5W8B
 
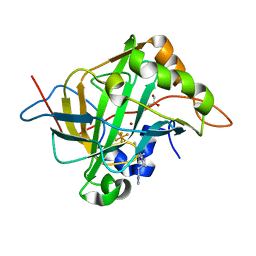 | | Carbonic anhydrase II in complex with activating histamine pyridinium derivative | | Descriptor: | 1-[2-(1H-imidazol-5-yl)ethyl]-4-methyl-2,6-di(propan-2-yl)pyridin-1-ium, Carbonic anhydrase 2, GLYCEROL, ... | | Authors: | Bhatt, A, Ilies, M, McKenna, R. | | Deposit date: | 2017-06-21 | | Release date: | 2018-05-30 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.601 Å) | | Cite: | Crystal Structure of Carbonic Anhydrase II in Complex with an Activating Ligand: Implications in Neuronal Function.
Mol. Neurobiol., 55, 2018
|
|
5SZ2
 
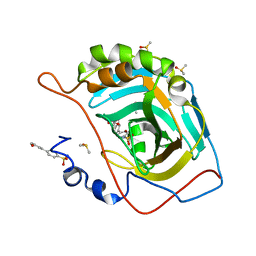 | | Carbonic anhydrase II in complex with 4-(3-formylphenyl)-benzenesulfonamide | | Descriptor: | 4-(3-formylphenyl)-benzenesulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Bhatt, A, Mahon, B.P, Cornelio, B, McKenna, R. | | Deposit date: | 2016-08-12 | | Release date: | 2016-12-21 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Structure-Activity Relationships of Benzenesulfonamide-Based Inhibitors towards Carbonic Anhydrase Isoform Specificity.
Chembiochem, 18, 2017
|
|
5SZ7
 
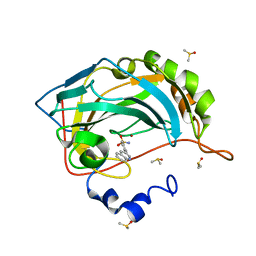 | | Carbonic anhydrase IX-mimic in complex with 4-(3-quinolinyl)-benzenesulfonamide | | Descriptor: | 4-(3-quinolinyl)-benzenesulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Bhatt, A, Mahon, B.P, Cornelio, B, McKenna, R. | | Deposit date: | 2016-08-12 | | Release date: | 2016-12-21 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.776 Å) | | Cite: | Structure-Activity Relationships of Benzenesulfonamide-Based Inhibitors towards Carbonic Anhydrase Isoform Specificity.
Chembiochem, 18, 2017
|
|
5SZ4
 
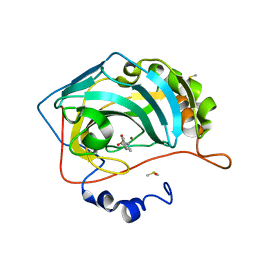 | | Carbonic anhydrase IX-mimic in complex with 4-(phenyl)-benzenesulfonamide | | Descriptor: | 4-(phenyl)-benzenesulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Bhatt, A, Mahon, B.P, Cornelio, B, McKenna, R. | | Deposit date: | 2016-08-12 | | Release date: | 2016-12-21 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure-Activity Relationships of Benzenesulfonamide-Based Inhibitors towards Carbonic Anhydrase Isoform Specificity.
Chembiochem, 18, 2017
|
|
5SZ0
 
 | | Carbonic anhydrase II in complex with 4-(phenyl)-benzenesulfonamide | | Descriptor: | 4-(phenyl)-benzenesulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Bhatt, A, Mahon, B.P, Cornelio, B, McKenna, R. | | Deposit date: | 2016-08-12 | | Release date: | 2016-12-21 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Structure-Activity Relationships of Benzenesulfonamide-Based Inhibitors towards Carbonic Anhydrase Isoform Specificity.
Chembiochem, 18, 2017
|
|
5SZ5
 
 | | Carbonic anhydrase IX-mimic in complex with 4-(2-methylphenyl)-benzenesulfonamide | | Descriptor: | 4-(2-methylphenyl)-benzenesulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Bhatt, A, Mahon, B.P, Cornelio, B, McKenna, R. | | Deposit date: | 2016-08-12 | | Release date: | 2016-12-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.274 Å) | | Cite: | Structure-Activity Relationships of Benzenesulfonamide-Based Inhibitors towards Carbonic Anhydrase Isoform Specificity.
Chembiochem, 18, 2017
|
|
5SZ1
 
