2XQW
 
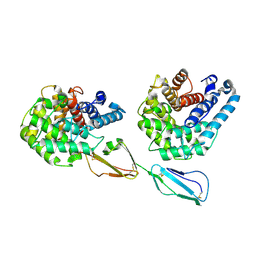 | | Structure of Factor H domains 19-20 in complex with complement C3d | | Descriptor: | COMPLEMENT C3, COMPLEMENT FACTOR H | | Authors: | Kajander, T, Lehtinen, M.J, Hyvarinen, S, Bhattacharjee, A, Leung, E, Isenman, D.E, Meri, S, Jokiranta, T.S, Goldman, A. | | Deposit date: | 2010-09-07 | | Release date: | 2011-02-02 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.306 Å) | | Cite: | Dual Interaction of Factor H with C3D and Glycosaminoglycans in Host-Nonhost Discrimination by Complement.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
1L1C
 
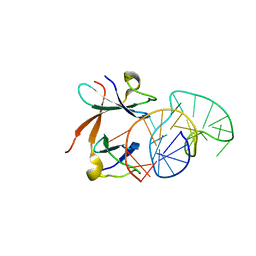 | | Structure of the LicT Bacterial Antiterminator Protein in Complex with its RNA Target | | Descriptor: | Transcription antiterminator licT, licT mRNA antiterminator hairpin | | Authors: | Yang, Y, Declerck, N, Manival, X, Aymerich, S, Kochoyan, M. | | Deposit date: | 2002-02-15 | | Release date: | 2002-03-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the LicT-RNA antitermination complex: CAT clamping RAT.
EMBO J., 21, 2002
|
|
6QXA
 
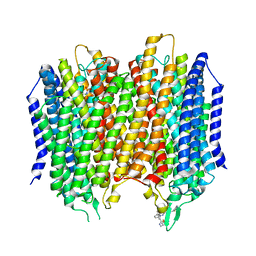 | | Structure of membrane bound pyrophosphatase from Thermotoga maritima in complex with imidodiphosphate and N-[(2-amino-6-benzothiazolyl)methyl]-1H-indole-2-carboxamide (ATC) | | Descriptor: | (4S,5S)-1,2-DITHIANE-4,5-DIOL, IMIDODIPHOSPHORIC ACID, K(+)-stimulated pyrophosphate-energized sodium pump, ... | | Authors: | Vidilaseris, K, Goldman, A. | | Deposit date: | 2019-03-07 | | Release date: | 2019-04-10 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (3.41 Å) | | Cite: | Asymmetry in catalysis byThermotoga maritimamembrane-bound pyrophosphatase demonstrated by a nonphosphorus allosteric inhibitor.
Sci Adv, 5, 2019
|
|
2G7I
 
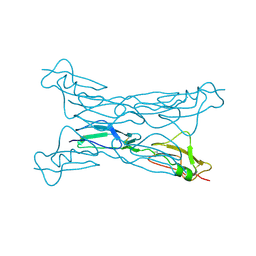 | |
6Y75
 
 | | BIL2 domain from T.thermophila BUBL1 locus (C1A-N143A) | | Descriptor: | DI(HYDROXYETHYL)ETHER, FORMIC ACID, NAD(P)(+)--arginine ADP-ribosyltransferase, ... | | Authors: | Ilari, A, Chiarini, V. | | Deposit date: | 2020-02-28 | | Release date: | 2021-02-03 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis of ubiquitination mediated by protein splicing in early Eukarya.
Biochim Biophys Acta Gen Subj, 1865, 2021
|
|
6TMM
 
 | | BIL2 domain from T.thermophila BUBL1 locus (C1A-N143A) | | Descriptor: | CALCIUM ION, DI(HYDROXYETHYL)ETHER, FORMIC ACID, ... | | Authors: | Chiarini, V, Ilari, A. | | Deposit date: | 2019-12-04 | | Release date: | 2020-12-16 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (2.398 Å) | | Cite: | Structural basis of ubiquitination mediated by protein splicing in early Eukarya.
Biochim Biophys Acta Gen Subj, 1865, 2021
|
|
3EII
 
 | | Zinc-bound glycoside hydrolase 61 E from Thielavia terrestris | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, SULFATE ION, ... | | Authors: | Salbo, R, Welner, D, Lo Leggio, L, Harris, P, McFarland, K. | | Deposit date: | 2008-09-16 | | Release date: | 2009-10-27 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Stimulation of lignocellulosic biomass hydrolysis by proteins of glycoside hydrolase family 61: structure and function of a large, enigmatic family.
Biochemistry, 49, 2010
|
|
3EJA
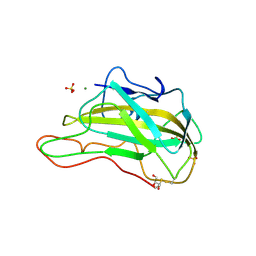 
 | | Magnesium-bound glycoside hydrolase 61 isoform E from Thielavia terrestris | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Salbo, R, Welner, D, Lo Leggio, L, Harris, P, McFarland, K. | | Deposit date: | 2008-09-18 | | Release date: | 2009-10-27 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.902 Å) | | Cite: | Stimulation of lignocellulosic biomass hydrolysis by proteins of glycoside hydrolase family 61: structure and function of a large, enigmatic family.
Biochemistry, 49, 2010
|
|
8CCN
 
