5TYT
 
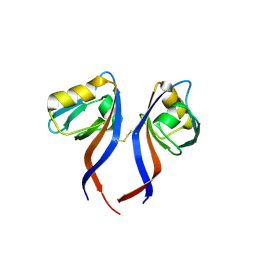 | | Crystal Structure of the PDZ domain of RhoGEF bound to CXCR2 C-terminal peptide | | Descriptor: | Rho guanine nucleotide exchange factor 11, C-X-C chemokine receptor type 2 chimera | | Authors: | Spellmon, N, Holcomb, J, Niu, A, Choudhary, V, Sun, X, Brunzelle, J, Li, C, Yang, Z. | | Deposit date: | 2016-11-21 | | Release date: | 2017-02-22 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.398 Å) | | Cite: | Structural basis of PDZ-mediated chemokine receptor CXCR2 scaffolding by guanine nucleotide exchange factor PDZ-RhoGEF.
Biochem. Biophys. Res. Commun., 485, 2017
|
|
8DN8
 
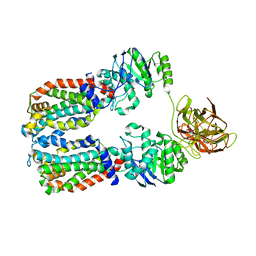 | |
8DKU
 
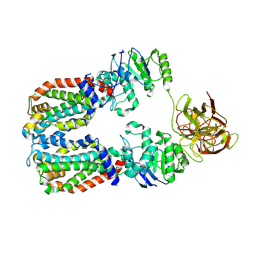 | | CryoEM structure of the A. aeolicus WzmWzt transporter bound to the native O antigen | | Descriptor: | 6-deoxy-3-O-methyl-alpha-D-mannopyranose-(1-3)-beta-D-rhamnopyranose-(1-2)-alpha-D-rhamnopyranose, ABC transporter, Transport permease protein | | Authors: | Spellmon, N, Muszynski, A, Vlach, J, Zimmer, J. | | Deposit date: | 2022-07-06 | | Release date: | 2022-09-21 | | Last modified: | 2025-05-14 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Molecular basis for polysaccharide recognition and modulated ATP hydrolysis by the O antigen ABC transporter.
Nat Commun, 13, 2022
|
|
8DNC
 
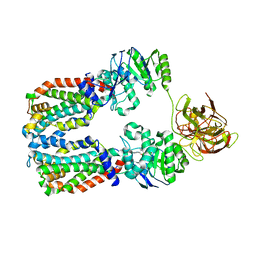 | | CryoEM structure of the A. aeolicus WzmWzt transporter bound to the native O antigen and ADP | | Descriptor: | 6-deoxy-3-O-methyl-alpha-D-mannopyranose-(1-3)-beta-D-rhamnopyranose-(1-2)-alpha-D-rhamnopyranose, ABC transporter, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Spellmon, N, Muszynski, A, Vlach, J, Zimmer, J. | | Deposit date: | 2022-07-11 | | Release date: | 2022-09-21 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Molecular basis for polysaccharide recognition and modulated ATP hydrolysis by the O antigen ABC transporter.
Nat Commun, 13, 2022
|
|
8DL0
 
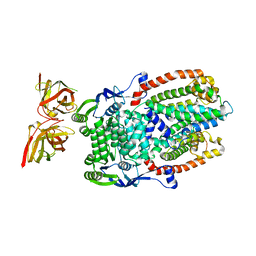 | |
8DKY
 
 | |
8DNE
 
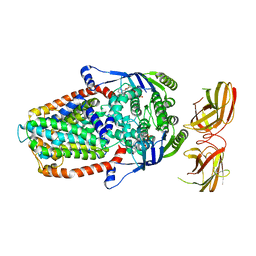 | |
6MON
 
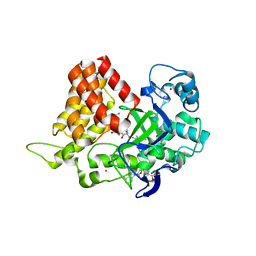 | | Crystal structure of human SMYD2 in complex with Nle-peptide inhibitor | | Descriptor: | GLYCEROL, LYS-LEU-NLE-SER-LYS-ARG-GLY, N-lysine methyltransferase SMYD2, ... | | Authors: | Spellmon, N, Cornett, E, Brunzelle, J, Rothbart, S, Yang, Z. | | Deposit date: | 2018-10-04 | | Release date: | 2018-12-12 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.711 Å) | | Cite: | A functional proteomics platform to reveal the sequence determinants of lysine methyltransferase substrate selectivity.
Sci Adv, 4, 2018
|
|
9E1T
 
