4RCX
 
 | |
3K4G
 
 | |
3N97
 
 | | RNA polymerase alpha C-terminal domain (E. coli) and sigma region 4 (T. aq. mutant) bound to (UP,-35 element) DNA | | Descriptor: | ACETATE ION, DNA (5'-D(*CP*CP*AP*TP*GP*TP*CP*AP*AP*GP*TP*AP*CP*TP*TP*TP*TP*TP*TP*CP*C)-3'), DNA (5'-D(*TP*GP*GP*AP*AP*AP*AP*AP*AP*GP*TP*AP*CP*TP*TP*GP*AP*CP*AP*TP*GP*G)-3'), ... | | Authors: | Lara-Gonzalez, S, Birktoft, J.J, Lawson, C.L. | | Deposit date: | 2010-05-28 | | Release date: | 2011-06-08 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.252 Å) | | Cite: | The RNA Polymerase alpha Subunit Recognizes the DNA Shape of the Upstream Promoter Element.
Biochemistry, 59, 2020
|
|
3N4M
 
 | | E. coli RNA polymerase alpha subunit C-terminal domain in complex with CAP and DNA | | Descriptor: | ADENOSINE-3',5'-CYCLIC-MONOPHOSPHATE, Catabolite gene activator, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Lara-Gonzalez, S, Birktoft, J.J, Lawson, C.L. | | Deposit date: | 2010-05-21 | | Release date: | 2011-05-25 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.987 Å) | | Cite: | The RNA Polymerase alpha Subunit Recognizes the DNA Shape of the Upstream Promoter Element.
Biochemistry, 59, 2020
|
|
4O53
 
 | | Crystal Structure of Trichomonas vaginalis Triosephosphate Isomerase Ile45-Leu mutant (Tvag_497370) | | Descriptor: | SODIUM ION, Triosephosphate isomerase | | Authors: | Lara-Gonzalez, S, Montero-Moran, G.M, Estrella-Hernandez, P, Benitez-Cardoza, C.G, Brieba, L.G. | | Deposit date: | 2013-12-19 | | Release date: | 2014-12-24 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Engineering mutants with altered dimer-monomer equilibrium reveal the existence of stable monomeric Triosephosphate isomerases
To be Published
|
|
4O4W
 
 | |
4O52
 
 | |
4O4V
 
 | |
4O50
 
 | |
4O57
 
 | | Crystal Structure of Trichomonas vaginalis Triosephosphate Isomerase Ile45-Tyr mutant (Tvag_497370) | | Descriptor: | SODIUM ION, Triosephosphate isomerase | | Authors: | Lara-Gonzalez, S, Estrella-Hernandez, P, Montero-Moran, G.M, Benitez-Cardoza, C.G, Brieba, L.G. | | Deposit date: | 2013-12-19 | | Release date: | 2014-12-24 | | Method: | X-RAY DIFFRACTION (1.793 Å) | | Cite: | Engineering mutants with altered dimer-monomer equilibrium reveal the existence of stable monomeric Triosephosphate isomerases
To be Published
|
|
4O54
 
 | |
3M86
 
 | | Crystal structure of the cysteine protease inhibitor, EhICP2, from Entamoeba histolytica | | Descriptor: | Amoebiasin-2, CHLORIDE ION, PENTAETHYLENE GLYCOL, ... | | Authors: | Lara-Gonzalez, S, Casados-Vazquez, L.E, Brieba, L.G. | | Deposit date: | 2010-03-17 | | Release date: | 2011-02-02 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structure of the cysteine protease inhibitor 2 from Entamoeba histolytica: functional convergence of a common protein fold.
Gene, 471, 2011
|
|
5VWN
 
 | | Triosephosphate isomerases deletion loop 3 from Trichomonas vaginalis | | Descriptor: | SODIUM ION, Triosephosphate isomerase | | Authors: | Lara-Gonzalez, S, Rojas-Mendez, K, Jimenez-Sandoval, P, Estrella-Hernandez, P, Brieba, L.G. | | Deposit date: | 2017-05-22 | | Release date: | 2018-04-04 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | A competent catalytic active site is necessary for substrate induced dimer assembly in triosephosphate isomerase.
Biochim. Biophys. Acta, 1865, 2017
|
|
3QST
 
