1E79
 
 | | Bovine F1-ATPase inhibited by DCCD (dicyclohexylcarbodiimide) | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, ATP SYNTHASE ALPHA CHAIN HEART ISOFORM, ... | | Authors: | Gibbons, C, Montgomery, M.G, Leslie, A.G.W, Walker, J.E. | | Deposit date: | 2000-08-25 | | Release date: | 2000-11-03 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The Structure of the Central Stalk in Bovine F(1)-ATPase at 2.4 A Resolution.
Nat.Struct.Biol., 7, 2000
|
|
1FSU
 
 | | Crystal Structure of 4-Sulfatase (human) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Bond, C, Guss, M. | | Deposit date: | 1996-07-29 | | Release date: | 1998-02-04 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of a human lysosomal sulfatase.
Structure, 5, 1997
|
|
8PWZ
 
 | | Crystal Structure of (3R)-hydroxyacyl-ACP dehydratase HadBD from Mycobacterium tuberculosis | | Descriptor: | (3R)-hydroxyacyl-ACP dehydratase subunit HadB, UPF0336 protein Rv0504c | | Authors: | Rima, J, Grimoire, Y, Bories, P, Bardou, F, Quemard, A, Bon, C, Mourey, L, Tranier, S. | | Deposit date: | 2023-07-22 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.00196719 Å) | | Cite: | HadBD dehydratase from Mycobacterium tuberculosis fatty acid synthase type II: A singular structure for a unique function.
Protein Sci., 33, 2024
|
|
3TZZ
 
 | | Crystal structure of a fragment containing the acyltransferase domain of Pks13 from Mycobacterium tuberculosis in the carboxypalmitoylated form at 2.5 A | | Descriptor: | 12-mer peptide, 2-(2-METHOXYETHOXY)ETHANOL, GLYCEROL, ... | | Authors: | Bergeret, F, Pedelacq, J.D, Mourey, L, Bon, C. | | Deposit date: | 2011-09-28 | | Release date: | 2012-08-29 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Biochemical and structural study of the atypical acyltransferase domain from the mycobacterial polyketide synthase pks13
J.Biol.Chem., 287, 2012
|
|
3TZW
 
 | | Crystal structure of a fragment containing the acyltransferase domain of Pks13 from Mycobacterium tuberculosis in the orthorhombic apoform at 2.6 A | | Descriptor: | 1,2-ETHANEDIOL, 12-mer peptide, Polyketide synthase PKS13, ... | | Authors: | Bergeret, F, Pedelacq, J.D, Mourey, L, Bon, C. | | Deposit date: | 2011-09-28 | | Release date: | 2012-08-29 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Biochemical and structural study of the atypical acyltransferase domain from the mycobacterial polyketide synthase pks13
J.Biol.Chem., 287, 2012
|
|
3TZY
 
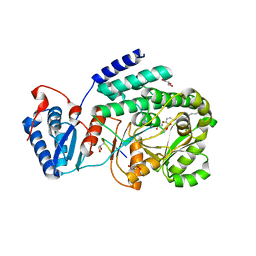 | | Crystal structure of a fragment containing the acyltransferase domain of Pks13 from Mycobacterium tuberculosis in the palmitoylated form at 2.2 A | | Descriptor: | 12-mer peptide, GLYCEROL, PALMITIC ACID, ... | | Authors: | Bergeret, F, Pedelacq, J.D, Mourey, L, Bon, C. | | Deposit date: | 2011-09-28 | | Release date: | 2012-08-29 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Biochemical and structural study of the atypical acyltransferase domain from the mycobacterial polyketide synthase pks13
J.Biol.Chem., 287, 2012
|
|
3TZX
 
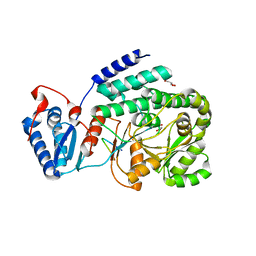 | | Crystal structure of a fragment containing the acyltransferase domain of Pks13 from Mycobacterium tuberculosis in tetragonal apo form at 2.3 A | | Descriptor: | 12-mer peptide, GLYCEROL, Polyketide synthase PKS13, ... | | Authors: | Bergeret, F, Pedelacq, J.D, Mourey, L, Bon, C. | | Deposit date: | 2011-09-28 | | Release date: | 2012-08-29 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Biochemical and structural study of the atypical acyltransferase domain from the mycobacterial polyketide synthase pks13
J.Biol.Chem., 287, 2012
|
|
1UOS
 
