6KYD
 
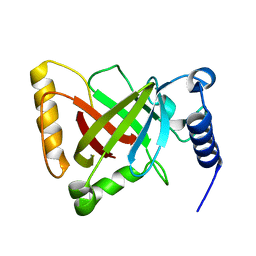 | | Structure of the R217A mutant of Clostridium difficile sortase B | | Descriptor: | Putative peptidase C60B, sortase B | | Authors: | Kang, C.Y, Huang, I.H, Wu, T.Y, Chang, J.C, Hsiao, Y.Y, Cheng, C.H, Tsai, W.J, Hsu, K.C, Wang, S.Y. | | Deposit date: | 2019-09-18 | | Release date: | 2020-02-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Functional analysis ofClostridium difficilesortase B reveals key residues for catalytic activity and substrate specificity.
J.Biol.Chem., 295, 2020
|
|
6MAA
 
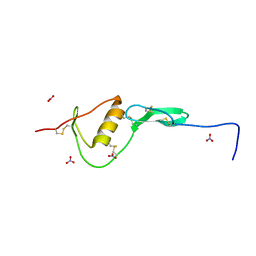 | | WFIKKN2 Follistatin Domain | | Descriptor: | NITRATE ION, WAP, Kazal, ... | | Authors: | McCoy, J.C, Walker, R.G, Thomas, T.B. | | Deposit date: | 2018-08-27 | | Release date: | 2019-03-06 | | Last modified: | 2019-12-04 | | Method: | X-RAY DIFFRACTION (1.394 Å) | | Cite: | Crystal structure of the WFIKKN2 follistatin domain reveals insight into how it inhibits growth differentiation factor 8 (GDF8) and GDF11.
J.Biol.Chem., 294, 2019
|
|
6LYT
 
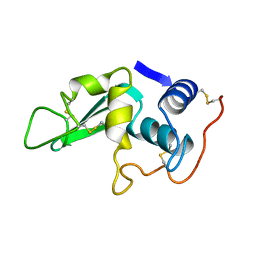 | |
6MJ1
 
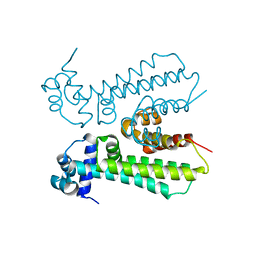 | |
6MJ0
 
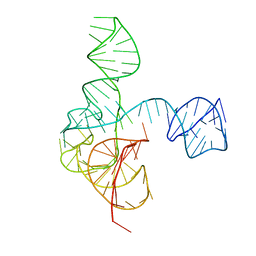 | | Crystal structure of the complete turnip yellow mosaic virus 3'UTR | | Descriptor: | RNA (101-MER) | | Authors: | Hartwick, E.W, Costantino, D.A, MacFadden, A, Nix, J.C, Tian, S, Das, R, Kieft, J.S. | | Deposit date: | 2018-09-20 | | Release date: | 2019-01-09 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Ribosome-induced RNA conformational changes in a viral 3'-UTR sense and regulate translation levels.
Nat Commun, 9, 2018
|
|
6MU4
 
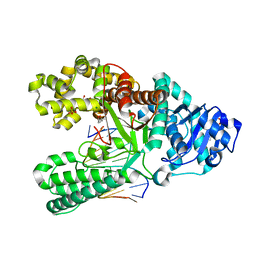 | | Bst DNA polymerase I FANA/DNA binary complex | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, DNA (5'-D(P*GP*CP*GP*AP*TP*CP*AP*CP*GP*T)-3'), DNA polymerase I, ... | | Authors: | Jackson, L.N, Chim, N, Chaput, J.C. | | Deposit date: | 2018-10-22 | | Release date: | 2019-06-05 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Crystal structures of a natural DNA polymerase that functions as an XNA reverse transcriptase.
Nucleic Acids Res., 47, 2019
|
|
6MTG
 
