4EI7
 
 | | Crystal structure of Bacillus cereus TubZ, GDP-form | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, Plasmid replication protein RepX | | Authors: | Hayashi, I, Hoshino, S. | | Deposit date: | 2012-04-05 | | Release date: | 2012-08-15 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Filament formation of the FtsZ/tubulin-like protein TubZ from the Bacillus cereus pXO1 plasmid.
J.Biol.Chem., 287, 2012
|
|
4EI9
 
 | | Crystal structure of Bacillus cereus TubZ, GTP-form | | Descriptor: | 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, GUANOSINE-5'-DIPHOSPHATE, Plasmid replication protein RepX | | Authors: | Hayashi, I, Hoshino, S. | | Deposit date: | 2012-04-05 | | Release date: | 2012-08-15 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Filament formation of the FtsZ/tubulin-like protein TubZ from the Bacillus cereus pXO1 plasmid.
J.Biol.Chem., 287, 2012
|
|
4EI8
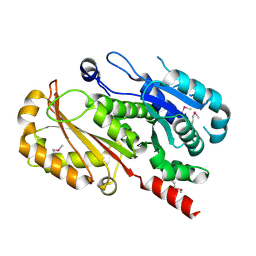 
 | |
5WQH
 
 | | Structure of fungal meroterpenoid isomerase Trt14 complexed with substrate analog and endo-terretonin D | | Descriptor: | CALCIUM ION, Isomerase trt14, methyl (2S,4aR,4bS,5S,6aS,10aS,10bS,12aS)-2,4b,7,7,10a,12,12a-heptamethyl-5-oxidanyl-1,4,6,8-tetrakis(oxidanylidene)-4a,5,6a,9,10,10b-hexahydronaphtho[1,2-h]isochromene-2-carboxylate, ... | | Authors: | Mori, T, Matsuda, Y, Abe, I. | | Deposit date: | 2016-11-26 | | Release date: | 2017-07-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.102 Å) | | Cite: | Molecular basis for the unusual ring reconstruction in fungal meroterpenoid biogenesis
Nat. Chem. Biol., 13, 2017
|
|
7T5K
 
 | | E. coli dihydroorotate dehydrogenase bound to the inhibitor HQNO | | Descriptor: | 2-HEPTYL-4-HYDROXY QUINOLINE N-OXIDE, Dihydroorotate dehydrogenase (quinone), FLAVIN MONONUCLEOTIDE, ... | | Authors: | Horwitz, S.M, Ambarian, J.A, Davis, K.M. | | Deposit date: | 2021-12-12 | | Release date: | 2022-03-02 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural insights into inhibition of the drug target dihydroorotate dehydrogenase by bacterial hydroxyalkylquinolines.
Rsc Chem Biol, 3, 2022
|
|
7T5Y
 
 | | E. coli dihydroorotate dehydrogenase bound to the inhibitor HMNQ | | Descriptor: | 3-methyl-2-[(2E)-non-2-en-1-yl]quinolin-4(1H)-one, Dihydroorotate dehydrogenase (quinone), FLAVIN MONONUCLEOTIDE, ... | | Authors: | Horwitz, S.M, Ambarian, J.A, Davis, K.M. | | Deposit date: | 2021-12-13 | | Release date: | 2022-03-02 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.62 Å) | | Cite: | Structural insights into inhibition of the drug target dihydroorotate dehydrogenase by bacterial hydroxyalkylquinolines.
Rsc Chem Biol, 3, 2022
|
|
7T6C
 
 | | E. coli dihydroorotate dehydrogenase bound to the ubiquinone surrogate DCIP | | Descriptor: | 2,6-bis(chloranyl)-4-[(4-hydroxyphenyl)amino]phenol, 2-{2-[2-(2-{2-[2-(2-ETHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, Dihydroorotate dehydrogenase (quinone), ... | | Authors: | Horwitz, S.M, Ambarian, J.A, Davis, K.M. | | Deposit date: | 2021-12-13 | | Release date: | 2022-03-02 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Structural insights into inhibition of the drug target dihydroorotate dehydrogenase by bacterial hydroxyalkylquinolines.
Rsc Chem Biol, 3, 2022
|
|
7T6H
 
