3F8N
 
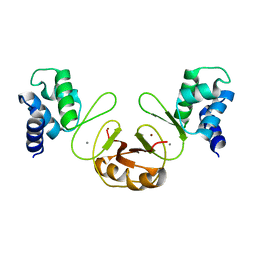 | | Crystal structure of PerR-Zn-Mn | | Descriptor: | MANGANESE (II) ION, Peroxide operon regulator, ZINC ION | | Authors: | Traore, D.A.K, Ferrer, J.-L, Jacquamet, L, Duarte, V, Latour, J.-M. | | Deposit date: | 2008-11-13 | | Release date: | 2009-06-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Structural characterization of the active form of PerR: insights into the metal-induced activation of PerR and Fur proteins for DNA binding
Mol.Microbiol., 73, 2009
|
|
3CIA
 
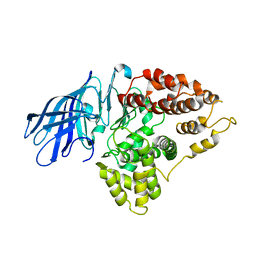 | | Crystal structure of cold-aminopeptidase from Colwellia psychrerythraea | | Descriptor: | ZINC ION, cold-active aminopeptidase | | Authors: | Bauvois, C, Jacquamet, L, Borel, F, Ferrer, J.-L. | | Deposit date: | 2008-03-11 | | Release date: | 2008-07-01 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structure of the Cold-active Aminopeptidase from Colwellia psychrerythraea, a Close Structural Homologue of the Human Bifunctional Leukotriene A4 Hydrolase.
J.Biol.Chem., 283, 2008
|
|
3CX3
 
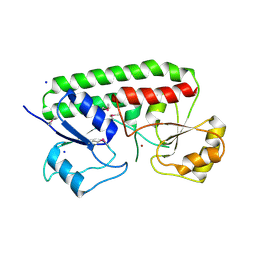 | | Crystal structure Analysis of the Streptococcus pneumoniae AdcAII protein | | Descriptor: | Lipoprotein, SODIUM ION, ZINC ION | | Authors: | Loisel, E, Durmort, C, Jacquamet, L. | | Deposit date: | 2008-04-23 | | Release date: | 2008-08-05 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | AdcAII, a new pneumococcal Zn-binding protein homologous with ABC transporters: biochemical and structural analysis.
J.Mol.Biol., 381, 2008
|
|
1ZLQ
 
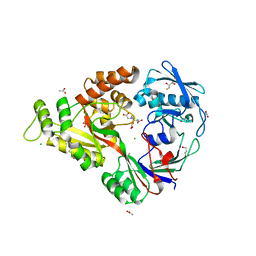 | | Crystallographic and spectroscopic evidence for high affinity binding of Fe EDTA (H2O)- to the periplasmic nickel transporter NikA | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, ACETATE ION, CHLORIDE ION, ... | | Authors: | Cherrier, M.V, Martin, L, Cavazza, C, Jacquamet, L, Lemaire, D, Gaillard, J, Fontecilla Camps, J.C. | | Deposit date: | 2005-05-09 | | Release date: | 2005-08-02 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystallographic and Spectroscopic Evidence for High Affinity Binding of FeEDTA(H(2)O)(-) to the Periplasmic Nickel Transporter NikA
J.Am.Chem.Soc., 127, 2005
|
|
1ZC2
 
 | | Crystal Structure of plasmid-encoded class C beta-lactamase CMY-2 complexed with citrate molecule | | Descriptor: | CITRIC ACID, beta-lactamase class C | | Authors: | Bauvois, C, Jacquamet, L, Fieulaine, S, Frere, J.-M, Galleni, M, Ferrer, J.-L. | | Deposit date: | 2005-04-10 | | Release date: | 2006-04-25 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Crystallographic structure of plasmid-encoded CMY-2 beta-lactamase revealed citrate molecule in the active site.
To be Published
|
|
2WCI
 
 | | Structure of E. coli monothiol glutaredoxin GRX4 homodimer | | Descriptor: | FE2/S2 (INORGANIC) CLUSTER, GLUTAREDOXIN-4, GLUTATHIONE, ... | | Authors: | Iwema, T, Picchiocci, A, Traore, D.A.K, Ferrer, J.-L, Chauvat, F, Jacquamet, L. | | Deposit date: | 2009-03-12 | | Release date: | 2009-06-23 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Basis for Delivery of the Intact [Fe2S2] Cluster by Monothiol Glutaredoxin.
Biochemistry, 48, 2009
|
|
2FU4
 