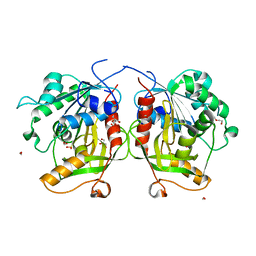 | | A Single Reactive Noncanonical Amino Acid is Able to Dramatically Stabilize Protein Structure | | Descriptor: | DI(HYDROXYETHYL)ETHER, FORMIC ACID, GLYCEROL, ... | | Authors: | Li, J.C, Nasertorabi, F, Xuan, W, Han, G.W, Stevens, R.C, Schultz, P.G. | | Deposit date: | 2018-10-19 | | Release date: | 2019-06-26 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | A Single Reactive Noncanonical Amino Acid Is Able to Dramatically Stabilize Protein Structure.
Acs Chem.Biol., 14, 2019
|
|
6MYD
 
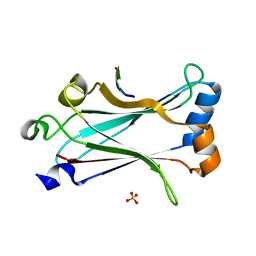 | | Structure of zebrafish TRAF6 in complex with STING CTT | | Descriptor: | STING CTT, Transmembrane protein 173, SULFATE ION, ... | | Authors: | de Oliveira Mann, C.C, Orzalli, M.H, King, D.S, Kagan, J.C, Lee, A.S.Y, Kranzusch, P.J. | | Deposit date: | 2018-11-01 | | Release date: | 2019-05-01 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.399 Å) | | Cite: | Modular Architecture of the STING C-Terminal Tail Allows Interferon and NF-kappa B Signaling Adaptation.
Cell Rep, 27, 2019
|
|
6MU5
 
 | | Bst DNA polymerase I TNA/DNA binary complex | | Descriptor: | DNA (5'-D(P*GP*CP*GP*AP*TP*CP*AP*CP*GP*T)-3'), DNA polymerase I, SULFATE ION, ... | | Authors: | Jackson, L.N, Chim, N, Chaput, J.C. | | Deposit date: | 2018-10-22 | | Release date: | 2019-06-05 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.912 Å) | | Cite: | Crystal structures of a natural DNA polymerase that functions as an XNA reverse transcriptase.
Nucleic Acids Res., 47, 2019
|
|
6F48
 
 | | Structure of quinolinate synthase with reaction intermediates X and Y | | Descriptor: | 2-imino,3-carboxy,5-hydroxy,6-oxo hexanoic acid, 5-hydroxy,-4,5-dihydroquinolinate, CHLORIDE ION, ... | | Authors: | Volbeda, A, Fontecilla-Camps, J.C. | | Deposit date: | 2017-11-29 | | Release date: | 2018-04-25 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Crystallographic Trapping of Reaction Intermediates in Quinolinic Acid Synthesis by NadA.
ACS Chem. Biol., 13, 2018
|
|
6F4L
 
 | |
6F4D
 
 | |
1VTQ
 
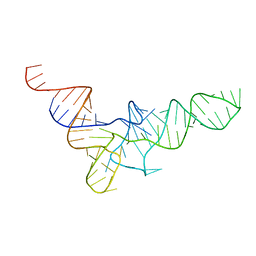 | | THREE-DIMENSIONAL STRUCTURE OF YEAST T-RNA-ASP. I. STRUCTURE DETERMINATION | | Descriptor: | T-RNA-ASP | | Authors: | Comarmond, M.B, Giege, R, Thierry, J.C, Moras, D, Fischer, J. | | Deposit date: | 1985-06-11 | | Release date: | 2011-07-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Three-Dimensional Structure of Yeast T-RNA-ASP. I. Structure Determination
Acta Crystallogr.,Sect.B, 42, 1986
|
|
6G74
 
