6MFL
 
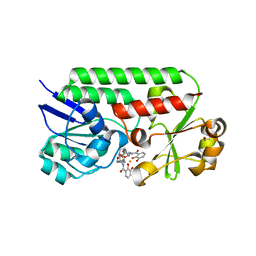 | | Structure of siderophore binding protein BauB bound to a complex between two molecules of acinetobactin and ferric iron. | | Descriptor: | 1,2-ETHANEDIOL, FE (III) ION, Siderophore Binding Protein BauB, ... | | Authors: | Gulick, A.M, Bailey, D.C, Wencewicz, T.A. | | Deposit date: | 2018-09-11 | | Release date: | 2018-11-21 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of the Siderophore Binding Protein BauB Bound to an Unusual 2:1 Complex Between Acinetobactin and Ferric Iron.
Biochemistry, 57, 2018
|
|
6MCW
 
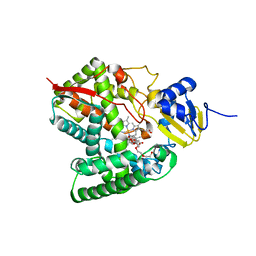 | | Crystal structure of the P450 domain of the CYP51-ferredoxin fusion protein from Methylococcus capsulatus, complex with the detergent Anapoe-X-114 | | Descriptor: | 23-[4-(2,4,4-trimethylpentan-2-yl)phenoxy]-3,6,9,12,15,18,21-heptaoxatricosan-1-ol, Cytochrome P450 51, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Hargrove, T, Wawrzak, Z, Lamb, D.C, Lepesheva, G.I. | | Deposit date: | 2018-09-02 | | Release date: | 2019-07-10 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Concerning P450 evolution: Structural Analyses Support Bacterial Origin of Sterol 14 alpha-Demethylases.
Mol.Biol.Evol., 2020
|
|
8TC3
 
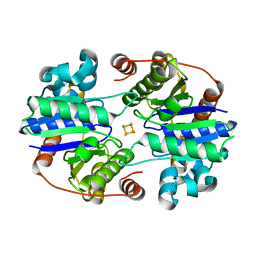 | |
8UMC
 
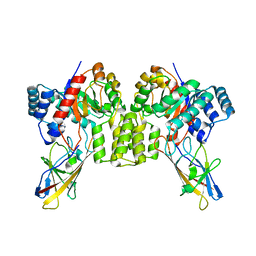 | | Deinococcus aerius TR0125 C-glucosyl deglycosidase (CGD), cryo-EM | | Descriptor: | DUF6379 domain-containing protein, MANGANESE (II) ION, Xylose isomerase-like TIM barrel domain-containing protein | | Authors: | Furlanetto, V, Kalyani, D.C, Kostelac, A, Haltrich, D, Hallberg, B.M, Divne, C. | | Deposit date: | 2023-10-17 | | Release date: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Structural and Functional Characterization of a Gene Cluster Responsible for Deglycosylation of C-glucosyl Flavonoids and Xanthonoids by Deinococcus aerius.
J.Mol.Biol., 436, 2024
|
|
8TQP
 | | HIV-CA Disulfide linked Hexamer bound to Quinazolin-4-one Scaffold inhibitor | | Descriptor: | 2-[4-(4-aminobenzene-1-sulfonyl)-2-oxopiperazin-1-yl]-N-{(1R)-2-(3,5-difluorophenyl)-1-[3-(4-methoxyphenyl)-4-oxo-3,4-dihydroquinazolin-2-yl]ethyl}acetamide, Gag polyprotein | | Authors: | Goldstone, D.C, Barnett, M.J, Taka, J.R.H. | | Deposit date: | 2023-08-08 | | Release date: | 2023-12-20 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Discovery, Crystallographic Studies, and Mechanistic Investigations of Novel Phenylalanine Derivatives Bearing a Quinazolin-4-one Scaffold as Potent HIV Capsid Modulators.
J.Med.Chem., 66, 2023
|
|
8TOV
 