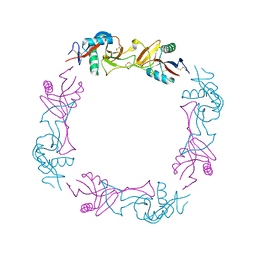 | | The Crystal Structure of the Snake Venom Toxin Convulxin | | Descriptor: | CONVULXIN ALPHA, CONVULXIN BETA | | Authors: | Batuwangala, T, Leduc, M, Gibbins, J.M, Bon, C, Jones, E.Y. | | Deposit date: | 2003-09-22 | | Release date: | 2003-10-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of the Snake-Venom Toxin Convulxin
Acta Crystallogr.,Sect.D, 60, 2004
|
|
1US0
 
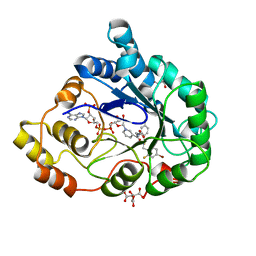 | | Human Aldose Reductase in complex with NADP+ and the inhibitor IDD594 at 0.66 Angstrom | | Descriptor: | ALDOSE REDUCTASE, CITRIC ACID, IDD594, ... | | Authors: | Howard, E.I, Sanishvili, R, Cachau, R.E, Mitschler, A, Chevrier, B, Barth, P, Lamour, V, Van Zandt, M, Sibley, E, Bon, C, Moras, D, Schneider, T.R, Joachimiak, A, Podjarny, A. | | Deposit date: | 2003-11-16 | | Release date: | 2004-05-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (0.66 Å) | | Cite: | Ultrahigh Resolution Drug Design I: Details of Interactions in Human Aldose Reductase-Inhibitor Complex at 0.66 A.
Proteins, 55, 2004
|
|
4ZAR
 
 | | Crystal Structure of Proteinase K from Engyodontium albuminhibited by METHOXYSUCCINYL-ALA-ALA-PRO-PHE-CHLOROMETHYL KETONE at 1.15 A resolution | | Descriptor: | CALCIUM ION, METHOXYSUCCINYL-ALA-ALA-PRO-PHE-CHLOROMETHYL KETONE, bound form, ... | | Authors: | Sawaya, M.R, Cascio, D, Collazo, M, Bond, C, Cohen, A, DeNicola, A, Eden, K, Jain, K, Leung, C, Lubock, N, McCormick, J, Rosinski, J, Spiegelman, L, Athar, Y, Tibrewal, N, Winter, J, Solomon, S. | | Deposit date: | 2015-04-14 | | Release date: | 2015-05-06 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Crystal Structure of Proteinase K from Engyodontium album inhibited by METHOXYSUCCINYL-ALA-ALA-PRO-PHE-CHLOROMETHYL KETONE at 1.15 A resolution
to be published
|
|
6CDX
 
 | | High-resolution crystal structure of fluoropropylated cystine knot, binding to alpha-5 beta-6 integrin | | Descriptor: | cystine knot (fluoropropylated) | | Authors: | Kimura, R, Nix, J, Bongura, C, Chakraborti, S, Gambhir, S, Filipp, F.V. | | Deposit date: | 2018-02-09 | | Release date: | 2019-08-14 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1 Å) | | Cite: | Evaluation of integrin alpha v beta6cystine knot PET tracers to detect cancer and idiopathic pulmonary fibrosis.
Nat Commun, 10, 2019
|
|
4BWU
 
 | | Three-dimensional structure of the K109A mutant of Paracoccus pantotrophus pseudoazurin at pH 5.5 | | Descriptor: | COPPER (II) ION, PSEUDOAZURIN, SULFATE ION | | Authors: | Freire, F, Mestre, A, Pinho, J, Najmudin, S, Bonifacio, C, Pauleta, S.R, Romao, M.J. | | Deposit date: | 2013-07-04 | | Release date: | 2014-07-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Exploring the Surface Determinants of Paracoccus Pantotrophus Pseudoazurin
To be Published
|
|
4BWT
 