 | |
3QSR
 
 | |
3P91
 
 | |
3M88
 
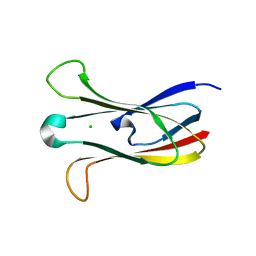 | | Crystal structure of the cysteine protease inhibitor, EhICP2, from Entamoeba histolytica | | Descriptor: | Amoebiasin-2, CHLORIDE ION, SULFATE ION | | Authors: | Lara-Gonzalez, S, Casados-Vazquez, L.E, Brieba, L.G. | | Deposit date: | 2010-03-17 | | Release date: | 2011-02-02 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the cysteine protease inhibitor 2 from Entamoeba histolytica: functional convergence of a common protein fold.
Gene, 471, 2011
|
|
4OHQ
 
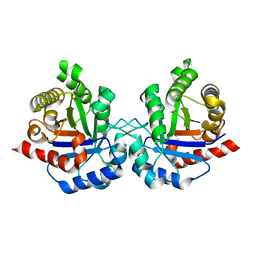 | |
1HE8
 
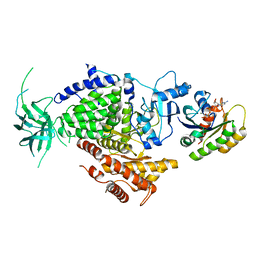 | | Ras G12V - PI 3-kinase gamma complex | | Descriptor: | MAGNESIUM ION, PHOSPHATIDYLINOSITOL 3-KINASE CATALYTIC SUBUNIT, GAMMA ISOFORM, ... | | Authors: | Pacold, M.E, Suire, S, Perisic, O, Lara-Gonzalez, S, Davis, C.T, Hawkins, P.T, Walker, E.H, Stephens, L, Eccleston, J.F, Williams, R.L. | | Deposit date: | 2000-11-20 | | Release date: | 2001-01-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal Structure and Functional Analysis of Ras Binding to its Effector Phosphoinositide 3-Kinase Gamma
Cell(Cambridge,Mass.), 103, 2000
|
|
4WJE
 
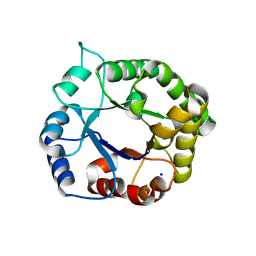 | |
4IYF
 
 | |
4IYD
 
 | |
4OBT
 
 | | Crystal structure of Arabidopsis thaliana cytosolic triose phosphate isomerase | | Descriptor: | Triosephosphate isomerase, cytosolic | | Authors: | Lopez-Castillo, M, Jimenez-Sandoval, P, Lara-Gonzalez, S, Brieba, L.G, Baruch, N. | | Deposit date: | 2014-01-07 | | Release date: | 2016-01-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural Basis for Redox Regulation of Cytoplasmic and Chloroplastic Triosephosphate Isomerases from Arabidopsis thaliana.
Front Plant Sci, 7, 2016
|
|
5CIZ
 
 | |
3IYD
 
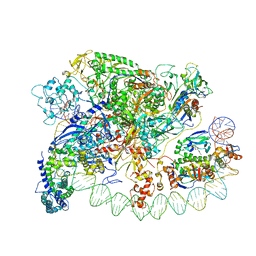 | | Three-dimensional EM structure of an intact activator-dependent transcription initiation complex | | Descriptor: | ADENOSINE-3',5'-CYCLIC-MONOPHOSPHATE, Catabolite gene activator, DNA (98-MER), ... | | Authors: | Hudson, B.P, Quispe, J, Lara, S, Kim, Y, Berman, H, Arnold, E, Ebright, R.H, Lawson, C.L. | | Deposit date: | 2009-08-01 | | Release date: | 2009-11-10 | | Last modified: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (19.799999 Å) | | Cite: | Three-dimensional EM structure of an intact activator-dependent transcription initiation complex
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