 | | Filamentous actin II from Plasmodium falciparum | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Actin-2, MAGNESIUM ION | | Authors: | Kursula, I, Lopez, A.J. | | Deposit date: | 2023-01-27 | | Release date: | 2023-02-22 | | Last modified: | 2023-08-23 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structure and function of Plasmodium actin II in the parasite mosquito stages.
Plos Pathog., 19, 2023
|
|
8CCO
 
 | |
3N2S
 
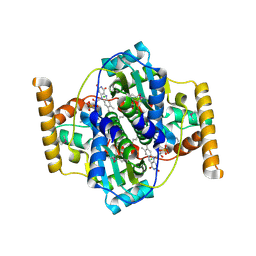 | | Structure of NfrA1 nitroreductase from B. subtilis | | Descriptor: | CHLORIDE ION, FLAVIN MONONUCLEOTIDE, NADPH-dependent nitro/flavin reductase | | Authors: | Morera, S, Gueguen-Chaignon, V, Meyer, P, Cortial, S, Ouazzani, J. | | Deposit date: | 2010-05-19 | | Release date: | 2010-09-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | NADH oxidase activity of Bacillus subtilis nitroreductase NfrA1: insight into its biological role.
Febs Lett., 584, 2010
|
|
3FWR
 
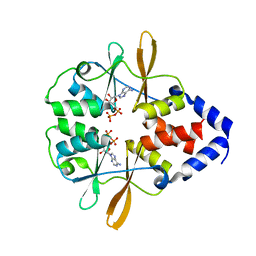 | |
3FWS
 
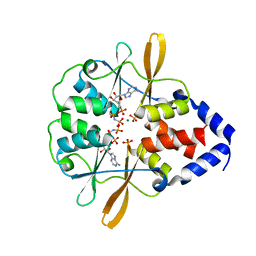 | | Crystal Structure of the CBS domains from the Bacillus subtilis CcpN repressor complexed with AppNp, phosphate and magnesium ions | | Descriptor: | MAGNESIUM ION, PHOSPHATE ION, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Chaix, D, Arold, S, Hoh, F, Declerck, N. | | Deposit date: | 2009-01-19 | | Release date: | 2010-01-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Ligand recognition by the energy sensor domain of the CcpN repressor
To be Published
|
|
3FV6
 
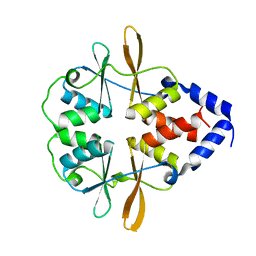 | |
6KHT
 
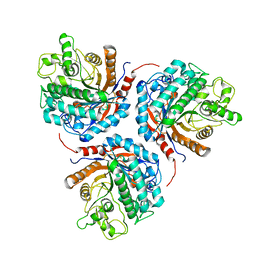 | | Chimeric beta-glucosidase Cel1b-H13 | | Descriptor: | Glycoside hydrolase family 1 | | Authors: | Niu, K.L. | | Deposit date: | 2019-07-16 | | Release date: | 2021-01-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.305 Å) | | Cite: | Chimeric beta-glucosidase Cel1b-H13
To Be Published
|
|
1AUU
 
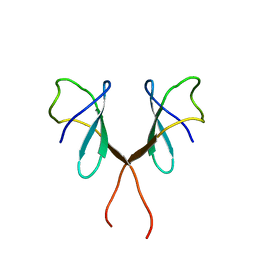 | |
4F7R
 
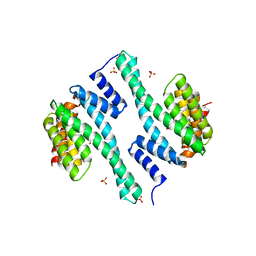 | |
4ZQ0
 
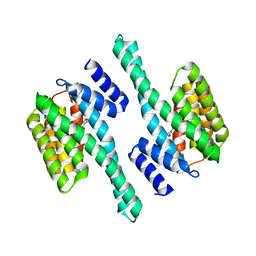 | |
2FCA
 
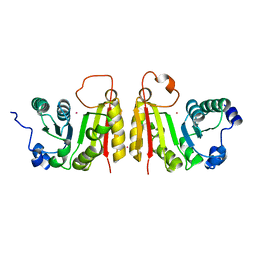 | | The structure of BsTrmB | | Descriptor: | POTASSIUM ION, tRNA (guanine-N(7)-)-methyltransferase | | Authors: | Zegers, I, Van Vliet, F, Bujnicki, J, Kosinski, J, Gigot, D, Droogmans, L. | | Deposit date: | 2005-12-12 | | Release date: | 2006-08-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of Bacillus subtilis TrmB, the tRNA (m7G46) methyltransferase.
Nucleic Acids Res., 34, 2006
|
|
