1AK0
 
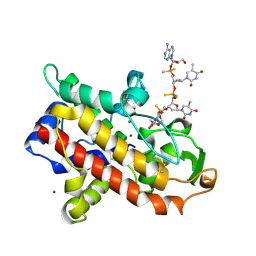 | | P1 NUCLEASE IN COMPLEX WITH A SUBSTRATE ANALOG | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ADENOSINE-5'-(DITHIO)PHOSPHATE, ... | | 著者 | Romier, C, Suck, D. | | 登録日 | 1997-05-28 | | 公開日 | 1997-12-03 | | 最終更新日 | 2020-07-29 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Recognition of single-stranded DNA by nuclease P1: high resolution crystal structures of complexes with substrate analogs.
Proteins, 32, 1998
|
|
8QV0
 
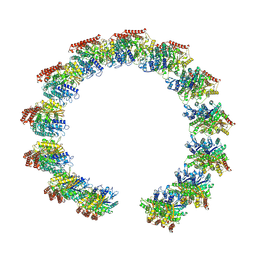 | | Structure of the native microtubule lattice nucleated from the yeast spindle pole body | | 分子名称: | Tubulin alpha-1 chain, Tubulin beta chain | | 著者 | Dendooven, T, Yatskevich, S, Burt, A, Bellini, D, Kilmartin, J, Barford, D. | | 登録日 | 2023-10-17 | | 公開日 | 2024-04-24 | | 実験手法 | ELECTRON MICROSCOPY (6.6 Å) | | 主引用文献 | Structure of the native gamma-tubulin ring complex capping spindle microtubules.
Nat.Struct.Mol.Biol., 2024
|
|
1R7D
 
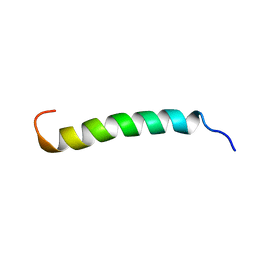 | | NMR structure of the membrane anchor domain (1-31) of the nonstructural protein 5A (NS5A) of hepatitis C virus (Ensemble of 51 structures, sample in 50% tfe) | | 分子名称: | Genome polyprotein | | 著者 | Penin, F, Brass, V, Appel, N, Ramboarina, S, Montserret, R, Ficheux, D, Blum, H.E, Bartenschlager, R, Moradpour, D. | | 登録日 | 2003-10-21 | | 公開日 | 2004-08-10 | | 最終更新日 | 2024-05-22 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Structure and function of the membrane anchor domain of hepatitis C virus nonstructural protein 5A.
J.Biol.Chem., 279, 2004
|
|
1YQ3
 
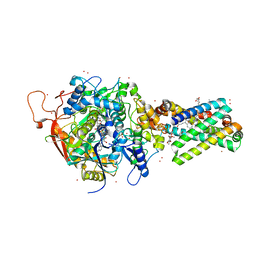 | | Avian respiratory complex ii with oxaloacetate and ubiquinone | | 分子名称: | 1,2-dioleoyl-sn-glycero-3-phosphoethanolamine, Coenzyme Q10, (2Z,6E,10Z,14E,18E,22E,26Z)-isomer, ... | | 著者 | Huang, L, Cobessi, D, Tung, E.Y, Berry, E.A. | | 登録日 | 2005-02-01 | | 公開日 | 2005-12-20 | | 最終更新日 | 2023-08-23 | | 実験手法 | X-RAY DIFFRACTION (2.2 Å) | | 主引用文献 | 3-Nitropropionic Acid Is a Suicide Inhibitor of Mitochondrial Respiration That, upon Oxidation by Complex II, Forms a Covalent Adduct with a Catalytic Base Arginine in the Active Site of the Enzyme
J.Biol.Chem., 281, 2006
|
|
1A5Y
 
 | |
6XF8
 
 | | DLP 5 fold | | 分子名称: | Inner capsid protein lambda-1, Inner capsid protein sigma-2, Outer capsid protein mu-1, ... | | 著者 | Sutton, G, Sun, D.P, Fu, X.F, Kotecha, A, Hecksel, G.W, Clare, D.K, Zhang, P, Stuart, D, Boyce, M. | | 登録日 | 2020-06-15 | | 公開日 | 2020-09-23 | | 実験手法 | ELECTRON MICROSCOPY (6.5 Å) | | 主引用文献 | Assembly intermediates of orthoreovirus captured in the cell.
Nat Commun, 11, 2020
|
|
7W3H
 
