1MKY
 
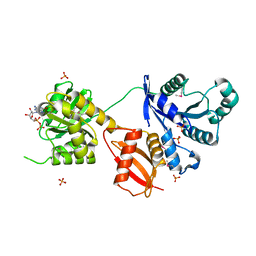 | | Structural Analysis of the Domain Interactions in Der, a Switch Protein Containing Two GTPase Domains | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, PHOSPHATE ION, Probable GTP-binding protein engA | | Authors: | Robinson, V.L, Hwang, J, Fox, E, Inouye, M, Stock, A.M. | | Deposit date: | 2002-08-29 | | Release date: | 2003-01-14 | | Last modified: | 2015-02-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Domain Arrangement of Der, a Switch Protein Containing Two GTPase Domains
Structure, 10, 2002
|
|
1NS6
 
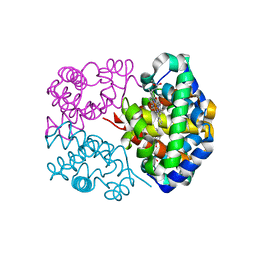 | |
1NS9
 
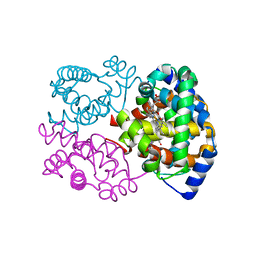 | |
1P2F
 
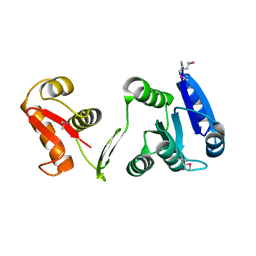 | |
3NNS
 
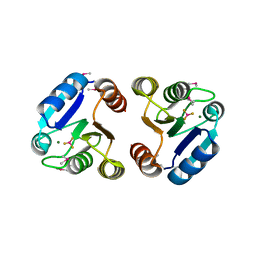 | | BeF3 Activated DrrB Receiver Domain | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, DNA BINDING RESPONSE REGULATOR B, MAGNESIUM ION | | Authors: | Robinson, V.L, Stock, A.M. | | Deposit date: | 2010-06-24 | | Release date: | 2010-08-11 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Regulation of response regulator autophosphorylation through interdomain contacts.
J.Biol.Chem., 285, 2010
|
|
3NNN
 
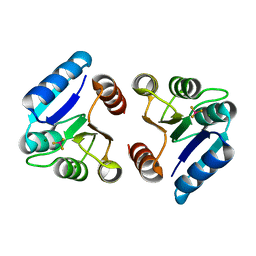 | | BeF3 Activated DrrD Receiver Domain | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, DNA BINDING RESPONSE REGULATOR D, MAGNESIUM ION | | Authors: | Robinson, V.L, Stock, A.M. | | Deposit date: | 2010-06-23 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Regulation of response regulator autophosphorylation through interdomain contacts.
J.Biol.Chem., 285, 2010
|
|
8EWH
 
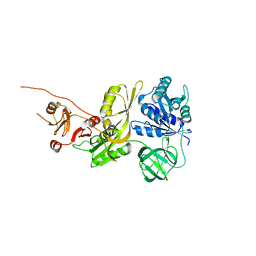 | | Salmonella typhimurium GTPase BIPA | | Descriptor: | 50S ribosomal subunit assembly factor BipA, SODIUM ION | | Authors: | Brown, R.S, deLivron, M.A, Robinson, V.L. | | Deposit date: | 2022-10-23 | | Release date: | 2022-11-16 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Crystallographic and Biochemical Characterization of the GTPase and Ribosome Binding Properties of Salmonella typhimuirum BipA
J Biomol Struct Dyn., 24:6, 2007
|
|
6D3K
 