 | | E. coli dihydroorotate dehydrogenase | | Descriptor: | Dihydroorotate dehydrogenase (quinone), FLAVIN MONONUCLEOTIDE, OROTIC ACID | | Authors: | Horwitz, S.M, Ambarian, J.A, Davis, K.M. | | Deposit date: | 2021-12-13 | | Release date: | 2022-03-02 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.42 Å) | | Cite: | Structural insights into inhibition of the drug target dihydroorotate dehydrogenase by bacterial hydroxyalkylquinolines.
Rsc Chem Biol, 3, 2022
|
|
4YLA
 
 | | Crystal structure of the indole prenyltransferase MpnD complexed with indolactam V and DMSPP | | Descriptor: | (2S,5S)-5-(hydroxymethyl)-1-methyl-2-(propan-2-yl)-1,2,4,5,6,8-hexahydro-3H-[1,4]diazonino[7,6,5-cd]indol-3-one, Aromatic prenyltransferase, DIMETHYLALLYL S-THIOLODIPHOSPHATE, ... | | Authors: | Mori, T, Morita, H, Abe, I. | | Deposit date: | 2015-03-05 | | Release date: | 2016-03-16 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Manipulation of prenylation reactions by structure-based engineering of bacterial indolactam prenyltransferases.
Nat Commun, 7, 2016
|
|
4YL7
 
 | |
6J82
 
 | | Crystal structure of TleB apo | | Descriptor: | Cytochrome P-450, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Alblova, M, Nakamura, H, Mori, T, Abe, I. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-07 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.202 Å) | | Cite: | Molecular basis for the P450-catalyzed C-N bond formation in indolactam biosynthesis.
Nat.Chem.Biol., 15, 2019
|
|
6J88
 
 | | Crystal structure of HinD with benzo[b]thiophen analog | | Descriptor: | N-[(2S)-1-(1-benzothiophen-3-yl)-3-hydroxypropan-2-yl]-N~2~-methyl-L-valinamide, Nocardicin N-oxygenase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Fei, H, Mori, T, Abe, I. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-07 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Molecular basis for the P450-catalyzed C-N bond formation in indolactam biosynthesis.
Nat.Chem.Biol., 15, 2019
|
|
6J86
 
 | | Crystal structure of HinD with NMFT | | Descriptor: | N-[(2S)-1-hydroxy-3-(1H-indol-3-yl)propan-2-yl]-Nalpha-methyl-L-phenylalaninamide, Nocardicin N-oxygenase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Fei, H, Mori, T, Abe, I. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-07 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Molecular basis for the P450-catalyzed C-N bond formation in indolactam biosynthesis.
Nat.Chem.Biol., 15, 2019
|
|
6J85
 
 | | Crystal structure of HinD apo | | Descriptor: | Nocardicin N-oxygenase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Fei, H, Mori, T, Abe, I. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-07 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Molecular basis for the P450-catalyzed C-N bond formation in indolactam biosynthesis.
Nat.Chem.Biol., 15, 2019
|
|
6J83
 
 | | Crystal structure of TleB with NMVT | | Descriptor: | Cytochrome P-450, N-[(2S)-1-hydroxy-3-(1H-indol-3-yl)propan-2-yl]-N~2~-methyl-L-valinamide, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Fei, H, Mori, T, Abe, I. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-07 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular basis for the P450-catalyzed C-N bond formation in indolactam biosynthesis.
Nat.Chem.Biol., 15, 2019
|
|
6J87
 
 | | Crystal structure of HinD with NMFT and NO | | Descriptor: | N-[(2S)-1-hydroxy-3-(1H-indol-3-yl)propan-2-yl]-Nalpha-methyl-L-phenylalaninamide, NITRIC OXIDE, Nocardicin N-oxygenase, ... | | Authors: | Fei, H, Mori, T, Abe, I. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-07 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Molecular basis for the P450-catalyzed C-N bond formation in indolactam biosynthesis.
Nat.Chem.Biol., 15, 2019
|
|
6J84
 
