4XYH
 
 | |
4XYI
 
 | | Mis16 with H4 peptide | | Descriptor: | Histone H4, Kinetochore protein Mis16 | | Authors: | An, S, Kim, H, Cho, U.-S. | | Deposit date: | 2015-02-02 | | Release date: | 2016-02-03 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Mis16 Independently Recognizes Histone H4 and the CENP-ACnp1-Specific Chaperone Scm3sp.
J.Mol.Biol., 427, 2015
|
|
7U5V
 
 | | Crystal structure of the Mixed Lineage Leukaemia (MLL1) SET Domain with the cofactor product S-Adenosylhomocysteine and Borealin peptide | | Descriptor: | Borealin, Histone-lysine N-methyltransferase 2A, S-ADENOSYL-L-HOMOCYSTEINE, ... | | Authors: | An, S, Cho, U.S, Oh, H, Sha, L, Xu, J, Kim, S, Yang, W, An, W, Dou, Y. | | Deposit date: | 2022-03-02 | | Release date: | 2023-09-27 | | Last modified: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Non-canonical MLL1 activity regulates centromeric phase separation and genome stability.
Nat.Cell Biol., 25, 2023
|
|
5W0V
 
 | | Crystal structure of full-length Kluyveromyces lactis Kap123 with histone H4 1-34 | | Descriptor: | Histone H4 1-34, Kap123 | | Authors: | An, S, Yoon, J, Song, J.-J, Cho, U.-S. | | Deposit date: | 2017-05-31 | | Release date: | 2017-11-01 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.821 Å) | | Cite: | Structure-based nuclear import mechanism of histones H3 and H4 mediated by Kap123.
Elife, 6, 2017
|
|
5WJC
 
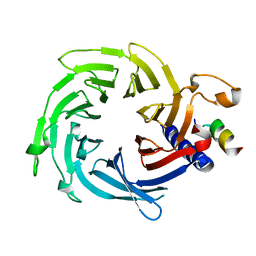 | | Crystal structure of Schizosaccharomyces pombe Mis16 in complex with Eic1 | | Descriptor: | Eic1 protein, Kinetochore protein Mis16 | | Authors: | An, S, Cho, U.-S, Koldewey, P, Chik, J, Subramanian, L. | | Deposit date: | 2017-07-21 | | Release date: | 2018-06-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.298 Å) | | Cite: | Mis16 Switches Function from a Histone H4 Chaperone to a CENP-ACnp1-Specific Assembly Factor through Eic1 Interaction.
Structure, 26, 2018
|
|
5VE8
 
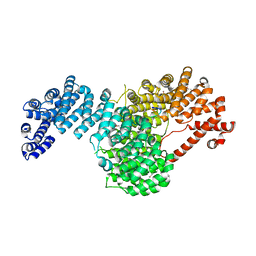 | | Crystal structure of full-length Kluyveromyces lactis Kap123 with histone H3 1-28 | | Descriptor: | Histone H3, Kap123 | | Authors: | An, S, Yoon, J, Song, J.-J, Cho, U.-S. | | Deposit date: | 2017-04-04 | | Release date: | 2017-11-01 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure-based nuclear import mechanism of histones H3 and H4 mediated by Kap123.
Elife, 6, 2017
|
|
5VCH
 
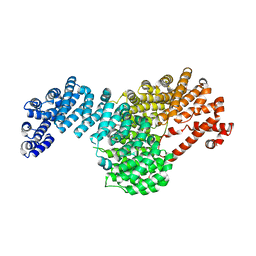 | |
6LKF
 
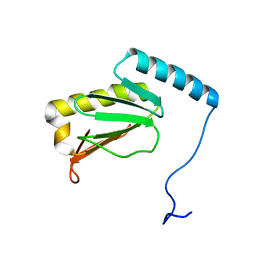 | |
3OPE
 
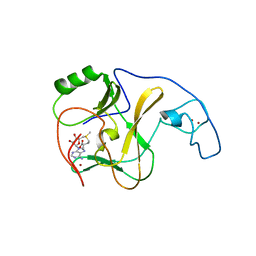 | | Structural Basis of Auto-inhibitory mechanism of Histone methyltransferase | | Descriptor: | Probable histone-lysine N-methyltransferase ASH1L, S-ADENOSYLMETHIONINE, ZINC ION | | Authors: | An, S, Song, J. | | Deposit date: | 2010-08-31 | | Release date: | 2011-01-12 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of the human histone methyltransferase ASH1L catalytic domain and its implications for the regulatory mechanism
J.Biol.Chem., 286, 2011
|
|
8JB9
 
