2CNB
 
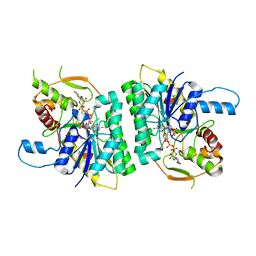 | | Trypanosoma brucei UDP-galactose-4-epimerase in ternary complex with NAD and the substrate analogue UDP-4-deoxy-4-fluoro-alpha-D-galactose | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, UDP-GALACTOSE-4-EPIMERASE, URIDINE-5'-DIPHOSPHATE-4-DEOXY-4-FLUORO-ALPHA-D-GALACTOSE | | Authors: | Alphey, M.S, Ferguson, M.A.J, Hunter, W.N. | | Deposit date: | 2006-05-18 | | Release date: | 2006-06-12 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Trypanosoma Brucei Udp-Galactose-4-Epimerase in Ternary Complex with Nad+ and the Substrate Analogue Udp-4-Deoxy-4-Fluoro-Alpha-D-Galactose
Acta Crystallogr.,Sect.F, 62, 2006
|
|
2CHO
 
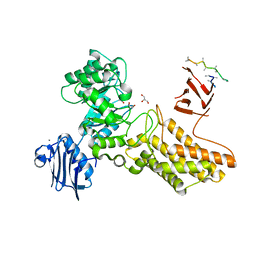 | | Bacteroides thetaiotaomicron hexosaminidase with O-GlcNAcase activity | | Descriptor: | ACETATE ION, CALCIUM ION, GLUCOSAMINIDASE, ... | | Authors: | Dennis, R.J, Taylor, E.J, Macauley, M.S, Stubbs, K.A, Turkenburg, J.P, Hart, S.J, Black, G.N, Vocadlo, D.J, Davies, G.J. | | Deposit date: | 2006-03-16 | | Release date: | 2006-06-19 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structure and Mechanism of a Bacterial B-Glucosaminidase Having O-Glcnacase Activity
Nat.Struct.Mol.Biol., 13, 2006
|
|
2CHN
 
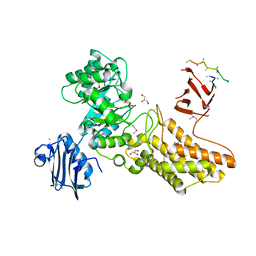 | | Bacteroides thetaiotaomicron hexosaminidase with O-GlcNAcase activity- NAG-thiazoline complex | | Descriptor: | 3AR,5R,6S,7R,7AR-5-HYDROXYMETHYL-2-METHYL-5,6,7,7A-TETRAHYDRO-3AH-PYRANO[3,2-D]THIAZOLE-6,7-DIOL, CALCIUM ION, GLUCOSAMINIDASE, ... | | Authors: | Dennis, R.J, Taylor, E.J, Macauley, M.S, Stubbs, K.A, Turkenburg, J.P, Hart, S.J, Black, G.N, Vocadlo, D.J, Davies, G.J. | | Deposit date: | 2006-03-15 | | Release date: | 2006-05-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structure and Mechanism of a Bacterial B-Glucosaminidase Having O-Glcnacase Activity
Nat.Struct.Mol.Biol., 13, 2006
|
|
2CXV
 
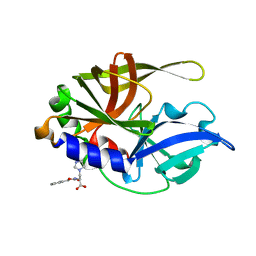 | | Dual Modes of Modification of Hepatitis A Virus 3C Protease by a Serine-Derived betaLactone: Selective Crystallization and High-resolution Structure of the His-102 Adduct | | Descriptor: | N-[(BENZYLOXY)CARBONYL]-L-ALANINE, Probable protein P3C | | Authors: | Yin, J, Bergmann, E.M, Cherney, M.M, Lall, M.S, Jain, R.P, Vederas, J.C, James, M.N.G. | | Deposit date: | 2005-07-01 | | Release date: | 2005-12-27 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Dual Modes of Modification of Hepatitis A Virus 3C Protease by a Serine-derived beta-Lactone: Selective Crystallization and Formation of a Functional Catalytic Triad in the Active Site
J.MOL.BIOL., 354, 2005
|
|
2C7Y
 