 | | Crystal structure of TleB with hydroxyl analog | | Descriptor: | (2S)-2-hydroxy-N-[(2S)-1-hydroxy-3-(1H-indol-3-yl)propan-2-yl]-3-methylbutanamide, Cytochrome P-450, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Nakamura, H, Mori, T, Abe, I. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-07 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Molecular basis for the P450-catalyzed C-N bond formation in indolactam biosynthesis.
Nat.Chem.Biol., 15, 2019
|
|
4YZK
 
 | |
4YZL
 
 | | Crystal structure of the indole prenyltransferase TleC complexed with indolactam V and DMSPP | | Descriptor: | (2S,5S)-5-(hydroxymethyl)-1-methyl-2-(propan-2-yl)-1,2,4,5,6,8-hexahydro-3H-[1,4]diazonino[7,6,5-cd]indol-3-one, DIMETHYLALLYL S-THIOLODIPHOSPHATE, Tryptophan dimethylallyltransferase | | Authors: | Mori, T, Morita, H, Abe, I. | | Deposit date: | 2015-03-25 | | Release date: | 2016-03-16 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.096 Å) | | Cite: | Manipulation of prenylation reactions by structure-based engineering of bacterial indolactam prenyltransferases.
Nat Commun, 7, 2016
|
|
4YZJ
 
 | |
3X27
 
 | | Structure of McbB in complex with tryptophan | | Descriptor: | Cucumopine synthase, TRYPTOPHAN | | Authors: | Mori, T, Sahashi, S, Morita, H, Abe, I. | | Deposit date: | 2014-12-10 | | Release date: | 2015-10-28 | | Last modified: | 2015-11-04 | | Method: | X-RAY DIFFRACTION (2.481 Å) | | Cite: | Structural Basis for beta-Carboline Alkaloid Production by the Microbial Homodimeric Enzyme McbB
Chem.Biol., 22, 2015
|
|
5WQF
 
 | | Structure of fungal meroterpenoid isomerase Trt14 | | Descriptor: | CALCIUM ION, Isomerase trt14, N-PROPANOL | | Authors: | Mori, T, Iwabuchi, T, Matsuda, Y, Abe, I. | | Deposit date: | 2016-11-26 | | Release date: | 2017-07-26 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | Molecular basis for the unusual ring reconstruction in fungal meroterpenoid biogenesis
Nat. Chem. Biol., 13, 2017
|
|
5WQG
 
 | | Structure of fungal meroterpenoid isomerase Trt14 complexed with terretonin D | | Descriptor: | CALCIUM ION, Isomerase trt14, methyl (2S,4aR,4bS,5S,6aS,10aS,10bS,12aS)-2,4b,7,7,10a,12a-hexamethyl-12-methylidene-5-oxidanyl-1,4,6,8-tetrakis(oxidanylidene) -5,6a,9,10,10b,11-hexahydro-4aH-naphtho[1,2-h]isochromene-2-carboxylate | | Authors: | Mori, T, Matsuda, Y, Abe, I. | | Deposit date: | 2016-11-26 | | Release date: | 2017-07-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Molecular basis for the unusual ring reconstruction in fungal meroterpenoid biogenesis
Nat. Chem. Biol., 13, 2017
|
|
5WQI
 
 | | Structure of fungal meroterpenoid isomerase Trt14 complexed with hydrolyzed product | | Descriptor: | (1R,2S,4aS,4bS,8aS,10S,10aS)-2-[(2S)-3-methoxy-2-methyl-2-oxidanyl-3-oxidanylidene-propanoyl]-2,3,4b,8,8,10a-hexamethyl-10-oxidanyl-7,9-bis(oxidanylidene)-1,4a,5,6,8a,10-hexahydrophenanthrene-1-carboxylic acid, CALCIUM ION, Isomerase trt14 | | Authors: | Mori, T, Matsuda, Y, Abe, I. | | Deposit date: | 2016-11-27 | | Release date: | 2017-07-26 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.899 Å) | | Cite: | Molecular basis for the unusual ring reconstruction in fungal meroterpenoid biogenesis
Nat. Chem. Biol., 13, 2017
|
|
5X9K
 
 | |