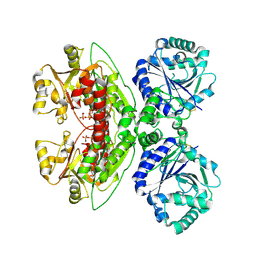 | | CryoEM structure of LARGE1 bound to UDP | | Descriptor: | MANGANESE (II) ION, URIDINE-5'-DIPHOSPHATE, Xylosyl- and glucuronyltransferase LARGE1 | | Authors: | Joesph, S, Spellmon, N, Campbell, K.P. | | Deposit date: | 2024-10-21 | | Release date: | 2025-02-12 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Prodystroglycan defines matriglycan lengths processively polymerized by LARGE1
To Be Published
|
|
9CKG
 
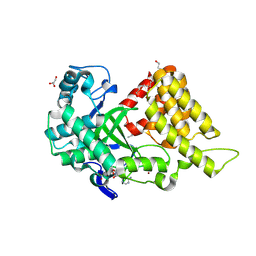 | | Crystal structure of SMYD2 active site mutant | | Descriptor: | DI(HYDROXYETHYL)ETHER, ETHANOL, GLYCEROL, ... | | Authors: | Sobota, J, Zhang, Y, Spellmon, N, Perry, E, Brunzelle, J, Yang, Z. | | Deposit date: | 2024-07-08 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Structure of the SMYD2-PARP1 Complex Reveals Both Productive and Allosteric Modes of Peptide Binding.
Biorxiv, 2024
|
|
9CKC
 
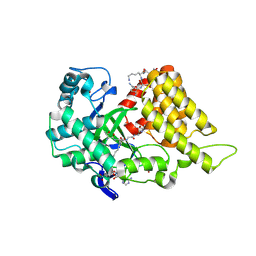 | | Crystal structure of SMYD2 in complex with two PARP1 peptides | | Descriptor: | N-lysine methyltransferase SMYD2, Poly [ADP-ribose] polymerase 1, processed C-terminus, ... | | Authors: | Zhang, Y, Alshammari, E, Spellmon, N, Brunzelle, J, Yang, Z. | | Deposit date: | 2024-07-08 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of the SMYD2-PARP1 Complex Reveals Both Productive and Allosteric Modes of Peptide Binding.
Biorxiv, 2024
|
|
9CKF
 
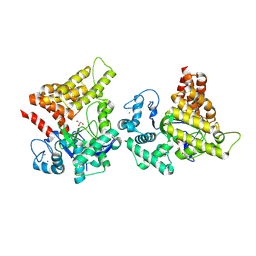 | | Crystal structure of SMYD2 secondary binding site mutant | | Descriptor: | GLYCEROL, N-lysine methyltransferase SMYD2, S-ADENOSYL-L-HOMOCYSTEINE, ... | | Authors: | Zhang, Y, Spellmon, N, Perry, E, Brunzelle, J, Yang, Z. | | Deposit date: | 2024-07-08 | | Release date: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the SMYD2-PARP1 Complex Reveals Both Productive and Allosteric Modes of Peptide Binding.
Biorxiv, 2024
|
|
8DOU
 
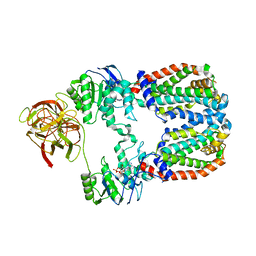 | |
6B6I
 
 | | 2.4A resolution structure of human Norovirus GII.4 protease | | Descriptor: | 3C-like protease | | Authors: | Muzzarelli, K.M, Kuiper, B.D, Spellmon, N.S, Hackett, J, Brunzelle, J.S, Kovari, I.A, Amblard, F, Yang, Z, Schinazi, R.F, Kovari, L.C. | | Deposit date: | 2017-10-02 | | Release date: | 2018-10-03 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Structural and Antiviral Studies of the Human Norovirus GII.4 Protease.
Biochemistry, 58, 2019
|
|
6N3G
 
 | | Crystal structure of histone lysine methyltransferase SmyD2 in complex with polyethylene glycol | | Descriptor: | DODECAETHYLENE GLYCOL, ETHANOL, N-lysine methyltransferase SMYD2, ... | | Authors: | Perry, E, Spellmon, N, Brunzelle, J, Yang, Z. | | Deposit date: | 2018-11-15 | | Release date: | 2019-11-20 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | Crystal structure of histone lysine methyltransferase SmyD2 in complex with polyethylene glycol
To be Published
|
|