 | | Crystal Structure of the DNA binding domain of E.coli FUR (Ferric Uptake Regulator) | | Descriptor: | CADMIUM ION, CHLORIDE ION, Ferric uptake regulation protein, ... | | Authors: | Pecqueur, L, D'Autreaux, B, Dupuy, J, Nicolet, Y, Jacquamet, L, Brutscher, B, Michaud-Soret, I, Bersch, B. | | Deposit date: | 2006-01-26 | | Release date: | 2006-05-16 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural changes of Escherichia coli ferric uptake regulator during metal-dependent dimerization and activation explored by NMR and X-ray crystallography
J.Biol.Chem., 281, 2006
|
|
2C8Q
 
 | | insuline(1sec) and UV laser excited fluorescence | | Descriptor: | INSULIN A CHAIN, INSULIN B CHAIN | | Authors: | Vernede, X, Lavault, B, Ohana, J, Nurizzo, D, Joly, J, Jacquamet, L, Felisaz, F, Cipriani, F, Bourgeois, D. | | Deposit date: | 2005-12-06 | | Release date: | 2006-03-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Uv Laser-Excited Fluorescence as a Tool for the Visualization of Protein Crystals Mounted in Loops.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2C8P
 
 | | lysozyme (60sec) and UV laser excited fluorescence | | Descriptor: | LYSOZYME C | | Authors: | Vernede, X, Lavault, B, Ohana, J, Nurizzo, D, Joly, J, Jacquamet, L, Felisaz, F, Cipriani, F, Bourgeois, D. | | Deposit date: | 2005-12-06 | | Release date: | 2006-03-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Uv Laser-Excited Fluorescence as a Tool for the Visualization of Protein Crystals Mounted in Loops.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2C8R
 
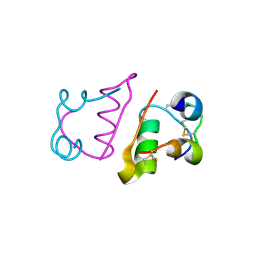 | | insuline(60sec) and UV laser excited fluorescence | | Descriptor: | INSULIN A CHAIN, INSULIN B CHAIN | | Authors: | Vernede, X, Lavault, B, Ohana, J, Nurizzo, D, Joly, J, Jacquamet, L, Felisaz, F, Cipriani, F, Bourgeois, D. | | Deposit date: | 2005-12-06 | | Release date: | 2006-03-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Uv Laser-Excited Fluorescence as a Tool for the Visualization of Protein Crystals Mounted in Loops.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2C8O
 
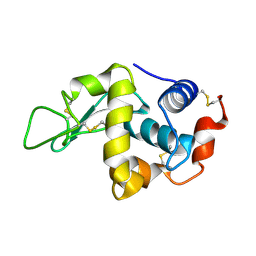 | | lysozyme (1sec) and UV lasr excited fluorescence | | Descriptor: | LYSOZYME C | | Authors: | Vernede, X, Lavault, B, Ohana, J, Nurizzo, D, Joly, J, Jacquamet, L, Felisaz, F, Cipriani, F, Bourgeois, D. | | Deposit date: | 2005-12-06 | | Release date: | 2006-03-08 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Uv Laser-Excited Fluorescence as a Tool for the Visualization of Protein Crystals Mounted in Loops.
Acta Crystallogr.,Sect.D, 62, 2006
|
|
2RGV
 
