3CV7
 
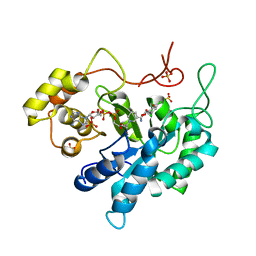 | | Crystal structure of porcine aldehyde reductase ternary complex | | Descriptor: | 3,5-dichloro-2-hydroxybenzoic acid, Alcohol dehydrogenase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Carbone, V, El-Kabbani, O. | | Deposit date: | 2008-04-18 | | Release date: | 2008-10-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.412 Å) | | Cite: | Structure of aldehyde reductase in ternary complex with coenzyme and the potent 20alpha-hydroxysteroid dehydrogenase inhibitor 3,5-dichlorosalicylic acid: Implications for inhibitor binding and selectivity
Arch.Biochem.Biophys., 479, 2008
|
|
3D3W
 
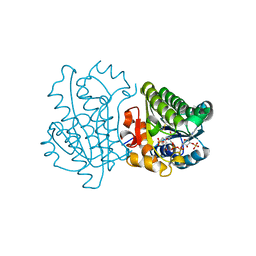 | |
3C3U
 
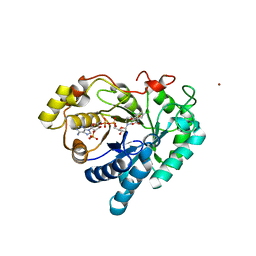 | | Crystal structure of AKR1C1 in complex with NADP and 3,5-dichlorosalicylic acid | | Descriptor: | 3,5-dichloro-2-hydroxybenzoic acid, Aldo-keto reductase family 1 member C1, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Dhagat, U, El-Kabbani, O. | | Deposit date: | 2008-01-28 | | Release date: | 2008-08-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Selectivity determinants of inhibitor binding to human 20alpha-hydroxysteroid dehydrogenase: crystal structure of the enzyme in ternary complex with coenzyme and the potent inhibitor 3,5-dichlorosalicylic acid
J.Med.Chem., 51, 2008
|
|
3CV6
 
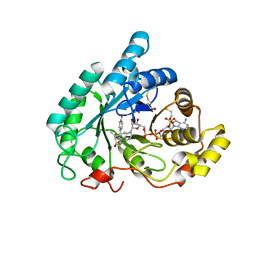 | | The crystal structure of mouse 17-alpha hydroxysteroid dehydrogenase GG225.226PP mutant in complex with inhibitor and cofactor NADP+. | | Descriptor: | 4-[(1R,2S)-1-ethyl-2-(4-hydroxyphenyl)butyl]phenol, Aldo-keto reductase family 1 member C21, BETA-MERCAPTOETHANOL, ... | | Authors: | Dhagat, U, El-Kabbani, O. | | Deposit date: | 2008-04-17 | | Release date: | 2009-03-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of the G225P/G226P mutant of mouse 3(17)alpha-hydroxysteroid dehydrogenase (AKR1C21) ternary complex: implications for the binding of inhibitor and substrate.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
3BCJ
 
 | | Crystal structure of Aldose Reductase complexed with 2S4R (Stereoisomer of Fidarestat, 2S4S) at 0.78 A | | Descriptor: | (2S,4R)-2-AMINOFORMYL-6-FLUORO-SPIRO[CHROMAN-4,4'-IMIDAZOLIDINE]-2',5'-DIONE, Aldose reductase, CITRIC ACID, ... | | Authors: | Zhao, H.T, El-Kabbani, O. | | Deposit date: | 2007-11-13 | | Release date: | 2008-04-08 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (0.78 Å) | | Cite: | Unusual Binding Mode of the 2S4R Stereoisomer of the Potent Aldose Reductase Cyclic Imide Inhibitor Fidarestat (2S4S) in the 15 K Crystal Structure of the Ternary Complex Refined at 0.78 A Resolution: Implications for the Inhibition Mechanism
J.Med.Chem., 51, 2008
|
|
