2L5R
 
 | |
2K10
 
 | |
9N2C
 
 | | Impacts of ribosomal RNA sequence variation on gene expression and phenotype: Cryo-EM structure of the rrsH ribosome (HBB-70S) | | Descriptor: | 1,4-DIAMINOBUTANE, 16S ribosomal RNA (rRNA) from the rrnH operon, 23S ribosomal RNA (rRNA) from the rrnB operon, ... | | Authors: | Welfer, G.A, Brady, R.A, Natchiar, S.K, Watson, Z.L, Rundlet, E.J, Alejo, J.L, Singh, A.P, Mishra, N.K, Altman, R.B, Blanchard, S.C. | | Deposit date: | 2025-01-28 | | Release date: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (2.4 Å) | | Cite: | Impacts of ribosomal RNA sequence variation on gene expression and phenotype.
Philos.Trans.R.Soc.Lond.B Biol.Sci., 380, 2025
|
|
9N2B
 
 | | Impacts of ribosomal RNA sequence variation on gene expression and phenotype: Cryo-EM structure of the rrsB ribosome (BBB-70S) | | Descriptor: | 1,4-DIAMINOBUTANE, 16S ribosomal RNA (rRNA) from the rrnB operon, 23S ribosomal RNA (rRNA) from the rrnB operon, ... | | Authors: | Welfer, G.A, Brady, R.A, Natchiar, S.K, Watson, Z.L, Rundlet, E.J, Alejo, J.L, Singh, A.P, Mishra, N.K, Altman, R.B, Blanchard, S.C. | | Deposit date: | 2025-01-28 | | Release date: | 2025-03-19 | | Method: | ELECTRON MICROSCOPY (2.2 Å) | | Cite: | Impacts of ribosomal RNA sequence variation on gene expression and phenotype.
Philos.Trans.R.Soc.Lond.B Biol.Sci., 380, 2025
|
|
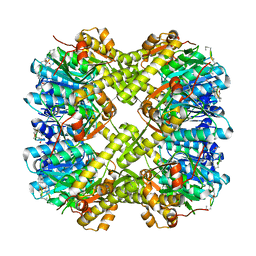
5VZ2
 | |
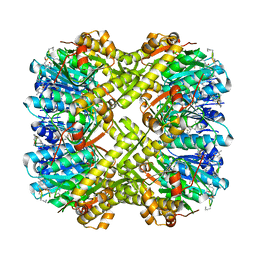
6PKA
 | |
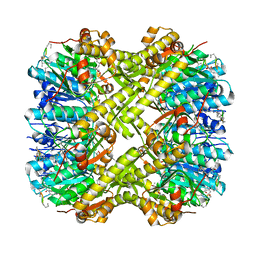
6PMD
 | |
6AXD
 
 | |
6ASR
 
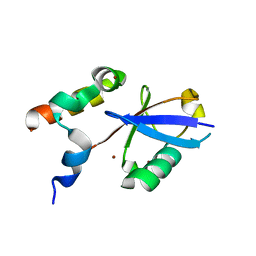 | | REV1 UBM2 domain complex with ubiquitin | | Descriptor: | DNA repair protein REV1, NICKEL (II) ION, Ubiquitin | | Authors: | Miller, D.J. | | Deposit date: | 2017-08-25 | | Release date: | 2018-06-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.356 Å) | | Cite: | Structures of REV1 UBM2 Domain Complex with Ubiquitin and with a Small-Molecule that Inhibits the REV1 UBM2-Ubiquitin Interaction.
J. Mol. Biol., 430, 2018
|
|
6CFD
 