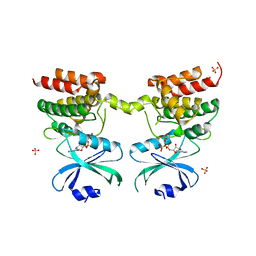 | | Crystal structure of unphosphorylated human PKR kinase domain in complex with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Interferon-induced, double-stranded RNA-activated protein kinase, ... | | Authors: | Erlandsen, H, Mayo, C.B, Robinson, V.L, Cole, J.L. | | Deposit date: | 2018-04-16 | | Release date: | 2019-07-10 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural Basis of Protein Kinase R Autophosphorylation.
Biochemistry, 58, 2019
|
|
6D3L
 
 | | Crystal structure of unphosphorylated human PKR | | Descriptor: | Interferon-induced, double-stranded RNA-activated protein kinase | | Authors: | Erlandsen, H, Mayo, C, Robinson, V.L, Cole, J.L. | | Deposit date: | 2018-04-16 | | Release date: | 2019-07-10 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural Basis of Protein Kinase R Autophosphorylation.
Biochemistry, 58, 2019
|
|
2FLK
 
 | | Crystal structure of CheY in complex with CheZ(200-214) solved from a F432 crystal grown in CAPS (pH 10.5) | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, C-terminal 15-mer from Chemotaxis protein cheZ, Chemotaxis protein cheY, ... | | Authors: | Guhaniyogi, J, Robinson, V.L, Stock, A.M. | | Deposit date: | 2006-01-06 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structures of Beryllium Fluoride-free and Beryllium Fluoride-bound CheY in Complex with the Conserved C-terminal Peptide of CheZ Reveal Dual Binding Modes Specific to CheY Conformation.
J.Mol.Biol., 359, 2006
|
|
2FMF
 
 | |
2FMK
 
 | | Crystal structure of Mg2+ and BeF3- bound CheY in complex with CheZ 200-214 solved from a P2(1)2(1)2 crystal grown in MES (pH 6.0) | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, C-terminal 15-mer from Chemotaxis protein cheZ, Chemotaxis protein cheY, ... | | Authors: | Guhaniyogi, J, Robinson, V.L, Stock, A.M. | | Deposit date: | 2006-01-09 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.999 Å) | | Cite: | Crystal Structures of Beryllium Fluoride-free and Beryllium Fluoride-bound CheY in Complex with the Conserved C-terminal Peptide of CheZ Reveal Dual Binding Modes Specific to CheY Conformation
J.Mol.Biol., 359, 2006
|
|
2FMH
 
 | | Crystal structure of Mg2+ and BeF3- bound CheY in complex with CheZ 200-214 solved from a F432 crystal grown in Tris (pH 8.4) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, BERYLLIUM TRIFLUORIDE ION, C-terminal 15-mer from Chemotaxis protein cheZ, ... | | Authors: | Guhaniyogi, J, Robinson, V.L, Stock, A.M. | | Deposit date: | 2006-01-09 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Crystal Structures of Beryllium Fluoride-free and Beryllium Fluoride-bound CheY in Complex with the Conserved C-terminal Peptide of CheZ Reveal Dual Binding Modes Specific to CheY Conformation.
J.Mol.Biol., 359, 2006
|
|
2FKA
 
 | | Crystal structure of Mg(2+) and BeF(3)(-)-bound CheY in complex with CheZ(200-214) solved from a F432 crystal grown in CAPS (pH 10.5) | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, BERYLLIUM TRIFLUORIDE ION, C-terminal 15-mer from Chemotaxis protein cheZ, ... | | Authors: | Guhaniyogi, J, Robinson, V.L, Stock, A.M. | | Deposit date: | 2006-01-04 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structures of Beryllium Fluoride-free and Beryllium Fluoride-bound CheY in Complex with the Conserved C-terminal Peptide of CheZ Reveal Dual Binding Modes Specific to CheY Conformation.
J.Mol.Biol., 359, 2006
|
|
2FLW
 
