4NL8
 
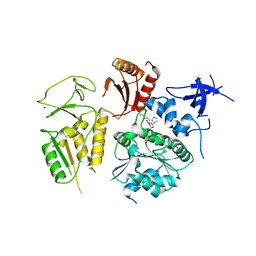 | | PriA Helicase Bound to SSB C-terminal Tail Peptide | | Descriptor: | Primosome assembly protein PriA, Single-stranded DNA-binding protein, ZINC ION | | Authors: | Bhattacharyya, B, George, N.P, Thurmes, T.M, Keck, J.L. | | Deposit date: | 2013-11-13 | | Release date: | 2014-01-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (4.08 Å) | | Cite: | Structural mechanisms of PriA-mediated DNA replication restart.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
4NL4
 
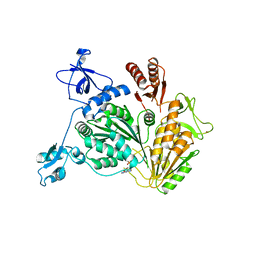 | | PriA Helicase Bound to ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Primosome assembly protein PriA, ZINC ION | | Authors: | Bhattacharyya, B, George, N.P, Keck, J.L. | | Deposit date: | 2013-11-13 | | Release date: | 2014-01-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structural mechanisms of PriA-mediated DNA replication restart.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
1EI7
 
 | |
3Q61
 
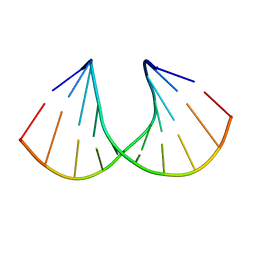 | | 3'-Fluoro Hexitol Nucleic Acid DNA Structure | | Descriptor: | DNA (5'-D(*GP*CP*GP*TP*AP*(F3H)P*AP*CP*GP*C)-3') | | Authors: | Seth, P.R, Allerson, C.R, Prakash, T.P, Siwkowski, A, Berdeja, A, Yu, J, Pallan, P.S, Watt, A.T, Gaus, H, Bhat, B, Egli, M, Swayze, E.E. | | Deposit date: | 2010-12-30 | | Release date: | 2012-01-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Synthesis, improved antisense activity and structural rationale for the divergent RNA affinities of 3'-fluoro hexitol nucleic acid (FHNA and Ara-FHNA) modified oligonucleotides.
J.Am.Chem.Soc., 133, 2011
|
|
2KWG
 
 | | Solution structure of a fully modified 2'-F/2'-OMe siRNA construct | | Descriptor: | 5'-R(*(GF2)P*(OMG)P*(GF2)P*(OMU)P*(AF2)P*(A2M)P*(AF2)P*(OMU)P*(AF2)P*(OMC)P*(AF2)P*(OMU)P*(UFT)P*(OMC)P*(UFT)P*(OMU)P*(CFZ)P*(A2M)P*(UFT)P*(OMU)P*(UFT))-3', 5'-R(P*(A2M)P*(UFT)P*(OMG)P*(AF2)P*(A2M)P*(GF2)P*(A2M)P*(AF2)P*(OMU)P*(GF2)P*(OMU)P*(AF2)P*(OMU)P*(UFT)P*(OMU)P*(AF2)P*(OMC)P*(CFZ)P*(OMC)P*(UFT)P*(OMU))-3' | | Authors: | Podbevsek, P, Bhat, B, Plavec, J. | | Deposit date: | 2010-04-09 | | Release date: | 2010-07-21 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution-state structure of a fully alternately 2'-F/2'-OMe modified 42-nt dimeric siRNA construct.
Nucleic Acids Res., 38, 2010
|
|
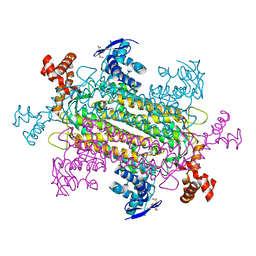
6P3C
 
 | |
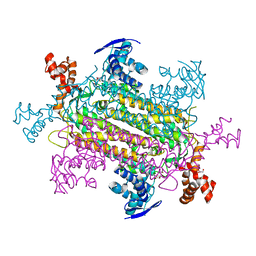
6OS7
 
 | |
6NZ9
 
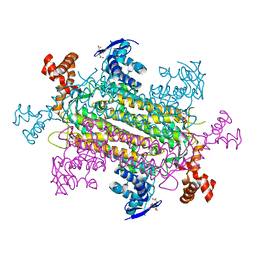 | | Crystal structure of E. coli fumarase C bound to citrate at 1.53 angstrom resolution | | Descriptor: | CITRIC ACID, Fumarate hydratase class II | | Authors: | Stuttgen, G.M, May, J.F, Bhattcharyya, B, Weaver, T.M. | | Deposit date: | 2019-02-13 | | Release date: | 2019-09-25 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.528 Å) | | Cite: | Closed fumarase C active-site structures reveal SS Loop residue contribution in catalysis.
Febs Lett., 594, 2020
|
|
8SBS
 