 | | plant enzyme | | Descriptor: | 3-KETOACYL-COA THIOLASE 2 | | Authors: | Sundaramoorthy, R, Micossi, E, Alphey, M.S, Germain, V, Bryce, J.H, Smith, S.M, Leonard, G.A, Hunter, W.N. | | Deposit date: | 2005-11-30 | | Release date: | 2006-05-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Crystal Structure of a Plant 3-Ketoacyl-Coa Thiolase Reveals the Potential for Redox Control of Peroxisomal Fatty Acid Beta-Oxidation.
J.Mol.Biol., 359, 2006
|
|
2CIO
 
 | |
2C7Z
 
 | | Plant enzyme crystal form II | | Descriptor: | 3-KETOACYL-COA THIOLASE 2 | | Authors: | Sundaramoorthy, R, Micossi, E, Alphey, M.S, Leonard, G.A, Hunter, W.N. | | Deposit date: | 2005-11-30 | | Release date: | 2006-05-17 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | The Crystal Structure of a Plant 3-Ketoacyl-Coa Thiolase Reveals the Potential for Redox Control of Peroxisomal Fatty Acid Beta-Oxidation.
J.Mol.Biol., 359, 2006
|
|
2EI3
 
 | | Anaerobic Crystal Structure Analysis of the 1,2-dihydroxynaphthalene dioxygenase from Pseudomonas sp. strain C18 complexes with 2,3-dihydroxybiphenyl | | Descriptor: | 1,2-dihydroxynaphthalene dioxygenase, BIPHENYL-2,3-DIOL, FE (II) ION, ... | | Authors: | Neau, D.B, Kelker, M.S, Colbert, C.L, Bolin, J.T. | | Deposit date: | 2007-03-10 | | Release date: | 2008-02-19 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural explanation for success and failure in the enzymatic ring-cleavage of 3,4 dihydroxybiphenyl and related PCB metabolites
To be Published
|
|
2EI0
 
 | | Anaerobic Crystal Structure Analysis of 1,2-dihydroxynaphthalene dioxygenase from Pseudomonas sp. strain C18 complexed with 3,4-dihydroxybiphenyl | | Descriptor: | 1,1'-BIPHENYL-3,4-DIOL, 1,2-dihydroxynaphthalene dioxygenase, FE (II) ION, ... | | Authors: | Neau, D.B, Kelker, M.S, Colbert, C.L, Bolin, J.T. | | Deposit date: | 2007-03-10 | | Release date: | 2008-02-19 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural explanation for success and failure in the enzymatic ring-cleavage of 3,4 dihydroxybiphenyl and related PCB metabolites
To be Published
|
|
2EI2
 
 | | Crystal Structure Analysis of the 1,2-dihydroxynaphthalene dioxygenase from Pseudomonas sp. stain C18 | | Descriptor: | 1,2-dihydroxynaphthalene dioxygenase, FE (II) ION, GLYCEROL, ... | | Authors: | Neau, D.B, Kelker, M.S, Colbert, C.L, Bolin, J.T. | | Deposit date: | 2007-03-10 | | Release date: | 2008-02-19 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Structural explanation for success and failure in the enzymatic ring-cleavage of 3,4 dihydroxybiphenyl and related PCB metabolites
To be Published
|
|
2EI1
 
 | | Anaerobic Crystal Structure Analysis of the 1,2-dihydroxynaphthalene dioxygeanse of Pseudomonas sp. strain C18 complexes to 1,2-dihydroxynaphthalene | | Descriptor: | 1,2-dihydroxynaphthalene dioxygenase, FE (II) ION, GLYCEROL, ... | | Authors: | Neau, D.B, Kelker, M.S, Colbert, C.L, Bolin, J.T. | | Deposit date: | 2007-03-10 | | Release date: | 2008-02-19 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Structural explanation for success and failure in the enzymatic ring-cleavage of 3,4 dihydroxybiphenyl and related PCB metabolites.
To be Published
|
|
2EHZ
 
 | | Anaerobic Crystal Structure Analysis of 1,2-dihydroxynaphthalene dioxygenase from Pseudomonas sp. strain C18 complexed with 4-methylcatechol | | Descriptor: | 1,2-ETHANEDIOL, 1,2-dihydroxynaphthalene dioxygenase, 4-METHYLCATECHOL, ... | | Authors: | Neau, D.B, Kelker, M.S, Colbert, C.L, Bolin, J.T. | | Deposit date: | 2007-03-10 | | Release date: | 2008-02-19 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structural explanation for success and failure in the enzymatic ring-cleavage of 3,4 dihydroxybiphenyl and related PCB metabolites
To be Published
|
|
7VFI
 
