4MX1
 
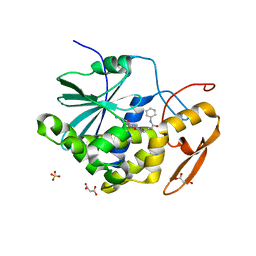 | | Structure of ricin A chain bound with 2-amino-4-oxo-N-(2-(3-phenylureido)ethyl)-3,4-dihydropteridine-7-carboxamide | | Descriptor: | 2-amino-4-oxo-N-{2-[(phenylcarbamoyl)amino]ethyl}-3,4-dihydropteridine-7-carboxamide, MALONIC ACID, Ricin A chain, ... | | Authors: | Robertus, J.D, Wiget, P.A, Manzano, L.A, Pruet, J.M, Gao, G, Saito, R, Jasheway, K.R, Monzingo, A.F, Anslyn, E.V. | | Deposit date: | 2013-09-25 | | Release date: | 2014-01-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Sulfur incorporation generally improves Ricin inhibition in pterin-appended glycine-phenylalanine dipeptide mimics.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
4HV3
 
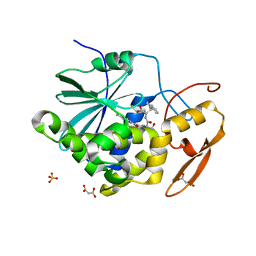 | | Structure of Ricin A chain bound with N-(N-(pterin-7-yl)carbonyl-L-serinyl)-L-tryptophan | | Descriptor: | (2S)-2-[[(2S)-2-[(2-azanyl-4-oxidanylidene-1H-pteridin-7-yl)carbonylamino]-3-oxidanyl-propanoyl]amino]-3-(1H-indol-3-yl)propanoic acid, MALONIC ACID, Ricin, ... | | Authors: | Robertus, J.D, Manzano, L.A, Jasheway, K.R, Monzingo, A.F, Saito, R, Pruet, J.M, Wiget, P.A, Anslyn, E.V. | | Deposit date: | 2012-11-05 | | Release date: | 2012-12-26 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Peptide-conjugated pterins as inhibitors of ricin toxin A.
J.Med.Chem., 56, 2013
|
|
2BAA
 
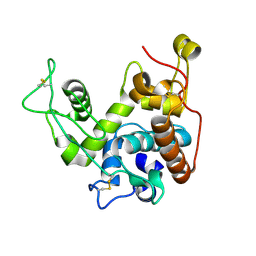 | | THE REFINED CRYSTAL STRUCTURE OF AN ENDOCHITINASE FROM HORDEUM VULGARE L. SEEDS TO 1.8 ANGSTROMS RESOLUTION | | Descriptor: | ENDOCHITINASE (26 KD) | | Authors: | Hart, P.J, Pfluger, H.D, Monzingo, A.F, Ready, M.P, Ernst, S.R, Hollis, T, Robertus, J.D. | | Deposit date: | 1995-01-26 | | Release date: | 1996-01-15 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The refined crystal structure of an endochitinase from Hordeum vulgare L. seeds at 1.8 A resolution.
J.Mol.Biol., 248, 1995
|
|
3BPB
 
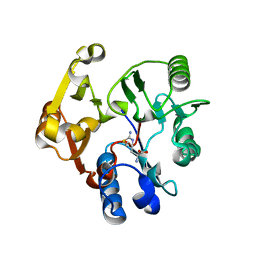 | | Crystal structure of the dimethylarginine dimethylaminohydrolase H162G adduct with S-methyl-L-thiocitrulline | | Descriptor: | N~5~-[(E)-imino(methylsulfanyl)methyl]-L-ornithine, dimethylarginine dimethylaminohydrolase | | Authors: | Monzingo, A.F, Linsky, T.W, Stone, E.M, Fast, W, Robertus, J.D. | | Deposit date: | 2007-12-18 | | Release date: | 2008-06-17 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | Promiscuous partitioning of a covalent intermediate common in the pentein superfamily.
Chem.Biol., 15, 2008
|
|
4QQU
 
 | | Crystal structure of the cobalamin-independent methionine synthase enzyme in a closed conformation | | Descriptor: | 2-AMINO-4-MERCAPTO-BUTYRIC ACID, 5-methyltetrahydropteroyltriglutamate--homocysteine methyltransferase, N-[4-({[(6S)-2-amino-4-oxo-3,4,5,6,7,8-hexahydropteridin-6-yl]methyl}amino)benzoyl]-L-gamma-glutamyl-L-gamma-glutamyl-L-glutamic acid, ... | | Authors: | Ubhi, D.K, Robertus, J.D. | | Deposit date: | 2014-06-29 | | Release date: | 2015-01-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.98 Å) | | Cite: | The cobalamin-independent methionine synthase enzyme captured in a substrate-induced closed conformation.
J.Mol.Biol., 427, 2015
|
|
3I2E
 
