5ZKA
 
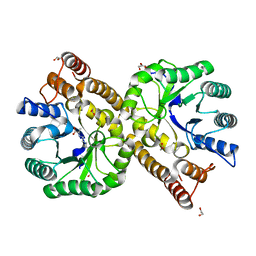 | | Crystal structure of N-acetylneuraminate lyase from Fusobacterium nucleatum complexed with Pyruvate | | Descriptor: | 1,2-ETHANEDIOL, N-acetylneuraminate lyase, TRIETHYLENE GLYCOL | | Authors: | Kumar, J.P, Rao, H, Nayak, V, Ramaswamy, S. | | Deposit date: | 2018-03-23 | | Release date: | 2018-12-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Crystal structures and kinetics of N-acetylneuraminate lyase from Fusobacterium nucleatum
Acta Crystallogr F Struct Biol Commun, 74, 2018
|
|
2KYQ
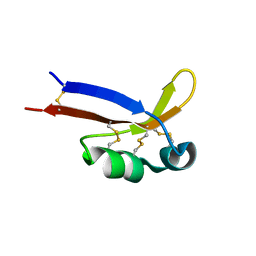 
 | | 1H, 15N, 13C chemical shifts and structure of CKR-brazzein | | Descriptor: | Defensin-like protein | | Authors: | Dittli, S.M, Assadi-Porter, F.M, Rao, H, Tonelli, M. | | Deposit date: | 2010-06-07 | | Release date: | 2011-05-18 | | Last modified: | 2011-11-30 | | Method: | SOLUTION NMR | | Cite: | Structural role of the terminal disulfide bond in the sweetness of brazzein.
Chem Senses, 36, 2011
|
|
2L07
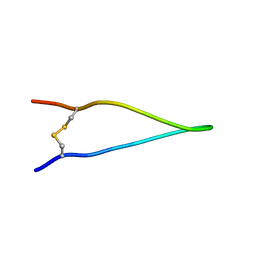 
 | | 1H, 13C, and 15N chemical shifts and structure of brazzein-derived peptide CKR-PNG | | Descriptor: | Defensin-like protein | | Authors: | Dittli, S.M, Assadi-Porter, F.M, Rao, H, Tonelli, M. | | Deposit date: | 2010-06-30 | | Release date: | 2011-06-22 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural role of the terminal disulfide bond in the sweetness of brazzein.
Chem Senses, 36, 2011
|
|
5ZJM
 
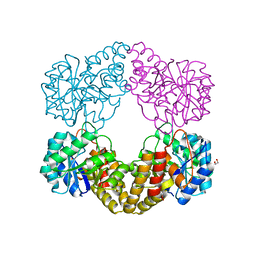 | | Crystal structure of N-acetylneuraminate lyase from Fusobacterium nucleatum | | Descriptor: | 1,2-ETHANEDIOL, N-acetylneuraminate lyase | | Authors: | Kumar, J.P, Rao, H, Nayak, V, Subramanian, R. | | Deposit date: | 2018-03-21 | | Release date: | 2019-01-23 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.323 Å) | | Cite: | Crystal structures and kinetics of N-acetylneuraminate lyase from Fusobacterium nucleatum
Acta Crystallogr F Struct Biol Commun, 74, 2018
|
|
4ZXO
 
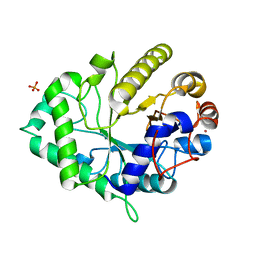 | | The structure of a GH26 beta-mannanase from Bacteroides ovatus, BoMan26A. | | Descriptor: | Glycosyl hydrolase family 26, PHOSPHATE ION, POTASSIUM ION | | Authors: | Bagenholm, V, Aurelius, O, Logan, D.T, Bouraoui, H, Stalbrand, H. | | Deposit date: | 2015-05-20 | | Release date: | 2016-06-29 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Galactomannan Catabolism Conferred by a Polysaccharide Utilization Locus of Bacteroides ovatus: ENZYME SYNERGY AND CRYSTAL STRUCTURE OF A beta-MANNANASE.
J. Biol. Chem., 292, 2017
|
|
8KHD
 
 | | The interface structure of Omicron RBD binding to 5817 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of 5817, Light chain of 5817, ... | | Authors: | Cao, L, Wang, X. | | Deposit date: | 2023-08-21 | | Release date: | 2024-04-17 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Identification of a broad sarbecovirus neutralizing antibody targeting a conserved epitope on the receptor-binding domain.
Cell Rep, 43, 2024
|
|
8KHC
 
 | | SARS-CoV-2 Omicron spike in complex with 5817 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of 5817 Fab, ... | | Authors: | Cao, L, Wang, X. | | Deposit date: | 2023-08-21 | | Release date: | 2024-04-17 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Identification of a broad sarbecovirus neutralizing antibody targeting a conserved epitope on the receptor-binding domain.
Cell Rep, 43, 2024
|
|
2LY6
 