 | | HIV-CA Disulfide linked Hexamer bound to Quinazolin-4-one Scaffold inhibitor | | Descriptor: | 2-[4-(4-aminobenzene-1-sulfonyl)-2-oxopiperazin-1-yl]-N-[(1R)-2-(3,5-difluorophenyl)-1-{3-[4-(morpholine-4-sulfonyl)phenyl]-4-oxo-3,4-dihydroquinazolin-2-yl}ethyl]acetamide, Matrix protein p17 | | Authors: | Goldstone, D.C, Barnett, M.J, Taka, J.R.H. | | Deposit date: | 2023-08-04 | | Release date: | 2023-12-20 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Discovery, Crystallographic Studies, and Mechanistic Investigations of Novel Phenylalanine Derivatives Bearing a Quinazolin-4-one Scaffold as Potent HIV Capsid Modulators.
J.Med.Chem., 66, 2023
|
|
8OZD
 | | cryoEM structure of SPARTA complex dimer-3 | | Descriptor: | DNA (16-MER), Piwi domain-containing protein, RNA (18-MER), ... | | Authors: | Ekundayo, B, Ni, D.C, Lu, X.H, Stahlberg, H. | | Deposit date: | 2023-05-09 | | Release date: | 2023-08-16 | | Last modified: | 2023-08-23 | | Method: | ELECTRON MICROSCOPY (3.89 Å) | | Cite: | Activation mechanism of a short argonaute-TIR prokaryotic immune system.
Sci Adv, 9, 2023
|
|
9CEQ
 
 | |
9BHW
 | |
9BHT
 | | Septin Hexameric Complex SEPT2/SEPT6/SEPT7 of Ciona intestinalis by Cryo-EM | | Descriptor: | CiSeptin-2, CiSeptin-6, CiSeptin-7, ... | | Authors: | Mendonca, D.C, Pereira, H.M, Garratt, R.C. | | Deposit date: | 2024-04-22 | | Release date: | 2024-07-10 | | Last modified: | 2024-07-24 | | Method: | ELECTRON MICROSCOPY (3.26 Å) | | Cite: | Structural Insights into Ciona intestinalis Septins: Complexes Suggest a Mechanism for Nucleotide-dependent Interfacial Cross-talk.
J.Mol.Biol., 436, 2024
|
|
8OZC
 
 | | cryoEM structure of SPARTA complex heterodimer apo | | Descriptor: | Piwi domain-containing protein, TIR domain-containing protein | | Authors: | Ekundayo, B, Ni, D.C, Lu, X.H, Stahlberg, H. | | Deposit date: | 2023-05-09 | | Release date: | 2023-08-16 | | Last modified: | 2023-08-23 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Activation mechanism of a short argonaute-TIR prokaryotic immune system.
Sci Adv, 9, 2023
|
|
4YL6
 
 | |
4YFN
 
 | | Escherichia coli RNA polymerase in complex with squaramide compound 14 (N-[3,4-dioxo-2-(4-{[4-(trifluoromethyl)benzyl]amino}piperidin-1-yl)cyclobut-1-en-1-yl]-3,5-dimethyl-1,2-oxazole-4-sulfonamide) | | Descriptor: | DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, DNA-directed RNA polymerase subunit beta', ... | | Authors: | Molodtsov, V, Fleming, P.R, Eyermann, C.J, Ferguson, A.D, Foulk, M.A, McKinney, D.C, Masse, C.E, Buurman, E.T, Murakami, K.S. | | Deposit date: | 2015-02-25 | | Release date: | 2015-03-11 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.817 Å) | | Cite: | X-ray Crystal Structures of Escherichia coli RNA Polymerase with Switch Region Binding Inhibitors Enable Rational Design of Squaramides with an Improved Fraction Unbound to Human Plasma Protein.
J.Med.Chem., 58, 2015
|
|
4Z7M
 