 | | Structure of USP14-bound human 26S proteasome in substrate-engaged state ED2.1_USP14 | | 分子名称: | 26S protease regulatory subunit 4, 26S protease regulatory subunit 6A, 26S protease regulatory subunit 6B, ... | | 著者 | Zhang, S, Zou, S, Yin, D, Wu, Z, Mao, Y. | | 登録日 | 2021-11-25 | | 公開日 | 2022-05-04 | | 最終更新日 | 2022-06-01 | | 実験手法 | ELECTRON MICROSCOPY (3.2 Å) | | 主引用文献 | USP14-regulated allostery of the human proteasome by time-resolved cryo-EM.
Nature, 605, 2022
|
|
8QRY
 
 | |
7W39
 
 | | Structure of USP14-bound human 26S proteasome in state EA2.1_UBL | | 分子名称: | 26S protease regulatory subunit 4, 26S protease regulatory subunit 6A, 26S protease regulatory subunit 6B, ... | | 著者 | Zhang, S, Zou, S, Yin, D, Wu, Z, Mao, Y. | | 登録日 | 2021-11-25 | | 公開日 | 2022-05-04 | | 最終更新日 | 2022-06-01 | | 実験手法 | ELECTRON MICROSCOPY (3.2 Å) | | 主引用文献 | USP14-regulated allostery of the human proteasome by time-resolved cryo-EM.
Nature, 605, 2022
|
|
7W3J
 
 | | Structure of USP14-bound human 26S proteasome in substrate-inhibited state SC_USP14 | | 分子名称: | 26S protease regulatory subunit 4, 26S protease regulatory subunit 6A, 26S protease regulatory subunit 6B, ... | | 著者 | Zhang, S, Zou, S, Yin, D, Wu, Z, Mao, Y. | | 登録日 | 2021-11-25 | | 公開日 | 2022-05-04 | | 最終更新日 | 2022-06-01 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | USP14-regulated allostery of the human proteasome by time-resolved cryo-EM.
Nature, 605, 2022
|
|
7W3B
 
 | | Structure of USP14-bound human 26S proteasome in substrate-engaged state ED5_USP14 | | 分子名称: | 26S protease regulatory subunit 4, 26S protease regulatory subunit 6A, 26S protease regulatory subunit 6B, ... | | 著者 | Zhang, S, Zou, S, Yin, D, Wu, Z, Mao, Y. | | 登録日 | 2021-11-25 | | 公開日 | 2022-05-04 | | 最終更新日 | 2022-06-01 | | 実験手法 | ELECTRON MICROSCOPY (3.6 Å) | | 主引用文献 | USP14-regulated allostery of the human proteasome by time-resolved cryo-EM.
Nature, 605, 2022
|
|
7W3A
 
 | | Structure of USP14-bound human 26S proteasome in substrate-engaged state ED4_USP14 | | 分子名称: | 26S protease regulatory subunit 4, 26S protease regulatory subunit 6A, 26S protease regulatory subunit 6B, ... | | 著者 | Zhang, S, Zou, S, Yin, D, Wu, Z, Mao, Y. | | 登録日 | 2021-11-25 | | 公開日 | 2022-05-04 | | 最終更新日 | 2022-06-01 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | USP14-regulated allostery of the human proteasome by time-resolved cryo-EM.
Nature, 605, 2022
|
|
7W3G
 
 | | Structure of USP14-bound human 26S proteasome in substrate-engaged state ED2.0_USP14 | | 分子名称: | 26S protease regulatory subunit 4, 26S protease regulatory subunit 6A, 26S protease regulatory subunit 6B, ... | | 著者 | Zhang, S, Zou, S, Yin, D, Wu, Z, Mao, Y. | | 登録日 | 2021-11-25 | | 公開日 | 2022-05-04 | | 最終更新日 | 2022-06-01 | | 実験手法 | ELECTRON MICROSCOPY (3.2 Å) | | 主引用文献 | USP14-regulated allostery of the human proteasome by time-resolved cryo-EM.
Nature, 605, 2022
|
|
7W37
 
 | | Structure of USP14-bound human 26S proteasome in state EA1_UBL | | 分子名称: | 26S protease regulatory subunit 4, 26S protease regulatory subunit 6A, 26S protease regulatory subunit 6B, ... | | 著者 | Zhang, S, Zou, S, Yin, D, Wu, Z, Mao, Y. | | 登録日 | 2021-11-25 | | 公開日 | 2022-05-04 | | 最終更新日 | 2022-06-01 | | 実験手法 | ELECTRON MICROSCOPY (3 Å) | | 主引用文献 | USP14-regulated allostery of the human proteasome by time-resolved cryo-EM.
Nature, 605, 2022
|
|
1AHO
 