 | | Three-dimensional structure of Paracoccus pantotrophus pseudoazurin at pH 6.5 | | Descriptor: | COPPER (II) ION, PSEUDOAZURIN, SULFATE ION | | Authors: | Freire, F, Mestre, A, Pinho, J, Najmudin, S, Bonifacio, C, Pauleta, S.R, Romao, M.J. | | Deposit date: | 2013-07-04 | | Release date: | 2014-07-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Exploring the Surface Determinants of Paracoccus Pantotrophus Pseudoazurin
To be Published
|
|
4BXV
 
 | | Three-dimensional structure of the mutant K109A of Paracoccus pantotrophus pseudoazurin at pH 7.0 | | Descriptor: | COPPER (II) ION, PSEUDOAZURIN, SULFATE ION | | Authors: | Freire, F, Mestre, A, Pinho, J, Najmudin, S, Bonifacio, C, Pauleta, S.R, Romao, M.J. | | Deposit date: | 2013-07-15 | | Release date: | 2014-07-30 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Exploring the Surface Determinants of Paracoccus Pantotrophus Pseudoazurin
To be Published
|
|
2OR2
 
 | | Structure of the W47A/W242A Mutant of Bacterial Phosphatidylinositol-Specific Phospholipase C | | Descriptor: | 1-phosphatidylinositol phosphodiesterase | | Authors: | Shao, C, Shi, X, Wehbi, H, Zambonelli, C, Head, J.F, Seaton, B.A, Roberts, M.F. | | Deposit date: | 2007-02-01 | | Release date: | 2007-02-13 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Dimer structure of an interfacially impaired phosphatidylinositol-specific phospholipase C.
J.Biol.Chem., 282, 2007
|
|
3EA1
 
 | | Crystal Structure of the Y247S/Y251S Mutant of Phosphatidylinositol-Specific Phospholipase C from Bacillus Thuringiensis | | Descriptor: | 1-phosphatidylinositol phosphodiesterase, ZINC ION | | Authors: | Shi, X, Shao, C, Zhang, X, Zambonelli, C, Redfied, A.G, Head, J.F, Seaton, B.A, Roberts, M.F. | | Deposit date: | 2008-08-24 | | Release date: | 2009-04-14 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Modulation of bacillus thuringiensis phosphatidylinositol-specific phospholipase C activity by mutations in the putative dimerization interface.
J.Biol.Chem., 284, 2009
|
|
3EA3
 
 | | Crystal Structure of the Y246S/Y247S/Y248S/Y251S Mutant of Phosphatidylinositol-Specific Phospholipase C from Bacillus Thuringiensis | | Descriptor: | 1-phosphatidylinositol phosphodiesterase, MANGANESE (II) ION | | Authors: | Shi, X, Shao, C, Zhang, X, Zambonelli, C, Redfied, A.G, Head, J.F, Seaton, B.A, Roberts, M.F. | | Deposit date: | 2008-08-24 | | Release date: | 2009-04-14 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Modulation of Bacillus thuringiensis Phosphatidylinositol-specific Phospholipase C Activity by Mutations in the Putative Dimerization Interface.
J.Biol.Chem., 284, 2009
|
|
2P1E
 
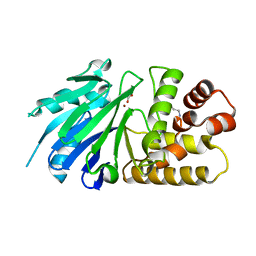 | | Crystal structure of the Leishmania infantum glyoxalase II with D-Lactate at the active site | | Descriptor: | Glyoxalase II, LACTIC ACID, SPERMIDINE, ... | | Authors: | Trincao, J, Barata, L, Najmudin, S, Bonifacio, C, Romao, M.J. | | Deposit date: | 2007-03-05 | | Release date: | 2008-01-15 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Catalysis and Structural Properties of Leishmania infantum Glyoxalase II: Trypanothione Specificity and Phylogeny.
Biochemistry, 47, 2008
|
|
3EA2
 
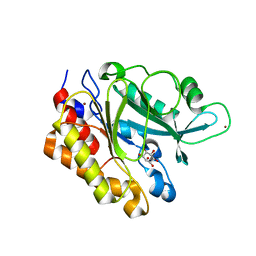 | | Crystal Structure of the Myo-inositol bound Y247S/Y251S Mutant of Phosphatidylinositol-Specific Phospholipase C from Bacillus Thuringiensis | | Descriptor: | 1,2,3,4,5,6-HEXAHYDROXY-CYCLOHEXANE, 1-phosphatidylinositol phosphodiesterase, ZINC ION | | Authors: | Shi, X, Shao, C, Zhang, X, Zambonelli, C, Redfied, A.G, Head, J.F, Seaton, B.A, Roberts, M.F. | | Deposit date: | 2008-08-24 | | Release date: | 2009-04-14 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Modulation of bacillus thuringiensis phosphatidylinositol-specific phospholipase C activity by mutations in the putative dimerization interface.
J.Biol.Chem., 284, 2009
|
|
2P18
 