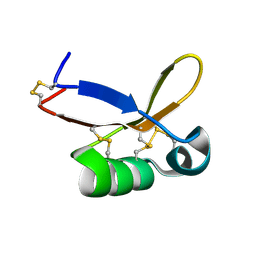 | | Refined solution structure of recombinant brazzein at low temperature | | Descriptor: | Defensin-like protein | | Authors: | Cornilescu, C.C, Cornilescu, G, Tonelli, M, Markley, J.L, Assadi-Porter, F.M. | | Deposit date: | 2012-09-12 | | Release date: | 2012-10-17 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Temperature-dependent conformational change affecting Tyr11 and sweetness loops of brazzein.
Proteins, 81, 2013
|
|
2LY5
 
 | | Refined solution structure of recombinant brazzein | | Descriptor: | Defensin-like protein | | Authors: | Cornilescu, C.C, Cornilescu, G, Tonelli, M, Markley, J.L, Assadi-Porter, F.M. | | Deposit date: | 2012-09-12 | | Release date: | 2013-01-30 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Temperature-dependent conformational change affecting Tyr11 and sweetness loops of brazzein.
Proteins, 81, 2013
|
|
2SFA
 
 | | SERINE PROTEINASE FROM STREPTOMYCES FRADIAE ATCC 14544 | | Descriptor: | SERINE PROTEINASE | | Authors: | Kitadokoro, K, Tsuzuki, H. | | Deposit date: | 1994-04-25 | | Release date: | 1996-06-20 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Purification, characterization, primary structure, crystallization and preliminary crystallographic study of a serine proteinase from Streptomyces fradiae ATCC 14544.
Eur.J.Biochem., 220, 1994
|
|
5ZR3
 
 | | Crystal structure of Hsp90-alpha N-terminal domain in complex with 4-(3-isopropyl-4-(4-(1-methyl-1H-pyrazol-4-yl)-1H-imidazol-1-yl)-1H-pyrazolo[3,4-b]pyridin-1-yl)-3-methylbenzamide | | Descriptor: | 3-methyl-4-{4-[4-(1-methyl-1H-pyrazol-4-yl)-1H-imidazol-1-yl]-3-(propan-2-yl)-1H-pyrazolo[3,4-b]pyridin-1-yl}benzamide, Heat shock protein HSP 90-alpha | | Authors: | Uno, T, Chong, K.T, Suzuki, T. | | Deposit date: | 2018-04-23 | | Release date: | 2019-01-02 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Discovery of 3-Ethyl-4-(3-isopropyl-4-(4-(1-methyl-1 H-pyrazol-4-yl)-1 H-imidazol-1-yl)-1 H-pyrazolo[3,4- b]pyridin-1-yl)benzamide (TAS-116) as a Potent, Selective, and Orally Available HSP90 Inhibitor.
J. Med. Chem., 62, 2019
|
|
7CYV
 
 | | Crystal structure of FD20, a neutralizing single-chain variable fragment (scFv) in complex with SARS-CoV-2 Spike receptor-binding domain (RBD) | | Descriptor: | Spike protein S1, The heavy chain variable region of the scFv FD20,The light chain variable region of the scFv FD20, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-3)][alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Li, Y, Li, T, Lai, Y, Cai, H, Yao, H, Li, D. | | Deposit date: | 2020-09-04 | | Release date: | 2021-09-15 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.13 Å) | | Cite: | Uncovering a conserved vulnerability site in SARS-CoV-2 by a human antibody.
Embo Mol Med, 13, 2021
|
|
7C8V
 
 | | Structure of sybody SR4 in complex with the SARS-CoV-2 S Receptor Binding domain (RBD) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, Spike protein S1, ... | | Authors: | Li, T, Yao, H, Cai, H, Qin, W, Li, D. | | Deposit date: | 2020-06-03 | | Release date: | 2020-06-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | A synthetic nanobody targeting RBD protects hamsters from SARS-CoV-2 infection.
Nat Commun, 12, 2021
|
|
7C8W
 
 | | Structure of sybody MR17 in complex with the SARS-CoV-2 S receptor-binding domain (RBD) | | Descriptor: | GLYCEROL, Spike protein S1, Synthetic nanobody MR17, ... | | Authors: | Li, T, Cai, H, Yao, H, Qin, W, Li, D. | | Deposit date: | 2020-06-03 | | Release date: | 2020-06-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.77 Å) | | Cite: | A synthetic nanobody targeting RBD protects hamsters from SARS-CoV-2 infection.
Nat Commun, 12, 2021
|
|
7CAN
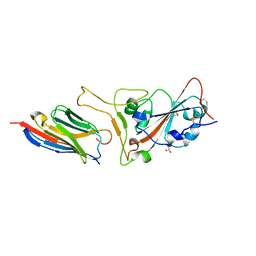 
 | | Structure of sybody MR17-K99Y in complex with the SARS-CoV-2 S Receptor-binding domain (RBD) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, Spike protein S1, ... | | Authors: | Li, T, Yao, H, Cai, H, Qin, W, Li, D. | | Deposit date: | 2020-06-09 | | Release date: | 2020-06-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.94 Å) | | Cite: | A synthetic nanobody targeting RBD protects hamsters from SARS-CoV-2 infection.
Nat Commun, 12, 2021
|
|
