3KCH
 
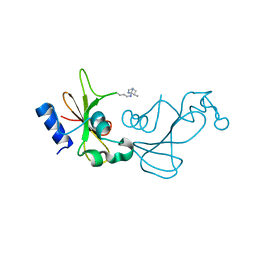 | | Baranase crosslinked by glutaraldehyde | | Descriptor: | PENTANEDIAL, Ribonuclease | | Authors: | Prange, T, Salem, M, Mauguen, Y. | | Deposit date: | 2009-10-21 | | Release date: | 2010-03-09 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Revisiting glutaraldehyde cross-linking: the case of the Arg-Lys intermolecular doublet
Acta Crystallogr.,Sect.F, 66, 2010
|
|
1BGS
 
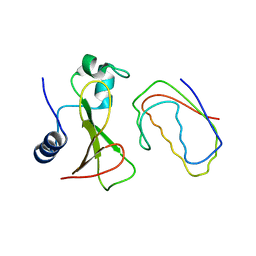 | | RECOGNITION BETWEEN A BACTERIAL RIBONUCLEASE, BARNASE, AND ITS NATURAL INHIBITOR, BARSTAR | | Descriptor: | BARNASE, BARSTAR | | Authors: | Guillet, V, Lapthorn, A, Mauguen, Y. | | Deposit date: | 1993-11-02 | | Release date: | 1994-04-30 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Recognition between a bacterial ribonuclease, barnase, and its natural inhibitor, barstar.
Structure, 1, 1993
|
|
2F2N
 
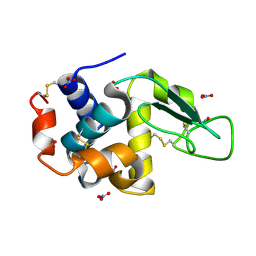 | | Triclinic hen egg lysozyme cross-linked by glutaraldehyde | | Descriptor: | Lysozyme C, NITRATE ION | | Authors: | Prange, T, Salem, M, Mauguen, Y. | | Deposit date: | 2005-11-17 | | Release date: | 2006-04-25 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.603 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
1A2P
 
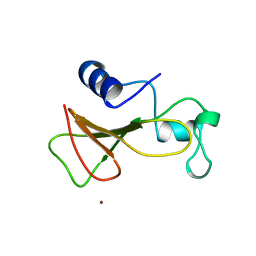 | | BARNASE WILDTYPE STRUCTURE AT 1.5 ANGSTROMS RESOLUTION | | Descriptor: | BARNASE, ZINC ION | | Authors: | Martin, C, Richard, V, Salem, M, Hartley, R.W, Mauguen, Y. | | Deposit date: | 1998-01-07 | | Release date: | 1998-04-29 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Refinement and structural analysis of barnase at 1.5 A resolution.
Acta Crystallogr.,Sect.D, 55, 1999
|
|
1B86
 
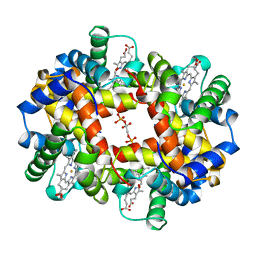 | | HUMAN DEOXYHAEMOGLOBIN-2,3-DIPHOSPHOGLYCERATE COMPLEX | | Descriptor: | (2R)-2,3-diphosphoglyceric acid, OXYGEN MOLECULE, PROTEIN (HEMOGLOBIN; ALPHA CHAIN), ... | | Authors: | Richard, V, Dodson, G.G, Mauguen, Y. | | Deposit date: | 1999-02-08 | | Release date: | 1999-02-14 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Human deoxyhaemoglobin-2,3-diphosphoglycerate complex low-salt structure at 2.5 A resolution.
J.Mol.Biol., 233, 1993
|
|
1RNB
 
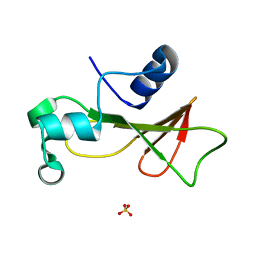 | |
1B27
 | |
1B3S
 | |
1B2S
 | |
1B2U
 | |
1BRS
 
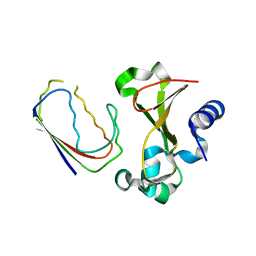 | |
1BNI
 
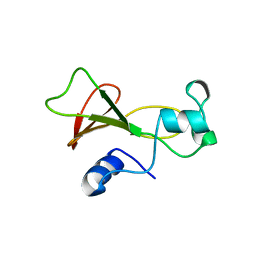 | | BARNASE WILDTYPE STRUCTURE AT PH 6.0 | | Descriptor: | BARNASE | | Authors: | Cameron, A, Henrick, K, Fersht, A.R, Dodson, G, Buckle, A.M. | | Deposit date: | 1995-05-17 | | Release date: | 1995-09-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structural analysis of mutations in the hydrophobic cores of barnase.
J.Mol.Biol., 234, 1993
|
|
2F4G
 
