4UD8
 
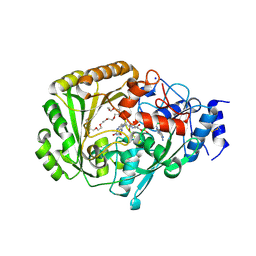 | | AtBBE15 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3,6,9,12,15,18,21,24-OCTAOXAHEXACOSAN-1-OL, ... | | Authors: | Daniel, B, Steiner, B, Pavkov-Keller, T, Dordic, A, Gutmann, A, Sensen, C.W, Nidetzky, B, van der Graaff, E, Wallner, S, Gruber, K, Macheroux, P. | | Deposit date: | 2014-12-09 | | Release date: | 2015-06-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.088 Å) | | Cite: | Oxidation of Monolignols by Members of the Berberine Bridge Enzyme Family Suggests a Role in Cell Wall Metabolism.
J.Biol.Chem., 290, 2015
|
|
5D79
 
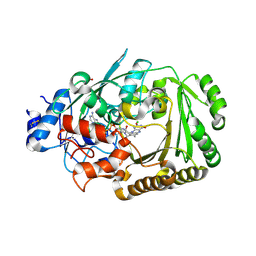 | | Structure of BBE-like #28 from Arabidopsis thaliana | | Descriptor: | Berberine bridge enzyme-like protein, CHLORIDE ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Daniel, B, Kumar, P, Gruber, K. | | Deposit date: | 2015-08-13 | | Release date: | 2016-06-22 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.849 Å) | | Cite: | Structure of a Berberine Bridge Enzyme-Like Enzyme with an Active Site Specific to the Plant Family Brassicaceae.
Plos One, 11, 2016
|
|
8AEP
 
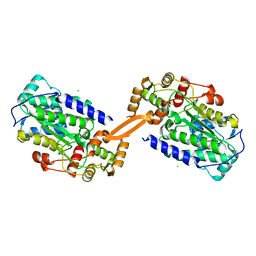 | | Reductase domain of the carboxylate reductase of Neurospora crassa | | Descriptor: | Acetyl-CoA synthetase-like protein, CHLORIDE ION, SULFATE ION | | Authors: | Daniel, B, Schrufer, A, Marlene, L, Sagmeister, T, Pavkov-Keller, T. | | Deposit date: | 2022-07-13 | | Release date: | 2023-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of the Reductase Domain of a Fungal Carboxylic Acid Reductase and Its Substrate Scope in Thioester and Aldehyde Reduction.
Acs Catalysis, 12, 2022
|
|
2ABM
 
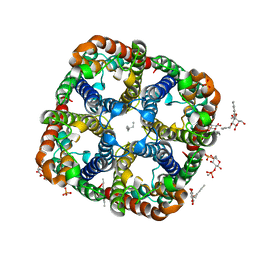 | | Crystal Structure of Aquaporin Z Tetramer Reveals both Open and Closed Water-conducting Channels | | Descriptor: | (1S)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PENTANOYLOXY)METHYL]ETHYL OCTANOATE, 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, 2-O-octyl-beta-D-glucopyranose, ... | | Authors: | Jiang, J, Daniels, B.V, Fu, D. | | Deposit date: | 2005-07-15 | | Release date: | 2005-09-20 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Crystal Structure of AqpZ Tetramer Reveals Two Distinct Arg-189 Conformations Associated with Water Permeation through the Narrowest Constriction of the Water-conducting Channel.
J.Biol.Chem., 281, 2006
|
|
8OIM
 
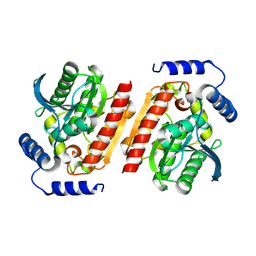 | |
1NKQ
 
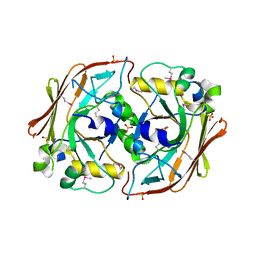 | | Crystal structure of yeast ynq8, a fumarylacetoacetate hydrolase family protein | | Descriptor: | ACETIC ACID, CALCIUM ION, Hypothetical 28.8 kDa protein in PSD1-SKO1 intergenic region, ... | | Authors: | Eswaramoorthy, S, Kumaran, D, Daniels, B, Studier, F.W, Swaminathan, S, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2003-01-03 | | Release date: | 2004-06-15 | | Last modified: | 2021-02-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crtystal Structure of Yeast Hypothetical Protein YNQ8_YEAST
To be Published
|
|
8A85
 
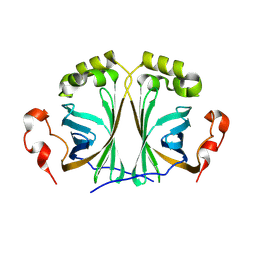 | |
8ADX
 
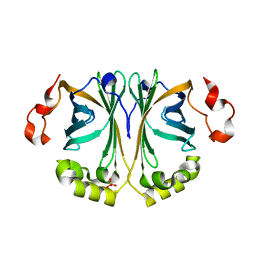 | |
8B30
 | |
8C66
 
 | |