 | | Carbonic anhydrase II in complex with 4-(2-methylphenyl)-benzenesulfonamide | | Descriptor: | 4-(2-methylphenyl)-benzenesulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Bhatt, A, Mahon, B.P, Cornelio, B, McKenna, R. | | Deposit date: | 2016-08-12 | | Release date: | 2016-12-21 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure-Activity Relationships of Benzenesulfonamide-Based Inhibitors towards Carbonic Anhydrase Isoform Specificity.
Chembiochem, 18, 2017
|
|
5SZ3
 
 | | Carbonic anhydrase II in complex with 4-(3-quinolinyl)-benzenesulfonamide | | Descriptor: | 4-(3-quinolinyl)-benzenesulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Bhatt, A, Mahon, B.P, Cornelio, B, McKenna, R. | | Deposit date: | 2016-08-12 | | Release date: | 2016-12-21 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.689 Å) | | Cite: | Structure-Activity Relationships of Benzenesulfonamide-Based Inhibitors towards Carbonic Anhydrase Isoform Specificity.
Chembiochem, 18, 2017
|
|
5SZ6
 
 | | Carbonic anhydrase IX-mimic in complex with 4-(3-formylphenyl)-benzenesulfonamide | | Descriptor: | 4-(3-formylphenyl)-benzenesulfonamide, Carbonic anhydrase 2, DIMETHYL SULFOXIDE, ... | | Authors: | Bhatt, A, Mahon, B.P, Cornelio, B, McKenna, R. | | Deposit date: | 2016-08-12 | | Release date: | 2016-12-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.15 Å) | | Cite: | Structure-Activity Relationships of Benzenesulfonamide-Based Inhibitors towards Carbonic Anhydrase Isoform Specificity.
Chembiochem, 18, 2017
|
|
7ONU
 
 | | Structure of human mitochondrial RNase P in complex with mitochondrial pre-tRNA-Tyr | | Descriptor: | 3-hydroxyacyl-CoA dehydrogenase type-2, MAGNESIUM ION, Mitochondrial Precursor tRNA-Tyr, ... | | Authors: | Bhatta, A, Dienemann, C, Cramer, P, Hillen, H.S. | | Deposit date: | 2021-05-26 | | Release date: | 2021-08-11 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structural basis of RNA processing by human mitochondrial RNase P.
Nat.Struct.Mol.Biol., 28, 2021
|
|
8RR3
 
 | | Human mitochondrial RNase Z complex with ELAC2-D550N catalytic mutant and tRNA-Gln precursor (Composite model) | | Descriptor: | 3-hydroxyacyl-CoA dehydrogenase type-2, Human mitochondrial tRNA-Gln precursor with 3' trailer RNA, S-ADENOSYL-L-HOMOCYSTEINE, ... | | Authors: | Bhatta, A, Yu, R.D, Kuhle, B, Hillen, H.S. | | Deposit date: | 2024-01-22 | | Release date: | 2025-01-08 | | Last modified: | 2025-04-23 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Molecular basis of human nuclear and mitochondrial tRNA 3' processing.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8RR1
 
 | | Human mitochondrial RNase Z complex with ELAC2-D550N catalytic mutant and tRNA-Tyr precursor (Composite model) | | Descriptor: | 3-hydroxyacyl-CoA dehydrogenase type-2, Human mitochondrial tRNA-Tyr precursor with 3' trailer, S-ADENOSYL-L-HOMOCYSTEINE, ... | | Authors: | Bhatta, A, Yu, R.D, Kuhle, B, Hillen, H.S. | | Deposit date: | 2024-01-22 | | Release date: | 2025-01-08 | | Last modified: | 2025-04-23 | | Method: | ELECTRON MICROSCOPY (2.93 Å) | | Cite: | Molecular basis of human nuclear and mitochondrial tRNA 3' processing.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8RR4
 