 | | THE AB INITIO STRUCTURE DETERMINATION AND REFINEMENT OF A SCORPION PROTEIN TOXIN | | 分子名称: | TOXIN II | | 著者 | Smith, G.D, Blessing, R.H, Ealick, S.E, Fontecilla-Camps, J.C, Hauptman, H.A, Housset, D, Langs, D.A, Miller, R. | | 登録日 | 1997-04-08 | | 公開日 | 1997-10-15 | | 最終更新日 | 2024-06-05 | | 実験手法 | X-RAY DIFFRACTION (0.96 Å) | | 主引用文献 | Ab initio structure determination and refinement of a scorpion protein toxin.
Acta Crystallogr.,Sect.D, 53, 1997
|
|
7W3K
 
 | | Structure of USP14-bound human 26S proteasome in substrate-inhibited state SD4_USP14 | | 分子名称: | 26S protease regulatory subunit 4, 26S protease regulatory subunit 6A, 26S protease regulatory subunit 6B, ... | | 著者 | Zhang, S, Zou, S, Yin, D, Wu, Z, Mao, Y. | | 登録日 | 2021-11-25 | | 公開日 | 2022-05-04 | | 最終更新日 | 2022-06-01 | | 実験手法 | ELECTRON MICROSCOPY (3.6 Å) | | 主引用文献 | USP14-regulated allostery of the human proteasome by time-resolved cryo-EM.
Nature, 605, 2022
|
|
1ABB
 
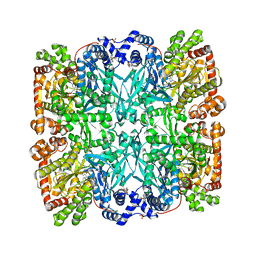 | | CONTROL OF PHOSPHORYLASE B CONFORMATION BY A MODIFIED COFACTOR: CRYSTALLOGRAPHIC STUDIES ON R-STATE GLYCOGEN PHOSPHORYLASE RECONSTITUTED WITH PYRIDOXAL 5'-DIPHOSPHATE | | 分子名称: | GLYCOGEN PHOSPHORYLASE B, INOSINIC ACID, PYRIDOXAL-5'-DIPHOSPHATE, ... | | 著者 | Leonidas, D.D, Oikonomakos, N.G, Papageorgiou, A.C, Acharya, K.R, Barford, D, Johnson, L.N. | | 登録日 | 1992-04-09 | | 公開日 | 1993-10-31 | | 最終更新日 | 2024-06-05 | | 実験手法 | X-RAY DIFFRACTION (2.8 Å) | | 主引用文献 | Control of phosphorylase b conformation by a modified cofactor: crystallographic studies on R-state glycogen phosphorylase reconstituted with pyridoxal 5'-diphosphate.
Protein Sci., 1, 1992
|
|
7W3C
 
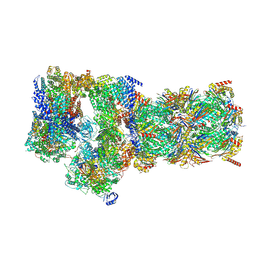 | | Structure of USP14-bound human 26S proteasome in substrate-engaged state ED0_USP14 | | 分子名称: | 26S protease regulatory subunit 4, 26S protease regulatory subunit 6A, 26S protease regulatory subunit 6B, ... | | 著者 | Zhang, S, Zou, S, Yin, D, Wu, Z, Mao, Y. | | 登録日 | 2021-11-25 | | 公開日 | 2022-05-04 | | 最終更新日 | 2022-06-01 | | 実験手法 | ELECTRON MICROSCOPY (3.4 Å) | | 主引用文献 | USP14-regulated allostery of the human proteasome by time-resolved cryo-EM.
Nature, 605, 2022
|
|
3WW7
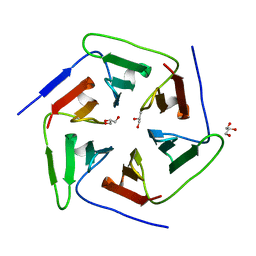 
 | | Crystal structure of the computationally designed Pizza2 protein | | 分子名称: | GLYCEROL, Pizza2 protein | | 著者 | Voet, A.R.D, Noguchi, H, Addy, C, Simoncini, D, Terada, D, Unzai, S, Park, S.Y, Zhang, K.Y.J, Tame, J.R.H. | | 登録日 | 2014-06-17 | | 公開日 | 2014-10-08 | | 最終更新日 | 2024-03-20 | | 実験手法 | X-RAY DIFFRACTION (1.697 Å) | | 主引用文献 | Computational design of a self-assembling symmetrical beta-propeller protein.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
8QV3
 