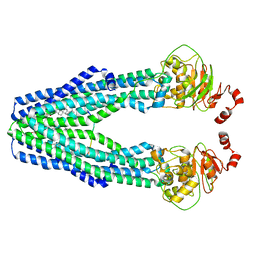 | | Cryo-EM structure of the mouse TAPL (9mer-peptide bound) | | Descriptor: | ABC-type oligopeptide transporter ABCB9, ARG-ARG-TYR-GLN-LYS-SER-THR-GLU-LEU, CHOLESTEROL HEMISUCCINATE | | Authors: | Park, J.G, Kim, S, Jang, E, Choi, S.H, Han, H, Kim, J.W, Ju, S, Min, D.S, Jin, M.S. | | Deposit date: | 2021-09-13 | | Release date: | 2022-10-19 | | Method: | ELECTRON MICROSCOPY (3.98 Å) | | Cite: | The lysosomal transporter TAPL has a dual role as peptide translocator and phosphatidylserine floppase.
Nat Commun, 13, 2022
|
|
7W9Z
 
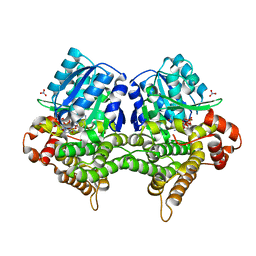 | | Crystal structure of Bacillus subtilis YugJ in complex with NADP and nitrate | | Descriptor: | Iron-containing alcohol dehydrogenase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NITRATE ION | | Authors: | Cho, H.Y, Nam, M.S, Hong, H.J, Song, W.S, Yoon, S.I. | | Deposit date: | 2021-12-11 | | Release date: | 2022-12-07 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural and Biochemical Analysis of the Furan Aldehyde Reductase YugJ from Bacillus subtilis.
Int J Mol Sci, 23, 2022
|
|
7W9Y
 
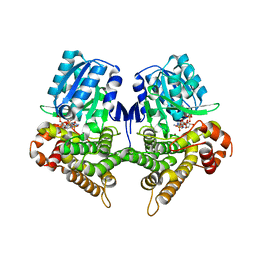 | | Crystal structure of Bacillus subtilis YugJ in complex with NADP and nickel | | Descriptor: | Iron-containing alcohol dehydrogenase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NICKEL (II) ION | | Authors: | Cho, H.Y, Nam, M.S, Hong, H.J, Song, W.S, Yoon, S.I. | | Deposit date: | 2021-12-11 | | Release date: | 2022-12-07 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Structural and Biochemical Analysis of the Furan Aldehyde Reductase YugJ from Bacillus subtilis.
Int J Mol Sci, 23, 2022
|
|
7W9X
 
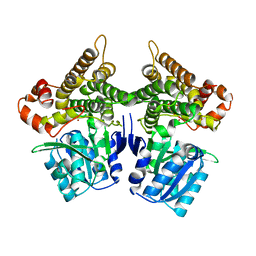 | | Crystal structure of Bacillus subtilis YugJ in complex with nickel | | Descriptor: | Iron-containing alcohol dehydrogenase, NICKEL (II) ION | | Authors: | Cho, H.Y, Nam, M.S, Hong, H.J, Song, W.S, Yoon, S.I. | | Deposit date: | 2021-12-11 | | Release date: | 2022-12-07 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.151 Å) | | Cite: | Structural and Biochemical Analysis of the Furan Aldehyde Reductase YugJ from Bacillus subtilis.
Int J Mol Sci, 23, 2022
|
|
3H5H
 
 | |
3H5E
 
 | |
3H5J
 
 | | LeuD_1-168 small subunit of isopropylmalate isomerase (Rv2987c) from mycobacterium tuberculosis | | Descriptor: | 1,2-ETHANEDIOL, 3-isopropylmalate dehydratase small subunit, SULFATE ION | | Authors: | Manikandan, K, Geerlof, A, Weiss, M.S. | | Deposit date: | 2009-04-22 | | Release date: | 2010-04-07 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Structural studies on the enzyme complex isopropylmalate isomerase (LeuCD) from Mycobacterium tuberculosis
Proteins, 79, 2011
|
|
7XLJ
 
 | | The crystal structure of ORE1(ANAC092) NAC domain | | Descriptor: | NAC domain-containing protein 92 | | Authors: | Chun, I.S, Kim, M.S. | | Deposit date: | 2022-04-21 | | Release date: | 2023-03-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structural basis of DNA binding by the NAC transcription factor ORE1, a master regulator of plant senescence.
Plant Commun., 4, 2023
|
|
7XP3
 