 | | Crystal structure of human dimethylarginine dymethylaminohydrolase-1 (DDAH-1) | | Descriptor: | N(G),N(G)-dimethylarginine dimethylaminohydrolase 1 | | Authors: | Monzingo, A.F, Wang, Y, Hu, S, Schaller, T.H, Robertus, J.D, Fast, W. | | Deposit date: | 2009-06-29 | | Release date: | 2009-08-25 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Developing dual and specific inhibitors of dimethylarginine dimethylaminohydrolase-1 and nitric oxide synthase: toward a targeted polypharmacology to control nitric oxide.
Biochemistry, 48, 2009
|
|
3I4A
 
 | | Crystal structure of dimethylarginine dimethylaminohydrolase-1 (DDAH-1) in complex with N5-(1-iminopropyl)-L-ornithine | | Descriptor: | N(G),N(G)-dimethylarginine dimethylaminohydrolase 1, N5-(1-iminopropyl)-L-ornithine | | Authors: | Monzingo, A.F, Wang, Y, Hu, S, Schaller, T.H, Fast, W, Robertus, J.D. | | Deposit date: | 2009-07-01 | | Release date: | 2009-08-25 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.898 Å) | | Cite: | Developing dual and specific inhibitors of dimethylarginine dimethylaminohydrolase-1 and nitric oxide synthase: toward a targeted polypharmacology to control nitric oxide.
Biochemistry, 48, 2009
|
|
1EDZ
 
 | | STRUCTURE OF THE NAD-DEPENDENT 5,10-METHYLENETETRAHYDROFOLATE DEHYDROGENASE FROM SACCHAROMYCES CEREVISIAE | | Descriptor: | 5,10-METHYLENETETRAHYDROFOLATE DEHYDROGENASE | | Authors: | Monzingo, A.F, Breksa, A, Ernst, S, Appling, D.R, Robertus, J.D. | | Deposit date: | 2000-01-28 | | Release date: | 2000-12-06 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The X-ray structure of the NAD-dependent 5,10-methylenetetrahydrofolate dehydrogenase from Saccharomyces cerevisiae.
Protein Sci., 9, 2000
|
|
1EE9
 
 | | CRYSTAL STRUCTURE OF THE NAD-DEPENDENT 5,10-METHYLENETETRAHYDROFOLATE DEHYDROGENASE FROM SACCHAROMYCES CEREVISIAE COMPLEXED WITH NAD | | Descriptor: | 5,10-METHYLENETETRAHYDROFOLATE DEHYDROGENASE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Monzingo, A.F, Breksa, A, Ernst, S, Appling, D.R, Robertus, J.D. | | Deposit date: | 2000-01-31 | | Release date: | 2000-12-06 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | The X-ray structure of the NAD-dependent 5,10-methylenetetrahydrofolate dehydrogenase from Saccharomyces cerevisiae.
Protein Sci., 9, 2000
|
|
1HQ6
 
 | | STRUCTURE OF PYRUVOYL-DEPENDENT HISTIDINE DECARBOXYLASE AT PH 8 | | Descriptor: | HISTIDINE DECARBOXYLASE | | Authors: | Schelp, E, Worley, S, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2000-12-14 | | Release date: | 2001-03-21 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | pH-induced structural changes regulate histidine decarboxylase activity in Lactobacillus 30a.
J.Mol.Biol., 306, 2001
|
|
1HWN
 
 | | EBULIN COMPLEXED WITH GALACTOSE, TRIGONAL CRYSTAL FORM | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, EBULIN, beta-D-galactopyranose | | Authors: | Pascal, J.M, Day, P.J, Monzingo, A.F, Ernst, S.R, Robertus, J.D. | | Deposit date: | 2001-01-09 | | Release date: | 2001-01-24 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | 2.8-A crystal structure of a nontoxic type-II ribosome-inactivating protein, ebulin l.
Proteins, 43, 2001
|
|
1HWO
 