 | |
5CUF
 
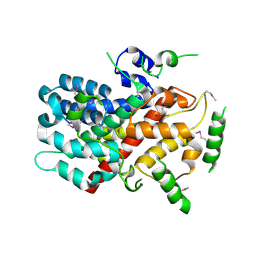 | | X-ray crystal structure of SeMet human Sestrin2 | | Descriptor: | Sestrin-2 | | Authors: | Kim, H, An, S, Ro, S.-H, Lee, J.H, Cho, U.-S. | | Deposit date: | 2015-07-24 | | Release date: | 2016-01-13 | | Last modified: | 2019-12-25 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Janus-faced Sestrin2 controls ROS and mTOR signalling through two separate functional domains.
Nat Commun, 6, 2015
|
|
5UCU
 
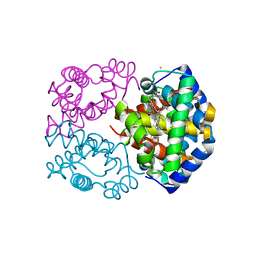 | | STRUCTURAL AND MECHANISTIC INSIGHTS INTO HEMOGLOBIN-CATALYZED HYDROGEN SULFIDE OXIDATION AND THE FATE OF POLYSULFIDE PRODUCTS | | Descriptor: | HYDROSULFURIC ACID, Hemoglobin subunit alpha, Hemoglobin subunit beta, ... | | Authors: | Vitvitsky, V, Yadav, P.K, An, S, Seravalli, J, Cho, U.-S, Banerjee, R. | | Deposit date: | 2016-12-22 | | Release date: | 2017-03-01 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.797 Å) | | Cite: | Structural and Mechanistic Insights into Hemoglobin-catalyzed Hydrogen Sulfide Oxidation and the Fate of Polysulfide Products.
J. Biol. Chem., 292, 2017
|
|
5Y6A
 
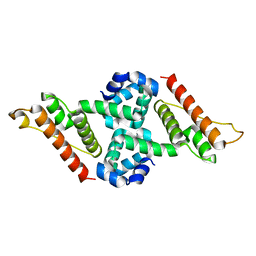 | | Crystal structure of the anti-CRISPR protein, AcrIIA1 | | Descriptor: | chain A and B | | Authors: | Ka, D, An, S.Y, Suh, J.Y, Bae, E. | | Deposit date: | 2017-08-11 | | Release date: | 2017-11-29 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of an anti-CRISPR protein, AcrIIA1
Nucleic Acids Res., 46, 2018
|
|
5Y69
 
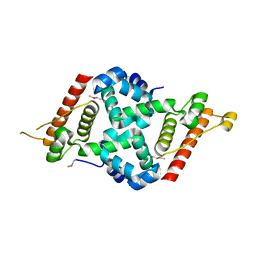 | | Crystal structure of the L52M mutant of AcrIIA1 | | Descriptor: | chain A | | Authors: | Ka, D, An, S.Y, Suh, J.Y, Bae, E. | | Deposit date: | 2017-08-11 | | Release date: | 2017-11-29 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure of an anti-CRISPR protein, AcrIIA1.
Nucleic Acids Res., 46, 2018
|
|
6M3N
 
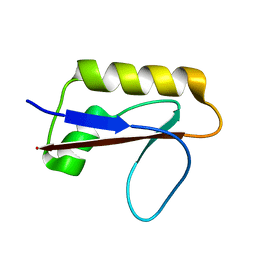 | | Solution structure of anti-CRISPR AcrIF7 | | Descriptor: | anti-CRIPSR AcrIF7 | | Authors: | Kim, I, An, S.Y, Koo, J, Bae, E, Suh, J.Y. | | Deposit date: | 2020-03-04 | | Release date: | 2020-08-26 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural and mechanistic insights into the CRISPR inhibition of AcrIF7.
Nucleic Acids Res., 48, 2020
|
|
2KNF
 
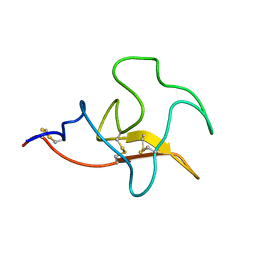 | | Solution structure and functional characterization of human plasminogen kringle 5 | | Descriptor: | Plasminogen | | Authors: | Battistel, M.D, Grishaev, A, An, S.A, Castellino, F.J, Llinas, M. | | Deposit date: | 2009-08-21 | | Release date: | 2009-10-27 | | Last modified: | 2021-10-13 | | Method: | SOLUTION NMR | | Cite: | Solution structure and functional characterization of human plasminogen kringle 5.
Biochemistry, 48, 2009
|
|