 | | Novel Inhibitors of Bacterial Methionine Aminopeptidase with Broad-Spectrum Biochemical Activity | | Descriptor: | MANGANESE (II) ION, Methionine aminopeptidase, N~2~-[(3,5-difluorophenyl)acetyl]-N-[(3S,7R)-1-methyl-2-oxo-7-phenyl-2,3,4,7-tetrahydro-1H-azepin-3-yl]-L-alaninamide | | Authors: | Rose, J.A, Lahiri, S.D, McKinney, D.C, Albert, R, Morningstar, M, Shapiro, A.B, Fisher, S.F, Fleming, P.R. | | Deposit date: | 2015-04-07 | | Release date: | 2016-04-13 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Novel Inhibitors of Bacterial Methionine Aminopeptidase with Broad-Spectrum Biochemical Activity
To be Published
|
|
4ZTC
 
 | | PglE Aminotransferase in complex with External Aldimine, Mutant K184A | | Descriptor: | Aminotransferase homolog, [(2R,3R,4R,5S,6R)-3-acetamido-6-methyl-5-[(E)-[2-methyl-3-oxidanyl-5-(phosphonooxymethyl)pyridin-4-yl]methylideneamino]-4-oxidanyl-oxan-2-yl] [[(2R,3S,4R,5R)-5-[2,4-bis(oxidanylidene)pyrimidin-1-yl]-3,4-bis(oxidanyl)oxolan-2-yl]methoxy-oxidanyl-phosphoryl] hydrogen phosphate | | Authors: | Riegert, A.S, Thoden, J.B, Young, N.M, Watson, D.C, Holden, H.M. | | Deposit date: | 2015-05-14 | | Release date: | 2015-07-29 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the external aldimine form of PglE, an aminotransferase required for N,N'-diacetylbacillosamine biosynthesis.
Protein Sci., 24, 2015
|
|
6QOZ
 
 | | CryoEM reconstruction of Cowpea Mosaic Virus (CPMV) bound to an Affimer reagent | | Descriptor: | Affimer binding protein, Cowpea mosaic virus large subunit, RNA2 polyprotein | | Authors: | Hesketh, E.L, Tiede, C, Adamson, H, Adams, T.L, Byrne, M.J, Meshcheriakova, Y, Lomonossoff, G.P, Kruse, I, McPherson, M.J, Tomlinson, D.C, Ranson, N.A. | | Deposit date: | 2019-02-13 | | Release date: | 2019-12-18 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Affimer reagents as tools in diagnosing plant virus diseases.
Sci Rep, 9, 2019
|
|
4YKC
 
 | |
4YKD
 
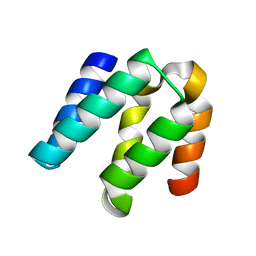 | |
4YFX
 
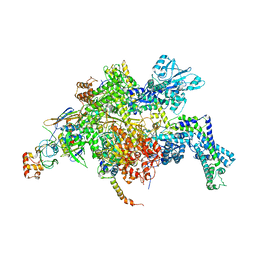 | | Escherichia coli RNA polymerase in complex with Myxopyronin B | | Descriptor: | DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, DNA-directed RNA polymerase subunit beta', ... | | Authors: | Molodtsov, V, Fleming, P.R, Eyermann, C.J, Ferguson, A.D, Foulk, M.A, McKinney, D.C, Masse, C.E, Buurman, E.T, Murakami, K.S. | | Deposit date: | 2015-02-25 | | Release date: | 2015-03-11 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.844 Å) | | Cite: | X-ray Crystal Structures of Escherichia coli RNA Polymerase with Switch Region Binding Inhibitors Enable Rational Design of Squaramides with an Improved Fraction Unbound to Human Plasma Protein.
J.Med.Chem., 58, 2015
|
|
4YFK
 