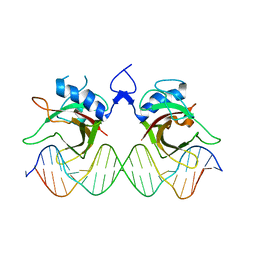 | | DNA complex form of ORESARA1(ANAC092) NAC Domain | | Descriptor: | DNA (5'-D(*AP*GP*TP*TP*AP*CP*GP*TP*AP*CP*GP*GP*CP*AP*CP*AP*CP*GP*TP*AP*AP*C)-3'), DNA (5'-D(*TP*GP*TP*TP*AP*CP*GP*TP*GP*TP*GP*CP*CP*GP*TP*AP*CP*GP*TP*AP*AP*C)-3'), DNA (5'-D(P*CP*AP*CP*AP*CP*GP*TP*AP*AP*C)-3'), ... | | Authors: | Chun, I.S, Kim, M.S. | | Deposit date: | 2022-05-03 | | Release date: | 2023-03-29 | | Last modified: | 2023-05-24 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Structural basis of DNA binding by the NAC transcription factor ORE1, a master regulator of plant senescence.
Plant Commun., 4, 2023
|
|
7YD4
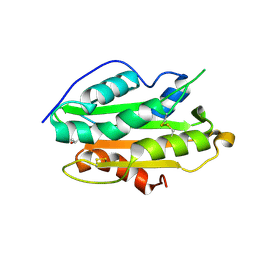 
 | | Crystal structure of an N terminal truncated secreted protein, Rv0398c from Mycobacterium tuberculosis | | Descriptor: | GLYCEROL, Possible secreted protein | | Authors: | Saha, R, Mukherjee, S, Singh, B.K, Weiss, M.S, De, S, Das, A.K. | | Deposit date: | 2022-07-03 | | Release date: | 2023-06-28 | | Method: | X-RAY DIFFRACTION (1.896 Å) | | Cite: | Crystal structure of a mycobacterial secretory protein Rv0398c and in silico prediction of its export pathway
Biochem.Biophys.Res.Commun., 672, 2023
|
|
7YJJ
 
 | |
7Z6R
 
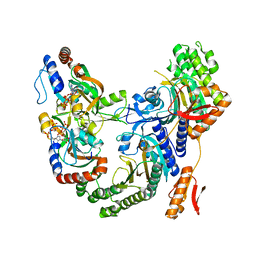 | | Psychrobacter arcticus ATPPRT (HisGZ) R56A mutant bound to ATP and PRPP | | Descriptor: | 1-O-pyrophosphono-5-O-phosphono-alpha-D-ribofuranose, ADENOSINE-5'-TRIPHOSPHATE, ATP phosphoribosyltransferase, ... | | Authors: | Alphey, M.S, Fisher, G, da Silva, R.G. | | Deposit date: | 2022-03-14 | | Release date: | 2022-03-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Allosteric rescue of catalytically impaired ATP phosphoribosyltransferase variants links protein dynamics to active-site electrostatic preorganisation.
Nat Commun, 13, 2022
|
|
7Z8U
 
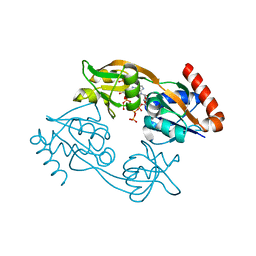 | | Catalytic subunit HisG R56A mutant from Psychrobacter arcticus ATPPRT (HisGZ) in complex with ATP and PRPP | | Descriptor: | 1-O-pyrophosphono-5-O-phosphono-alpha-D-ribofuranose, ADENOSINE-5'-TRIPHOSPHATE, ATP phosphoribosyltransferase, ... | | Authors: | Alphey, M.S, Fisher, G, da Silva, R.G. | | Deposit date: | 2022-03-18 | | Release date: | 2022-03-30 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Allosteric rescue of catalytically impaired ATP phosphoribosyltransferase variants links protein dynamics to active-site electrostatic preorganisation.
Nat Commun, 13, 2022
|
|