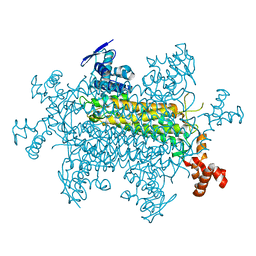 | |
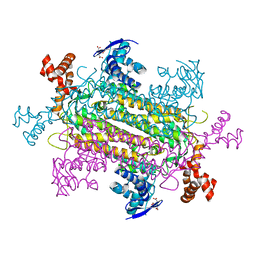
6NZC
 
 | |
6NZA
 
 | |
6NZB
 
 | |
5KKD
 
 | |
6PZL
 
 | |
6PYK
 
 | |
3INM
 
 | | Crystal structure of human cytosolic NADP(+)-dependent isocitrate dehydrogenase R132H mutant in complex with NADPH, ALPHA-KETOGLUTARATE and CALCIUM(2+) | | Descriptor: | 2-OXOGLUTARIC ACID, CALCIUM ION, GLYCEROL, ... | | Authors: | Fontano, E, Brown, R.S, Suto, R.K, Bhyravbhatla, B. | | Deposit date: | 2009-08-12 | | Release date: | 2009-11-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Cancer-associated IDH1 mutations produce 2-hydroxyglutarate.
Nature, 462, 2009
|
|
5KDK
 
 | |
5KEH
 
 | |
5KF3
 
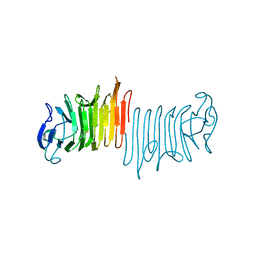 | |
6E8A
 
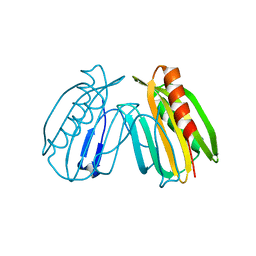 | | Crystal structure of DcrB from Salmonella enterica at 1.92 Angstroms resolution | | Descriptor: | DUF1795 domain-containing protein | | Authors: | Rasmussen, D.M, Soens, R.W, Bhattacharyya, B, May, J.F. | | Deposit date: | 2018-07-27 | | Release date: | 2018-10-31 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | The structure of DcrB, a lipoprotein from Salmonella enterica, reveals flexibility in the N-terminal segment of the Mog1p/PsbP-like fold.
J. Struct. Biol., 204, 2018
|
|
5SZ8
 
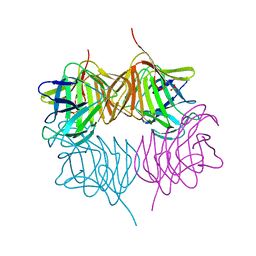 | |
3IYK
 
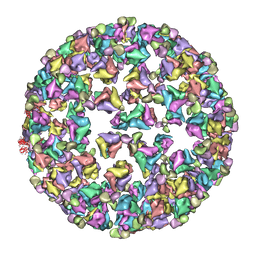 | | Bluetongue virus structure reveals a sialic acid binding domain, amphipathic helices and a central coiled coil in the outer capsid proteins | | Descriptor: | 2-O-methyl-5-N-acetyl-alpha-D-neuraminic acid, VP2, VP5 | | Authors: | Zhang, X, Boyce, M, Bhattacharya, B, Zhang, X, Schein, S, Roy, P, Zhou, Z.H. | | Deposit date: | 2010-01-25 | | Release date: | 2010-04-07 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (7 Å) | | Cite: | Bluetongue virus coat protein VP2 contains sialic acid-binding domains, and VP5 resembles enveloped virus fusion proteins.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
1X9F
 
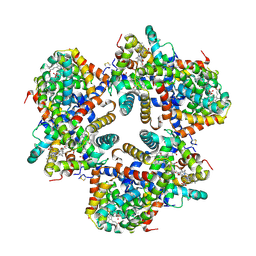 | | Hemoglobin Dodecamer from Lumbricus Erythrocruorin | | Descriptor: | CARBON MONOXIDE, Globin II, extracellular, ... | | Authors: | Strand, K, Knapp, J.E, Bhyravbhatla, B, Royer Jr, W.E. | | Deposit date: | 2004-08-20 | | Release date: | 2004-11-30 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of the hemoglobin dodecamer from lumbricus erythrocruorin: allosteric core of giant annelid respiratory complexes
J.Mol.Biol., 344, 2004
|
|
2GTL
 
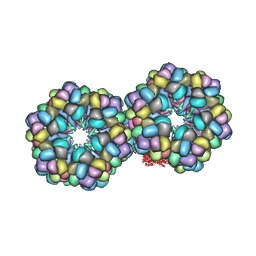 | | Lumbricus Erythrocruorin at 3.5A resolution | | Descriptor: | CALCIUM ION, CARBON MONOXIDE, Extracellular globin 2, ... | | Authors: | Royer Jr, W.E, Sharma, H, Strand, K, Knapp, J.E, Bhyravbhatla, B. | | Deposit date: | 2006-04-28 | | Release date: | 2006-07-18 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Lumbricus erythrocruorin at 3.5 a resolution: architecture of a megadalton respiratory complex.
Structure, 14, 2006
|
|
3QH1
 
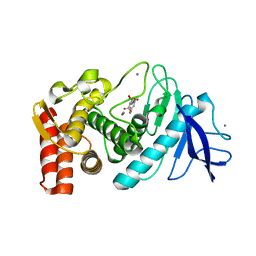 | | Structure of Thermolysin in complex with N-benzyloxycarbonyl-L-aspartic acid | | Descriptor: | CALCIUM ION, N-[(benzyloxy)carbonyl]-L-aspartic acid, Thermolysin, ... | | Authors: | Birrane, G, Bhyravbhatla, B, Navia, M. | | Deposit date: | 2011-01-25 | | Release date: | 2012-01-04 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Synthesis of Aspartame by Thermolysin: An X-ray Structural Study.
ACS MED.CHEM.LETT., 5, 2014
|
|
