ION, ... | | Authors: | Hatakeyama, T, Unno, H, Eto, S, Hidemura, H, Uchida, T, Kouzuma, Y. | | Deposit date: | 2007-06-13 | | Release date: | 2007-10-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | C-type lectin-like carbohydrate-recognition of the hemolytic lectin CEL-III containing ricin-type beta-trefoil folds
J.Biol.Chem., 282, 2007
|
|
2Z49
 
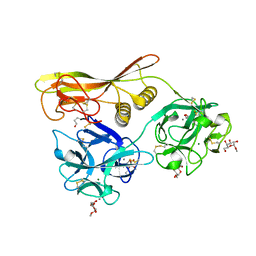 | | Crystal Structure of Hemolytic Lectin CEL-III Complexed with methyl-alpha-D-galactopylanoside | | Descriptor: | CALCIUM ION, Hemolytic lectin CEL-III, MAGNESIUM ION, ... | | Authors: | Hatakeyama, T, Unno, H, Eto, S, Hidemura, H, Uchida, T, Kouzuma, Y. | | Deposit date: | 2007-06-13 | | Release date: | 2007-10-30 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | C-type lectin-like carbohydrate-recognition of the hemolytic lectin CEL-III containing ricin-type beta-trefoil folds
J.Biol.Chem., 282, 2007
|
|
1VCL
 
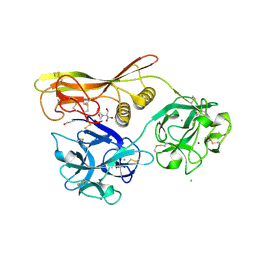 | | Crystal Structure of Hemolytic Lectin CEL-III | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Uchida, T, Yamasaki, T, Eto, S, Sugawara, H, Kurisu, G, Nakagawa, A, Kusunoki, M, Hatakeyama, T. | | Deposit date: | 2004-03-09 | | Release date: | 2004-09-07 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structure of the Hemolytic Lectin CEL-III Isolated from the Marine Invertebrate Cucumaria echinata: IMPLICATIONS OF DOMAIN STRUCTURE FOR ITS MEMBRANE PORE-FORMATION MECHANISM
J.Biol.Chem., 279, 2004
|
|
3KFY
 
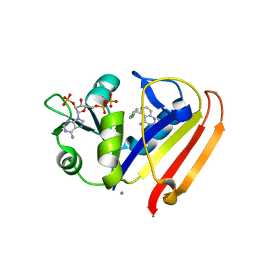 | | Dynamic switching and partial occupancies of a small molecule inhibitor complex of DHFR | | Descriptor: | 5-[(4-chlorophenyl)sulfanyl]quinazoline-2,4-diamine, CALCIUM ION, Dihydrofolate reductase, ... | | Authors: | Collins, E.J, Lee, A.L, Carroll, M.J, Gromova, A.V, Miller, K.R, Singleton, S.F. | | Deposit date: | 2009-10-28 | | Release date: | 2010-12-08 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Dynamic switching and partial occupancies of a small molecule inhibitor complex of DHFR
To be Published
|
|
3R33
 
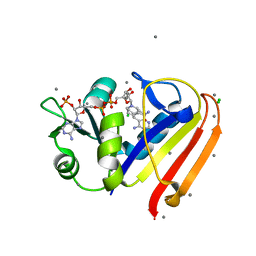 | | Evidence for dynamic motion in proteins as a mechanism for ligand dissociation | | Descriptor: | (6S)-6-methyl-5,6,7,8-tetrahydroquinazoline-2,4-diamine, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Collins, E.J, Lee, A.L, Carroll, M.J, Mauldin, R.V, Gromova, A.V, Singleton, S.F. | | Deposit date: | 2011-03-15 | | Release date: | 2012-01-25 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Evidence for dynamics in proteins as a mechanism for ligand dissociation.
Nat.Chem.Biol., 8, 2012
|
|
3QYL
 | | Sensitivity of receptor internal motions to ligand binding affinity and kinetic off-rate | | Descriptor: | (7S)-7-methyl-5,6,7,8-tetrahydroquinazoline-2,4-diamine, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Collins, E.J, Lee, A.L, Carroll, M.J, Mauldin, R.V, Gromova, A.V, Singleton, S.F. | | Deposit date: | 2011-03-03 | | Release date: | 2012-01-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Evidence for dynamics in proteins as a mechanism for ligand dissociation.
Nat.Chem.Biol., 8, 2012
|
|
3QYO
 | | Sensitivity of receptor internal motions to ligand binding affinity and kinetic off-rate | | Descriptor: | CALCIUM ION, Dihydrofolate reductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Collins, E.J, Lee, A.L, Carroll, M.J, Mauldin, R.V, Gromova, A.V, Singleton, S.F. | | Deposit date: | 2011-03-03 | | Release date: | 2012-01-18 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Evidence for dynamics in proteins as a mechanism for ligand dissociation.
Nat.Chem.Biol., 8, 2012
|
|
2CN4
 
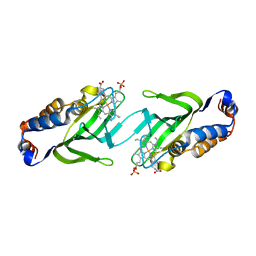 | | The crystal structure of the secreted dimeric form of the hemophore HasA reveals a domain swapping with an exchanged heme ligand | | Descriptor: | HEMOPHORE HASA, PHOSPHATE ION, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Czjzek, M, Letoffe, S, Wandersman, C, Delepierre, M, Lecroisey, A, Izadi-Pruneyre, N. | | Deposit date: | 2006-05-18 | | Release date: | 2006-05-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Crystal Structure of the Secreted Dimeric Form of the Hemophore Hasa Reveals a Domain Swapping with an Exchanged Heme Ligand
J.Mol.Biol., 365, 2007
|
|
1SMP
 
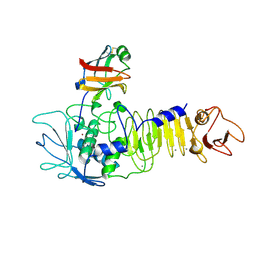 | | CRYSTAL STRUCTURE OF A COMPLEX BETWEEN SERRATIA MARCESCENS METALLO-PROTEASE AND AN INHIBITOR FROM ERWINIA CHRYSANTHEMI | | Descriptor: | CALCIUM ION, ERWINIA CHRYSANTHEMI INHIBITOR, SERRATIA METALLO PROTEINASE, ... | | Authors: | Baumann, U, Bauer, M, Letoffe, S, Delepelaire, P, Wandersman, C. | | Deposit date: | 1995-01-13 | | Release date: | 1996-04-03 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of a complex between Serratia marcescens metallo-protease and an inhibitor from Erwinia chrysanthemi.
J.Mol.Biol., 248, 1995
|
|