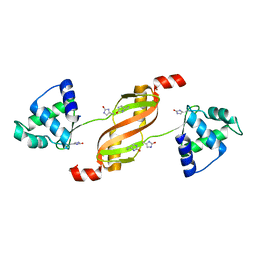 | |
2RL3
 
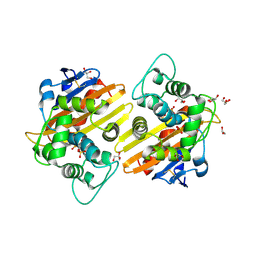 | | Crystal structure of the OXA-10 W154H mutant at pH 7 | | Descriptor: | 1,2-ETHANEDIOL, Beta-lactamase PSE-2, GLYCEROL, ... | | Authors: | Vercheval, L, Kerff, F, Herman, R, Sauvage, E, Guiet, R, Charlier, P, Frere, J.-M, Galleni, M. | | Deposit date: | 2007-10-18 | | Release date: | 2008-10-28 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Biochemistry, 48, 2009
|
|
2WGI
 
 | | Crystal structure of the acyl-enzyme OXA-10 W154A-benzylpenicillin at pH 6 | | Descriptor: | BETA-LACTAMASE OXA-10, GLYCEROL, OPEN FORM - PENICILLIN G | | Authors: | Vercheval, L, Falzone, C, Sauvage, E, Herman, R, Charlier, P, Galleni, M, Kerff, F. | | Deposit date: | 2009-04-20 | | Release date: | 2009-11-10 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Critical Role of Tryptophan 154 for the Activity and Stability of Class D Beta-Lactamases.
Biochemistry, 48, 2009
|
|
2HPB
 
 | | Crystal structure of the OXA-10 W154A mutant at pH 9.0 | | Descriptor: | Beta-lactamase PSE-2, SULFATE ION | | Authors: | Kerff, F, Falzone, C, Herman, R, Sauvage, E, Charlier, P. | | Deposit date: | 2006-07-17 | | Release date: | 2007-07-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Biochemistry, 48, 2009
|
|
2HP6
 
 | | Crystal structure of the OXA-10 W154A mutant at pH 7.5 | | Descriptor: | Beta-lactamase PSE-2, SULFATE ION | | Authors: | Kerff, F, Falzone, C, Herman, R, Sauvage, E, Charlier, P. | | Deposit date: | 2006-07-17 | | Release date: | 2007-07-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Biochemistry, 48, 2009
|
|
2HP9
 
 | | Crystal Structure of the OXA-10 W154A mutant at pH 6.0 | | Descriptor: | Beta-lactamase PSE-2, SULFATE ION | | Authors: | Kerff, F, Falzone, C, Herman, R, Sauvage, E, Charlier, P. | | Deposit date: | 2006-07-17 | | Release date: | 2007-07-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Biochemistry, 48, 2009
|
|
2HP5
 
 | | Crystal Structure of the OXA-10 W154G mutant at pH 7.0 | | Descriptor: | Beta-lactamase PSE-2, COBALT (II) ION, SULFATE ION | | Authors: | Kerff, F, Falzone, C, Herman, R, Sauvage, E, Charlier, P. | | Deposit date: | 2006-07-17 | | Release date: | 2007-07-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Critical role of tryptophan 154 for the activity and stability of class D beta-lactamases.
Biochemistry, 48, 2009
|
|
1QVZ
 
 | | Crystal structure of the S. cerevisiae YDR533c protein | | Descriptor: | YDR533c protein | | Authors: | Graille, M, Leulliot, N, Quevillon-Cheruel, S, van Tilbeurgh, H. | | Deposit date: | 2003-08-29 | | Release date: | 2004-03-30 | | Last modified: | 2021-11-10 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal structure of the YDR533c S. cerevisiae protein, a class II member of the Hsp31 family
STRUCTURE, 12, 2004
|
|
1QVV
 
 | | Crystal structure of the S. cerevisiae YDR533c protein | | Descriptor: | YDR533c protein | | Authors: | Graille, M, Leulliot, N, Quevillon-Cheruel, S, van Tilbeurgh, H. | | Deposit date: | 2003-08-29 | | Release date: | 2004-03-30 | | Last modified: | 2020-07-15 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Crystal structure of the YDR533c S. cerevisiae protein, a class II member of the Hsp31 family
STRUCTURE, 12, 2004
|
|
1QVW
 
 | | Crystal structure of the S. cerevisiae YDR533c protein | | Descriptor: | GLYCEROL, YDR533c protein | | Authors: | Graille, M, Leulliot, N, Quevillon-Cheruel, S, van Tilbeurgh, H. | | Deposit date: | 2003-08-29 | | Release date: | 2004-03-30 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the YDR533c S. cerevisiae protein, a class II member of the Hsp31 family
STRUCTURE, 12, 2004
|
|
2FE3
 
 | |
3ZEK
 
 | |
3ZEJ
 
 | |