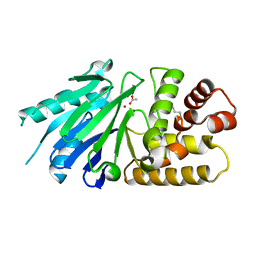 | | Crystal structure of the Leishmania infantum glyoxalase II | | Descriptor: | ACETIC ACID, Glyoxalase II, SPERMIDINE, ... | | Authors: | Trincao, J, Barata, L, Najmudin, S, Bonifacio, C, Romao, M.J. | | Deposit date: | 2007-03-02 | | Release date: | 2008-01-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Catalysis and Structural Properties of Leishmania infantum Glyoxalase II: Trypanothione Specificity and Phylogeny.
Biochemistry, 47, 2008
|
|
2H4O
 
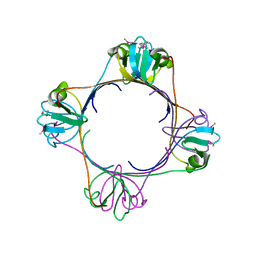 | | X-ray Crystal Structure of Protein yonK from Bacillus subtilis. Northeast Structural Genomics Consortium Target SR415 | | Descriptor: | YonK protein | | Authors: | Seetharaman, J, Sue, M, Forouhar, F, Ken, C, Bonnie, C, Ma, L, Xiao, R, Acton, T.B, Hunt, J.F, Tong, L, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2006-05-24 | | Release date: | 2006-07-25 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of the hypothetical protein from bacillus subtilis (yonk).
To be Published
|
|
2WFB
 
 | | High resolution structure of the apo form of the orange protein (ORP) from Desulfovibrio gigas | | Descriptor: | ACETATE ION, PHOSPHATE ION, PUTATIVE UNCHARACTERIZED PROTEIN ORP | | Authors: | Najmudin, S, Bonifacio, C, Duarte, A.G, Pereira, A.S, Moura, I, Moura, J.M, Romao, M.J. | | Deposit date: | 2009-04-03 | | Release date: | 2010-05-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | High Resolution Crystal Structure of the Apo Form of the Orange Protein (Apo-Orp) from Desulfovibrio Gigas
To be Published
|
|
2JJB
 
 | | Family 37 trehalase from Escherichia coli in complex with casuarine-6- O-alpha-glucopyranose | | Descriptor: | 1,2-ETHANEDIOL, CASUARINE, PERIPLASMIC TREHALASE, ... | | Authors: | Gloster, T.M, Roberts, S, Davies, G.J, Cardona, F, Parmeggiani, C, Bonaccini, C, Gratteri, P, Sim, L, Rose, D.R, Goti, A. | | Deposit date: | 2008-03-28 | | Release date: | 2009-01-13 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Total Syntheses of Casuarine and its 6-O-Alpha-Glucoside: Complementary Inhibition Towards Glycoside Hydrolases of the Gh31 and Gh37 Families.
Chemistry, 15, 2009
|
|
1F0I
 
 | | THE FIRST CRYSTAL STRUCTURE OF A PHOSPHOLIPASE D | | Descriptor: | PHOSPHATE ION, PHOSPHOLIPASE D | | Authors: | Leiros, I, Secundo, F, Zambonelli, C, Servi, S, Hough, E. | | Deposit date: | 2000-05-16 | | Release date: | 2001-05-16 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The first crystal structure of a phospholipase D.
Structure Fold.Des., 8, 2000
|
|
1RZ5
 
 | | Di-haem Cytochrome c Peroxidase, Form OUT | | Descriptor: | CALCIUM ION, Cytochrome c peroxidase, HEME C | | Authors: | Dias, J.M, Alves, T, Bonifacio, C, Pereira, A.S, Bourgeois, D, Moura, I, Romao, M.J. | | Deposit date: | 2003-12-24 | | Release date: | 2004-06-29 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for the mechanism of Ca(2+) activation of the di-heme cytochrome c peroxidase from Pseudomonas nautica 617.
Structure, 12, 2004
|
|
