1V2F
 | |
1V72
 
 | |
1VC3
 
 | |
1V2D
 
 | |
1V7C
 
 | |
1V2E
 
 | |
1VB3
 
 | |
1VCO
 
 | |
1VE4
 
 | |
1VCM
 
 | |
1VCN
 
 | |
1X2A
 
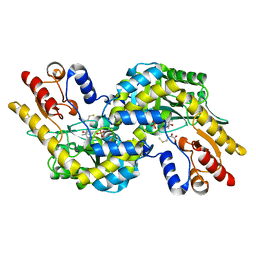
 | | Crystal Structure of e.coli AspAT complexed with N-phosphopyridoxyl-D-glutamic acid | | Descriptor: | Aspartate aminotransferase, N-({3-HYDROXY-2-METHYL-5-[(PHOSPHONOOXY)METHYL]PYRIDIN-4-YL}METHYL)-D-GLUTAMIC ACID | | Authors: | Goto, M. | | Deposit date: | 2005-04-21 | | Release date: | 2005-06-14 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Binding of C5-dicarboxylic substrate to aspartate aminotransferase: implications for the conformational change at the transaldimination step.
Biochemistry, 44, 2005
|
|
1X28
 | | Crystal Structure of e.coli AspAT complexed with N-phosphopyridoxyl-L-glutamic acid | | Descriptor: | Aspartate aminotransferase, N-({3-hydroxy-2-methyl-5-[(phosphonooxy)methyl]pyridin-4-yl}methyl)-L-glutamic acid | | Authors: | Goto, M. | | Deposit date: | 2005-04-21 | | Release date: | 2005-06-14 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Binding of C5-dicarboxylic substrate to aspartate aminotransferase: implications for the conformational change at the transaldimination step.
Biochemistry, 44, 2005
|
|
1WRV
 
 | |
1WLY
 
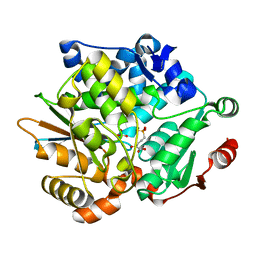
 | | Crystal Structure of 2-Haloacrylate Reductase | | Descriptor: | 2-haloacrylate reductase | | Authors: | Omi, R. | | Deposit date: | 2004-06-30 | | Release date: | 2005-10-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal Structure of 2-Haloacrylate Reductase
To be Published
|
|
1X29
 
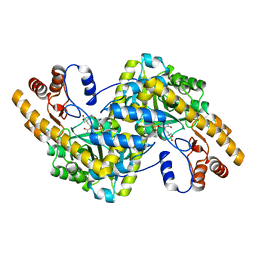 | | Crystal Structure of e.coli AspAT complexed with N-phosphopyridoxyl-2-methyl-L-glutamic acid | | Descriptor: | Aspartate aminotransferase, N-({3-HYDROXY-2-METHYL-5-[(PHOSPHONOOXY)METHYL]PYRIDIN-4-YL}METHYL)-2-METHYL-L-GLUTAMIC ACID | | Authors: | Goto, M. | | Deposit date: | 2005-04-21 | | Release date: | 2005-06-14 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Binding of C5-dicarboxylic substrate to aspartate aminotransferase: implications for the conformational change at the transaldimination step.
Biochemistry, 44, 2005
|
|
2COJ
 
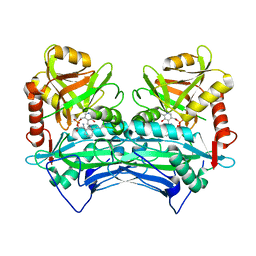 | |
2COG
 
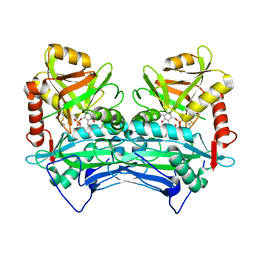 | |
2COI
 
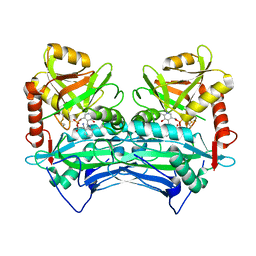 | |
2DVX
 | |
2DVT
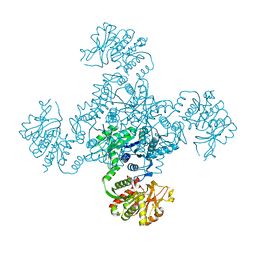
 | | Crystal Structure of 2,6-Dihydroxybenzoate Decarboxylase from Rhizobium | | Descriptor: | Thermophilic reversible gamma-resorcylate decarboxylase, ZINC ION | | Authors: | Goto, M. | | Deposit date: | 2006-08-01 | | Release date: | 2006-09-19 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structures of Nonoxidative Zn-Dependent 2,6-Dihydroxybenzoate (gamma-Resorcylate) Decarboxylase
from Rhizobium sp. Strain MTP-10005
To be Published
|
|
2DVU
 | |
2EEO
 
 | |