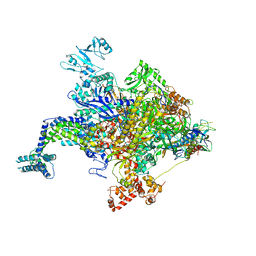 | | Escherichia coli RNA polymerase in complex with squaramide compound 8. | | Descriptor: | 3,5-dimethyl-N-{2-[4-(4-methylbenzyl)piperidin-1-yl]-3,4-dioxocyclobut-1-en-1-yl}-1,2-oxazole-4-sulfonamide, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | Authors: | Molodtsov, V, Fleming, P.R, Eyermann, C.J, Ferguson, A.D, Foulk, M.A, McKinney, D.C, Masse, C.E, Buurman, E.T, Murakami, K.S. | | Deposit date: | 2015-02-25 | | Release date: | 2015-03-11 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.571 Å) | | Cite: | X-ray Crystal Structures of Escherichia coli RNA Polymerase with Switch Region Binding Inhibitors Enable Rational Design of Squaramides with an Improved Fraction Unbound to Human Plasma Protein.
J.Med.Chem., 58, 2015
|
|
5A0O
 
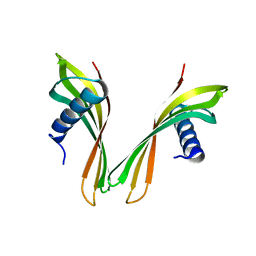 | | adhiron raised against p300 | | Descriptor: | ADHIRON | | Authors: | Kyle, H.F, Wickson, K.F, Stott, J, Burslem, G.M, Breeze, A.L, Tiede, C, Tomlinson, D.C, Warriner, S.L, Nelson, A, Wilson, A.J, Edwards, T.A. | | Deposit date: | 2015-04-21 | | Release date: | 2015-07-15 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.73 Å) | | Cite: | Exploration of the Hif-1Alpha.P300 Interface Using Peptide and Adhiron Phage Display Technologies
Mol.Biosyst., 11, 2015
|
|
4ZRZ
 
 | | PlyCB mutant R66E | | Descriptor: | PlyCB | | Authors: | Gallagher, D.T, Nelson, D.C, Shen, Y. | | Deposit date: | 2015-05-12 | | Release date: | 2016-09-14 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | A bacteriophage endolysin that eliminates intracellular streptococci.
Elife, 5, 2016
|
|
4YAH
 
 | | Crystal Structure of the Methionine Binding Protein, MetQ | | Descriptor: | D-methionine-binding lipoprotein MetQ, METHIONINE | | Authors: | Lai, J.Y, Kadaba, N.S, Kaiser, J.T, Rees, D.C. | | Deposit date: | 2015-02-17 | | Release date: | 2015-04-08 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The contribution of methionine to the stability of the Escherichia coli MetNIQ ABC transporter-substrate binding protein complex.
Biol.Chem., 396, 2015
|
|
5BNM
 
 | | E. coli FabH with Small Molecule Inhibitor 1 | | Descriptor: | 3-oxoacyl-[acyl-carrier-protein] synthase 3, N-{[3'-(hydroxymethyl)biphenyl-4-yl]methyl}benzenesulfonamide, SULFATE ION, ... | | Authors: | Kazmirski, S.L, McKinney, D.C. | | Deposit date: | 2015-05-26 | | Release date: | 2016-05-18 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Antibacterial FabH Inhibitors with Mode of Action Validated in Haemophilus influenzae by in Vitro Resistance Mutation Mapping.
Acs Infect Dis., 2, 2016
|
|
5AO5
 
 | | Endo180 D1-4, monoclinic form | | Descriptor: | C-TYPE MANNOSE RECEPTOR 2, SODIUM ION, SULFATE ION | | Authors: | Paracuellos, P, Briggs, D.C, Carafoli, F, Loncar, T, Hohenester, E. | | Deposit date: | 2015-09-09 | | Release date: | 2015-10-28 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Insights Into Collagen Uptake by C-Type Mannose Receptors from the Crystal Structure of Endo180 Domains 1-4.
Structure, 23, 2015
|
|