 | | EBULIN COMPLEXED WITH LACTOSE, TRIGONAL CRYSTAL FORM | | Descriptor: | EBULIN, beta-D-galactopyranose-(1-4)-alpha-D-glucopyranose, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Pascal, J.M, Day, P.J, Monzingo, A.F, Ernst, S.R, Robertus, J.D. | | Deposit date: | 2001-01-09 | | Release date: | 2001-01-24 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | 2.8-A crystal structure of a nontoxic type-II ribosome-inactivating protein, ebulin l.
Proteins, 43, 2001
|
|
1IBT
 
 | | STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE AT-170 C | | Descriptor: | HISTIDINE DECARBOXYLASE ALPHA CHAIN, HISTIDINE DECARBOXYLASE BETA CHAIN | | Authors: | Worley, S, Schelp, E, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2001-03-29 | | Release date: | 2002-03-13 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure and cooperativity of a T-state mutant of histidine decarboxylase from Lactobacillus 30a.
Proteins, 46, 2002
|
|
1IBW
 
 | | STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE BOUND WITH HISTIDINE METHYL ESTER AT 25 C | | Descriptor: | HISTIDINE DECARBOXYLASE BETA CHAIN, HISTIDINE-METHYL-ESTER, Histidine decarboxylase alpha chain | | Authors: | Worley, S, Schelp, E, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2001-03-29 | | Release date: | 2002-03-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structure and cooperativity of a T-state mutant of histidine decarboxylase from Lactobacillus 30a.
Proteins, 46, 2002
|
|
1IBU
 
 | | STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE AT 25 C | | Descriptor: | HISTIDINE DECARBOXYLASE ALPHA CHAIN, HISTIDINE DECARBOXYLASE BETA CHAIN | | Authors: | Worley, S, Schelp, E, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2001-03-29 | | Release date: | 2002-03-13 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structure and cooperativity of a T-state mutant of histidine decarboxylase from Lactobacillus 30a.
Proteins, 46, 2002
|
|
1IBV
 
 | | STRUCTURE OF THE D53,54N MUTANT OF HISTIDINE DECARBOXYLASE BOUND WITH HISTIDINE METHYL ESTER AT-170 C | | Descriptor: | HISTIDINE DECARBOXYLASE BETA CHAIN, HISTIDINE-METHYL-ESTER, Histidine decarboxylase alpha chain | | Authors: | Worley, S, Schelp, E, Monzingo, A.F, Ernst, S, Robertus, J.D. | | Deposit date: | 2001-03-29 | | Release date: | 2002-03-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure and cooperativity of a T-state mutant of histidine decarboxylase from Lactobacillus 30a.
Proteins, 46, 2002
|
|
2AAI
 
 | | Crystallographic refinement of ricin to 2.5 Angstroms | | Descriptor: | RICIN (A CHAIN), RICIN (B CHAIN), alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Rutenber, E, Katzin, B.J, Montfort, W, Villafranca, J.E, Ernst, S.R, Collins, E.J, Mlsna, D, Monzingo, A.F, Ready, M.P, Robertus, J.D. | | Deposit date: | 1993-09-07 | | Release date: | 1994-01-31 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystallographic refinement of ricin to 2.5 A.
Proteins, 10, 1991
|
|
1SBT
 
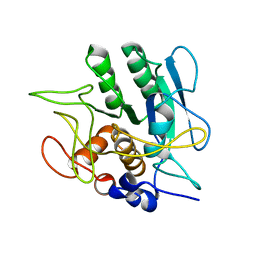 | | ATOMIC COORDINATES FOR SUBTILISIN BPN (OR NOVO) | | Descriptor: | SUBTILISIN BPN' | | Authors: | Alden, R.A, Birktoft, J.J, Kraut, J, Robertus, J.D, Wright, C.S. | | Deposit date: | 1972-08-11 | | Release date: | 1977-01-06 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Atomic coordinates for subtilisin BPN' (or Novo).
Biochem.Biophys.Res.Commun., 45, 1971
|
|
1RTC
 
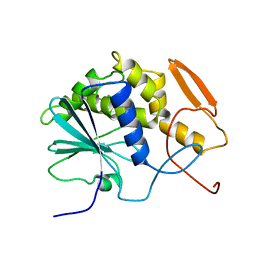 | | THE STRUCTURE OF RECOMBINANT RICIN A CHAIN AT 2.3 ANGSTROMS | | Descriptor: | RICIN | | Authors: | Mlsna, D, Monzingo, A.F, Katzin, B.J, Ernst, S, Robertus, J.D. | | Deposit date: | 1992-10-29 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of recombinant ricin A chain at 2.3 A.
Protein Sci., 2, 1993
|
|
1DU5
 