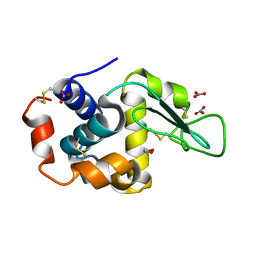 | | Triclinic cross-linked lysozyme soaked in bromoethanol 1M | | Descriptor: | 2-BROMOETHANOL, Lysozyme C, NITRATE ION | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-23 | | Release date: | 2006-04-25 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.654 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2F4Y
 
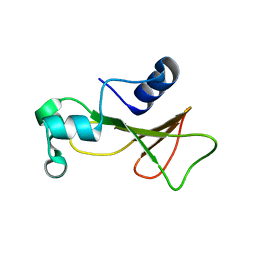 | | Barnase cross-linked with glutaraldehyde | | Descriptor: | Ribonuclease, ZINC ION | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-24 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2F56
 
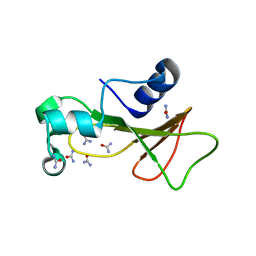 | | Barnase cross-linked with glutaraldehyde soaked in 6M urea | | Descriptor: | Ribonuclease, UREA, ZINC ION | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-25 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.955 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2F5M
 
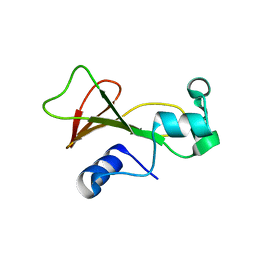 | | Cross-linked barnase soaked in bromo-ethanol | | Descriptor: | 2-BROMOETHANOL, Ribonuclease | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-26 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2F5W
 
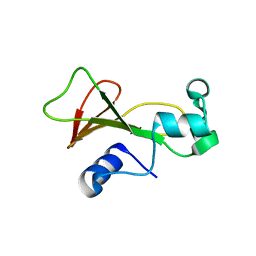 | | Cross-linked barnase soaked in 3 M thiourea | | Descriptor: | Ribonuclease, THIOUREA, ZINC ION | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-28 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2F30
 
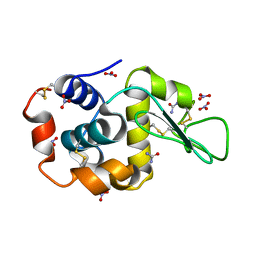 | | Triclinic cross-linked Lysozyme soaked with 4.5M urea | | Descriptor: | Lysozyme C, NITRATE ION, UREA | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-18 | | Release date: | 2006-04-25 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
2F4A
 
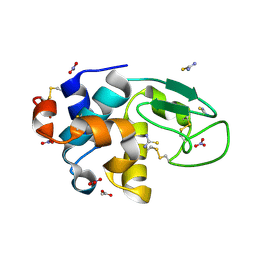 | | Triclinic cross-linked lysozyme soaked with thiourea 1.5M | | Descriptor: | ACETATE ION, Lysozyme C, NITRATE ION, ... | | Authors: | Prange, T, Salem, M. | | Deposit date: | 2005-11-23 | | Release date: | 2006-04-25 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | On the edge of the denaturation process: Application of X-ray diffraction to barnase and lysozyme cross-linked crystals with denaturants in molar concentrations.
Biochim.Biophys.Acta, 1764, 2006
|
|
1BRG
 
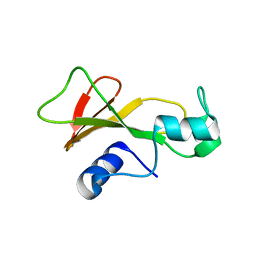 | |
1B21
 
 | | DELETION OF A BURIED SALT BRIDGE IN BARNASE | | Descriptor: | PROTEIN (BARNASE), ZINC ION | | Authors: | Vaughan, C.K, Harryson, P, Buckle, A.M, Oliveberg, M, Fersht, A.R. | | Deposit date: | 1998-12-03 | | Release date: | 1998-12-09 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A structural double-mutant cycle: estimating the strength of a buried salt bridge in barnase.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1BAN
 
 | |
1B20
 
 | | DELETION OF A BURIED SALT-BRIDGE IN BARNASE | | Descriptor: | PROTEIN (BARNASE), ZINC ION | | Authors: | Vaughan, C.K, Harryson, P, Buckle, A.M, Oliveberg, M, Fersht, A.R. | | Deposit date: | 1998-12-03 | | Release date: | 1998-12-09 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | A structural double-mutant cycle: estimating the strength of a buried salt bridge in barnase.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1B2Z
 
 | | DELETION OF A BURIED SALT BRIDGE IN BARNASE | | Descriptor: | PROTEIN (BARNASE), ZINC ION | | Authors: | Vaughan, C.K, Harryson, P, Buckle, A.M, Oliveberg, M, Fersht, A.R. | | Deposit date: | 1998-12-03 | | Release date: | 1998-12-09 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | A structural double-mutant cycle: estimating the strength of a buried salt bridge in barnase.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1BAO
 
 | |