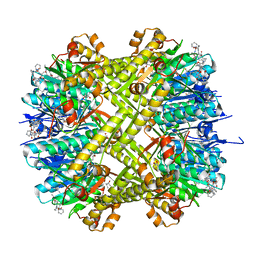 | | ADEP4 bound to E. faecium ClpP | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, ATP-dependent Clp protease proteolytic subunit, N-[(6aS,12S,15aS,17R,21R,23aS)-17,21-dimethyl-6,11,15,20,23-pentaoxooctadecahydro-2H,6H,11H,15H-pyrido[2,1-i]dipyrrolo[2,1-c:2',1'-l][1,4,7,10,13]oxatetraazacyclohexadecin-12-yl]-3,5-difluoro-Nalpha-[(2E)-hept-2-enoyl]-L-phenylalaninamide | | Authors: | Lee, R.E, Griffith, E.C. | | Deposit date: | 2018-02-14 | | Release date: | 2018-05-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | In VivoandIn VitroEffects of a ClpP-Activating Antibiotic against Vancomycin-Resistant Enterococci.
Antimicrob. Agents Chemother., 62, 2018
|
|
5W18
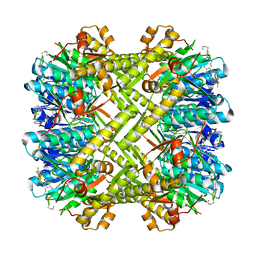 
 | | Staphylococcus aureus ClpP in complex with (S)-N-((2R,6S,8aS,14aS,20S,23aS)-2,6-dimethyl-5,8,14,19,23-pentaoxooctadecahydro-1H,5H,14H,19H-pyrido[2,1-i]dipyrrolo[2,1-c:2',1'-l][1]oxa[4,7,10,13]tetraazacyclohexadecin-20-yl)-3-phenyl-2-(3-phenylureido)propanamide | | Descriptor: | 9V7-PHE-SER-PRO-YCP-ALA-MP8, ATP-dependent Clp protease proteolytic subunit | | Authors: | Lee, R.E, Griffith, E.C. | | Deposit date: | 2017-06-02 | | Release date: | 2017-08-09 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Ureadepsipeptides as ClpP Activators.
Acs Infect Dis., 2019
|
|
4NXX
 
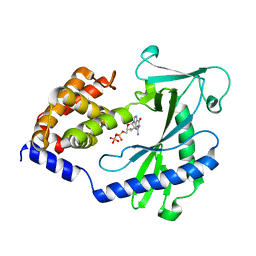 | |
4NXU
 
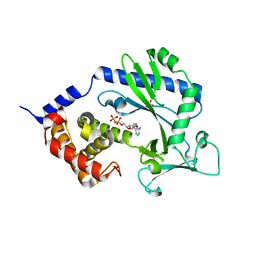 | | Crystal structure of the cytosolic domain of human MiD51 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, GLYCEROL, Mitochondrial dynamic protein MID51, ... | | Authors: | Richter, V, Kvansakul, M, Ryan, M.T. | | Deposit date: | 2013-12-09 | | Release date: | 2013-12-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and functional analysis of MiD51, a dynamin receptor required for mitochondrial fission.
J.Cell Biol., 204, 2014
|
|
4NXW
 
 | |
4NXT
 
 | | Crystal structure of the cytosolic domain of human MiD51 | | Descriptor: | GLYCEROL, Mitochondrial dynamic protein MID51, SULFATE ION | | Authors: | Richter, V, Ryan, M.T, Kvansakul, M. | | Deposit date: | 2013-12-09 | | Release date: | 2013-12-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Structural and functional analysis of MiD51, a dynamin receptor required for mitochondrial fission.
J.Cell Biol., 204, 2014
|
|
4NXV
 
 | | Crystal structure of the cytosolic domain of human MiD51 | | Descriptor: | GLYCEROL, GUANOSINE-5'-DIPHOSPHATE, Mitochondrial dynamic protein MID51, ... | | Authors: | Richter, V, Kvansakul, M, Ryan, M.T. | | Deposit date: | 2013-12-09 | | Release date: | 2013-12-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and functional analysis of MiD51, a dynamin receptor required for mitochondrial fission.
J.Cell Biol., 204, 2014
|
|