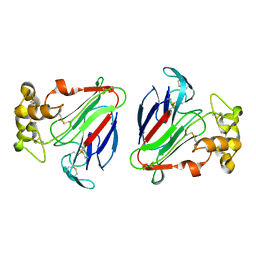 | | THE CRYSTAL STRUCTURE OF ZEAMATIN. | | Descriptor: | ZEAMATIN | | Authors: | Batalia, M.A, Monzingo, A.F, Ernst, S, Roberts, W, Robertus, J.D. | | Deposit date: | 2000-01-14 | | Release date: | 2000-02-02 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The crystal structure of the antifungal protein zeamatin, a member of the thaumatin-like, PR-5 protein family.
Nat.Struct.Biol., 3, 1996
|
|
1CHG
 
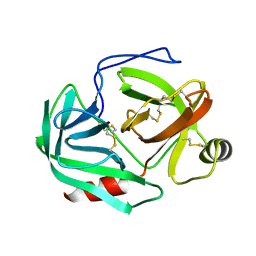 | | CHYMOTRYPSINOGEN,2.5 ANGSTROMS CRYSTAL STRUCTURE, COMPARISON WITH ALPHA-CHYMOTRYPSIN,AND IMPLICATIONS FOR ZYMOGEN ACTIVATION | | Descriptor: | CHYMOTRYPSINOGEN A | | Authors: | Freer, S.T, Kraut, J, Robertus, J.D, Wright, H.T, Xuong, N.H. | | Deposit date: | 1975-03-01 | | Release date: | 1976-11-22 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Chymotrypsinogen: 2.5-angstrom crystal structure, comparison with alpha-chymotrypsin, and implications for zymogen activation.
Biochemistry, 9, 1970
|
|
1HWP
 
 | | EBULIN COMPLEXED WITH PTEROIC ACID, TRIGONAL CRYSTAL FORM | | Descriptor: | EBULIN, PTEROIC ACID, beta-D-galactopyranose-(1-4)-beta-D-glucopyranose, ... | | Authors: | Pascal, J.M, Day, P.J, Monzingo, A.F, Ernst, S.R, Robertus, J.D. | | Deposit date: | 2001-01-09 | | Release date: | 2001-01-24 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | 2.8-A crystal structure of a nontoxic type-II ribosome-inactivating protein, ebulin l.
Proteins, 43, 2001
|
|
1HWM
 
 | | EBULIN,ORTHORHOMBIC CRYSTAL FORM MODEL | | Descriptor: | EBULIN, alpha-D-mannopyranose-(1-3)-[beta-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, beta-D-galactopyranose | | Authors: | Pascal, J.M, Day, P.J, Monzingo, A.F, Ernst, S.R, Robertus, J.D. | | Deposit date: | 2001-01-09 | | Release date: | 2001-01-24 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | 2.8-A crystal structure of a nontoxic type-II ribosome-inactivating protein, ebulin l.
Proteins, 43, 2001
|
|
3P8P
 
 | | Crystal Structure of Human Dimethylarginine Dimethylaminohydrolase-1 (DDAH-1) variant C274S bound with N5-(1-iminopentyl)-L-ornithine | | Descriptor: | N(G),N(G)-dimethylarginine dimethylaminohydrolase 1, N~5~-[(1E)-pentanimidoyl]-L-ornithine | | Authors: | Monzingo, A.F, Lluis, M, Wang, Y, Fast, W, Robertus, J.D. | | Deposit date: | 2010-10-14 | | Release date: | 2010-11-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Characterization of C-Alkyl Amidines as Bioavailable Covalent Reversible Inhibitors of Human DDAH-1.
Chemmedchem, 6, 2011
|
|
3P8E
 
 | | Crystal structure of human DIMETHYLARGININE DIMETHYLAMINOHYDROLASE-1 (DDAH-1) covalently bound with N5-(1-iminopentyl)-L-ornithine | | Descriptor: | N(G),N(G)-dimethylarginine dimethylaminohydrolase 1, N~5~-[(1S)-1-aminopentyl]-L-ornithine | | Authors: | Lluis, M, Wang, Y, Monzingo, A.F, Fast, W, Robertus, J.D. | | Deposit date: | 2010-10-13 | | Release date: | 2010-11-10 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.4946 Å) | | Cite: | Characterization of C-Alkyl Amidines as Bioavailable Covalent Reversible Inhibitors of Human DDAH-1.
Chemmedchem, 6, 2011
|
|