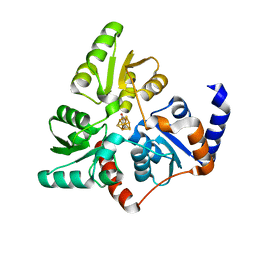 | |
2IY9
 
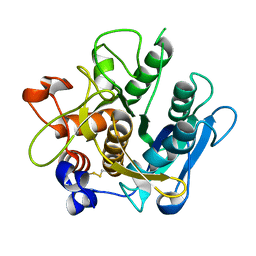 | | Crystal structure of the A-subunit of the AB5 toxin from E. coli | | Descriptor: | SUBA | | Authors: | Paton, A.W, Beddoe, T, Thorpe, C.M, Whisstock, J.C, Wilce, M.C.J, Rossjohn, J, Talbot, U.M, Paton, J.C. | | Deposit date: | 2006-07-13 | | Release date: | 2006-10-04 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Ab5 Subtilase Cytotoxin Inactivates the Endoplasmic Reticulum Chaperone Bip
Nature, 443, 2006
|
|
6XRB
 
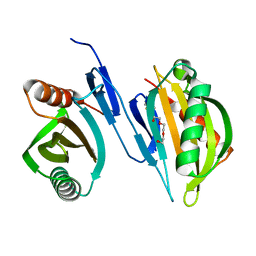 | | Crystal structure of SciW from Salmonella typhimurium | | Descriptor: | SciW, TRIETHYLENE GLYCOL | | Authors: | Sachar, K, Ahmad, S, Whitney, J.C, Prehna, G. | | Deposit date: | 2020-07-11 | | Release date: | 2020-12-30 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Structural basis for effector transmembrane domain recognition by type VI secretion system chaperones.
Elife, 9, 2020
|
|
6XRF
 
 | | EagT6 Tse6 NT complex | | Descriptor: | Effector EagT6, PAAR motif family protein | | Authors: | Sachar, K, Ahmad, S, Whitney, J.C, Prehna, G. | | Deposit date: | 2020-07-12 | | Release date: | 2020-12-30 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.56 Å) | | Cite: | Structural basis for effector transmembrane domain recognition by type VI secretion system chaperones.
Elife, 9, 2020
|
|
6X90
 
 | |
5H8P
 
 | |
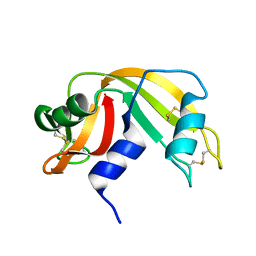
4RAT
 | |
6YS8
 
 | |
5H8M
 
 | | Crystal structure of Mycobacterium tuberculosis malate synthase C619A, G459A mutant in complex with product malate | | Descriptor: | (2S)-2-hydroxybutanedioic acid, MAGNESIUM ION, Malate synthase G | | Authors: | Krieger, I.V, Huang, H.-L, Sacchettini, J.C. | | Deposit date: | 2015-12-23 | | Release date: | 2016-10-26 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Mycobacterium tuberculosis Malate Synthase Structures with Fragments Reveal a Portal for Substrate/Product Exchange.
J. Biol. Chem., 291, 2016
|
|
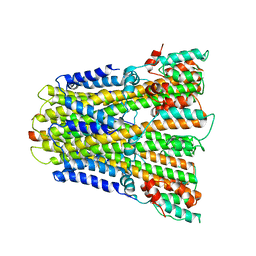
6YSF
 | |
5H8U
 
 | | Crystal structure of mycobacterium tuberculosis wild-type malate synthase in complex with product malate | | Descriptor: | (2S)-2-hydroxybutanedioic acid, GLYOXYLIC ACID, MAGNESIUM ION, ... | | Authors: | Krieger, I.V, Huang, H.-L, Sacchettini, J.C. | | Deposit date: | 2015-12-23 | | Release date: | 2016-10-26 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Mycobacterium tuberculosis Malate Synthase Structures with Fragments Reveal a Portal for Substrate/Product Exchange.
J. Biol. Chem., 291, 2016
|
|
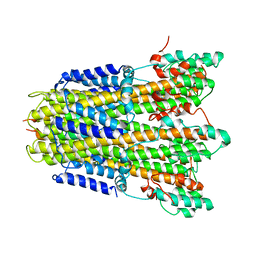
6YSL
 | |