 | | Crystal structure of Mg2+ and BeF3- ound CheY in complex with CheZ 200-214 solved from a F432 crystal grown in Hepes (pH 7.5) | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, C-terminal 15-mer from Chemotaxis protein cheZ, Chemotaxis protein cheY, ... | | Authors: | Guhaniyogi, J, Robinson, V.L, Stock, A.M. | | Deposit date: | 2006-01-06 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structures of Beryllium Fluoride-free and Beryllium Fluoride-bound CheY in Complex with the Conserved C-terminal Peptide of CheZ Reveal Dual Binding Modes Specific to CheY Conformation.
J.Mol.Biol., 359, 2006
|
|
2FMI
 
 | | Crystal structure of CheY in complex with CheZ 200-214 solved from a F432 crystal grown in Tris (pH 8.4) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, C-terminal 15-mer from Chemotaxis protein cheZ, Chemotaxis protein cheY, ... | | Authors: | Guhaniyogi, J, Robinson, V.L, Stock, A.M. | | Deposit date: | 2006-01-09 | | Release date: | 2006-05-23 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.302 Å) | | Cite: | Crystal Structures of Beryllium Fluoride-free and Beryllium Fluoride-bound CheY in Complex with the Conserved C-terminal Peptide of CheZ Reveal Dual Binding Modes Specific to CheY Conformation.
J.Mol.Biol., 359, 2006
|
|
3NHZ
 
 | | Structure of N-terminal Domain of MtrA | | Descriptor: | MAGNESIUM ION, Two component system transcriptional regulator mtrA | | Authors: | Barbieri, C.M, Mack, T.R, Robinson, V.L, Miller, M.T, Stock, A.M. | | Deposit date: | 2010-06-14 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Regulation of response regulator autophosphorylation through interdomain contacts.
J.Biol.Chem., 285, 2010
|
|
6NPF
 
 | | Structure of E.coli enolase in complex with an analog of the natural product SF-2312 metabolite. | | Descriptor: | Enolase, GLYCEROL, L(+)-TARTARIC ACID, ... | | Authors: | Erlandsen, H, Krucinska, J, Lombardo, M, Wright, D. | | Deposit date: | 2019-01-17 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | Functional and structural basis of E. coli enolase inhibition by SF2312: a mimic of the carbanion intermediate.
Sci Rep, 9, 2019
|
|
6BFZ
 
 | | Crystal structure of enolase from E. coli with a mixture of apo form, substrate, and product form | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-PHOSPHOGLYCERIC ACID, Enolase, ... | | Authors: | Erlandsen, H, Wright, D, Krucinska, J. | | Deposit date: | 2017-10-27 | | Release date: | 2018-10-31 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | Structural and Functional Studies of Bacterial Enolase, a Potential Target against Gram-Negative Pathogens.
Biochemistry, 58, 2019
|
|
6BFY
 
 | | Crystal structure of enolase from Escherichia coli with bound 2-phosphoglycerate substrate | | Descriptor: | 2-PHOSPHOGLYCERIC ACID, Enolase, GLYCEROL, ... | | Authors: | Erlandsen, H, Wright, D, Krucinska, J. | | Deposit date: | 2017-10-27 | | Release date: | 2018-10-31 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Structural and Functional Studies of Bacterial Enolase, a Potential Target against Gram-Negative Pathogens.
Biochemistry, 58, 2019
|
|
2LPM
 
 | |
6D3Q
 
 | | Crystal structure of Escherichia coli enolase complexed with a natural inhibitor SF2312. | | Descriptor: | Enolase, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Erlandsen, H, Krucinska, J, Hazeen, A, Wright, D. | | Deposit date: | 2018-04-16 | | Release date: | 2019-11-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Functional and structural basis of E. coli enolase inhibition by SF2312: a mimic of the carbanion intermediate.
Sci Rep, 9, 2019
|
|