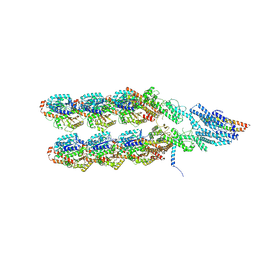 | | Structure of the y-Tubulin Small Complex (yTuSC) as part of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body | | 分子名称: | GUANOSINE-5'-TRIPHOSPHATE, Spindle pole body component, Spindle pole body component 110, ... | | 著者 | Dendooven, T, Yatskevich, S, Burt, A, Bellini, D, Kilmartin, J, Barford, D. | | 登録日 | 2023-10-17 | | 公開日 | 2024-04-24 | | 実験手法 | ELECTRON MICROSCOPY (8.2 Å) | | 主引用文献 | Structure of the native gamma-tubulin ring complex capping spindle microtubules.
Nat.Struct.Mol.Biol., 2024
|
|
8QV2
 
 | | Structure of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body | | 分子名称: | GUANOSINE-5'-DIPHOSPHATE, GUANOSINE-5'-TRIPHOSPHATE, Spindle pole body component, ... | | 著者 | Dendooven, T, Yatskevich, S, Burt, A, Bellini, D, Kilmartin, J, Barford, D. | | 登録日 | 2023-10-17 | | 公開日 | 2024-04-24 | | 実験手法 | ELECTRON MICROSCOPY (9.2 Å) | | 主引用文献 | Structure of the native gamma-tubulin ring complex capping spindle microtubules.
Nat.Struct.Mol.Biol., 2024
|
|
1AYE
 
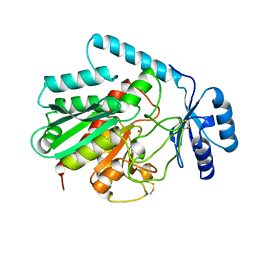 | | HUMAN PROCARBOXYPEPTIDASE A2 | | 分子名称: | PROCARBOXYPEPTIDASE A2, ZINC ION | | 著者 | Garcia-Saez, I, Reverte, D, Vendrell, J, Aviles, F.X, Coll, M. | | 登録日 | 1997-11-03 | | 公開日 | 1999-01-13 | | 最終更新日 | 2023-08-02 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | The three-dimensional structure of human procarboxypeptidase A2. Deciphering the basis of the inhibition, activation and intrinsic activity of the zymogen.
EMBO J., 16, 1997
|
|
1U9C
 
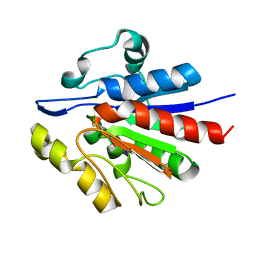 | | Crystallographic structure of APC35852 | | 分子名称: | APC35852 | | 著者 | Borek, D, Chen, Y, Shao, D, Collart, F, Joachimiak, A, Otwinowski, Z, Midwest Center for Structural Genomics (MCSG) | | 登録日 | 2004-08-09 | | 公開日 | 2004-10-05 | | 最終更新日 | 2023-08-23 | | 実験手法 | X-RAY DIFFRACTION (1.35 Å) | | 主引用文献 | Structural analysis of DJI superfamily
To be Published
|
|
5MVQ
 
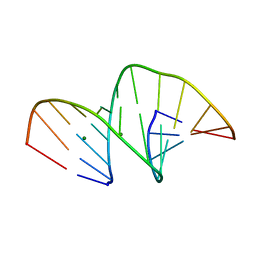 | | Crystal structure of an unmodified, self-complementary dodecamer. | | 分子名称: | DNA, MAGNESIUM ION | | 著者 | Hardwick, J.S, Ptchelkine, D, Phillips, S.E.V, Brown, T. | | 登録日 | 2017-01-17 | | 公開日 | 2017-05-10 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (1.604 Å) | | 主引用文献 | 5-Formylcytosine does not change the global structure of DNA.
Nat. Struct. Mol. Biol., 24, 2017
|
|
5MVU
 
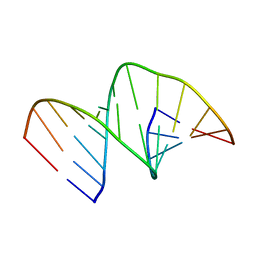 | |