 | | Human mitochondrial RNase Z complex with ELAC2-D550N catalytic mutant with ordered flexible arm and tRNA-Tyr precursor - (Composite model) | | Descriptor: | 3-hydroxyacyl-CoA dehydrogenase type-2, Human mitochondria tRNA-Tyr precursor with 3' trailer, S-ADENOSYL-L-HOMOCYSTEINE, ... | | Authors: | Bhatta, A, Yu, R.D, Kuhle, B, Hillen, H.S. | | Deposit date: | 2024-01-22 | | Release date: | 2025-01-08 | | Last modified: | 2025-04-23 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Molecular basis of human nuclear and mitochondrial tRNA 3' processing.
Nat.Struct.Mol.Biol., 32, 2025
|
|
4J38
 
 | | Structure of Borrelia burgdorferi Outer surface protein E in complex with Factor H domains 19-20 | | Descriptor: | Complement factor H, Outer surface protein E, SULFATE ION | | Authors: | Bhattacharjee, A, Kolodziejczyk, R, Kajander, T, Goldman, A, Jokiranta, T.S. | | Deposit date: | 2013-02-05 | | Release date: | 2013-05-15 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | Structural Basis for Complement Evasion by Lyme Disease Pathogen Borrelia burgdorferi
J.Biol.Chem., 288, 2013
|
|
3KZJ
 
 | | Structure of complement Factor H variant R1203A | | Descriptor: | Complement factor H, SULFATE ION | | Authors: | Bhattacharjee, A, Lehtinen, M.J, Kajander, T, Goldman, A, Jokiranta, T.S. | | Deposit date: | 2009-12-08 | | Release date: | 2010-05-19 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Both domain 19 and domain 20 of factor H are involved in binding to complement C3b and C3d
Mol.Immunol., 47, 2010
|
|
1Q0P
 
 | | A domain of Factor B | | Descriptor: | Complement factor B, MANGANESE (II) ION | | Authors: | Bhattacharya, A.A, Liddington, R.C. | | Deposit date: | 2003-07-17 | | Release date: | 2004-03-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structure of the A Domain from Complement Factor B Reveals an Integrin-like Open Conformation.
STRUCTURE, 12, 2004
|
|
1E7E
 
 | |
1E7C
 
 | | HUMAN SERUM ALBUMIN COMPLEXED WITH MYRISTIC ACID and the general anesthetic halothane | | Descriptor: | 2-BROMO-2-CHLORO-1,1,1-TRIFLUOROETHANE, MYRISTIC ACID, SERUM ALBUMIN | | Authors: | Bhattacharya, A.A, Curry, S, Franks, N.P. | | Deposit date: | 2000-08-26 | | Release date: | 2001-01-17 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Binding of the General Anesthetics Propofol and Halothane to Human Serum Albumin. High Resolution Crystal Structures
J.Biol.Chem., 275, 2000
|
|
1E7F
 
 | |
1E7H
 
 | |
1E7A
 | |
1E7B
 
 | | Crystal structure of human serum albumin complexed with the general anesthetic halothane | | Descriptor: | 2-BROMO-2-CHLORO-1,1,1-TRIFLUOROETHANE, SERUM ALBUMIN | | Authors: | Bhattacharya, A.A, Curry, S, Franks, N.P. | | Deposit date: | 2000-08-26 | | Release date: | 2001-01-17 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Binding of the General Anesthetics Propofol and Halothane to Human Serum Albumin. High Resolution Crystal Structures
J.Biol.Chem., 275, 2000
|
|
1E78
 
 | |
1E7I
 
 | |
4MUC
 
 | | The 4th and 5th C-terminal domains of Factor H related protein 1 | | Descriptor: | Complement factor H-related protein 1, SULFATE ION | | Authors: | Bhattacharjee, A, Goldman, A, Kolodziejczyk, R, Jokiranta, T.S. | | Deposit date: | 2013-09-21 | | Release date: | 2015-02-18 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.897 Å) | | Cite: | The Major Autoantibody Epitope on Factor H in Atypical Hemolytic Uremic Syndrome Is Structurally Different from Its Homologous Site in Factor H-related Protein 1, Supporting a Novel Model for Induction of Autoimmunity in This Disease.
J.Biol.Chem., 290, 2015
|
|
