4N9O
 
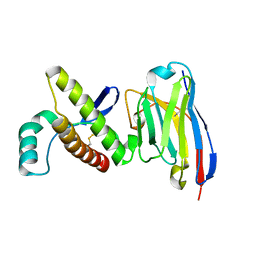 | | Probing the N-terminal beta-sheet conversion in the crystal structure of the human prion protein bound to a Nanobody | | Descriptor: | Major prion protein, Nanobody Nb484 | | Authors: | Abskharon, R.N.N, Giachin, G, Wohlkonig, A, Soror, S.H, Pardon, E, Legname, G, Steyaert, J. | | Deposit date: | 2013-10-21 | | Release date: | 2014-01-22 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Probing the N-Terminal beta-Sheet Conversion in the Crystal Structure of the Human Prion Protein Bound to a Nanobody.
J.Am.Chem.Soc., 136, 2014
|
|
5LKN
 
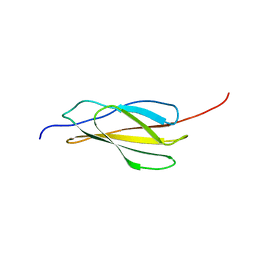 | | NMR solution structure of human FNIII domain 2 of NCAM | | Descriptor: | Neural cell adhesion molecule 1 | | Authors: | Slapsak, U, Salzano, G, Amin, L, Abskharon, R.N.N, Ilc, G, Zupancic, B, Biljan, I, Plavec, J, Giachin, G, Legname, G. | | Deposit date: | 2016-07-22 | | Release date: | 2016-09-14 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The N Terminus of the Prion Protein Mediates Functional Interactions with the Neuronal Cell Adhesion Molecule (NCAM) Fibronectin Domain.
J.Biol.Chem., 291, 2016
|
|
4KML
 
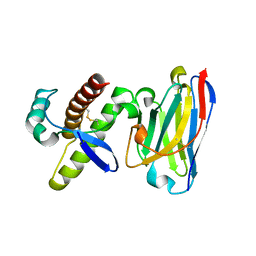 | | Probing the N-terminal beta-sheet conversion in the crystal structure of the full-length human prion protein bound to a Nanobody | | Descriptor: | Major prion protein, Nanobody | | Authors: | Abskharon, R.N.N, Giachin, G, Wohlkonig, A, Soror, S.H, Pardon, E, Legname, G, Steyaert, J. | | Deposit date: | 2013-05-08 | | Release date: | 2014-02-19 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Probing the N-Terminal beta-Sheet Conversion in the Crystal Structure of the Human Prion Protein Bound to a Nanobody.
J.Am.Chem.Soc., 136, 2014
|
|
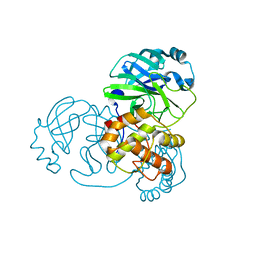
7NIJ
 
 | |
2LV1
 
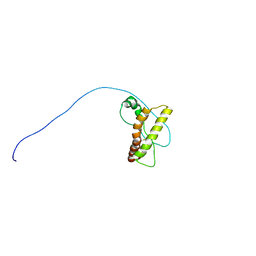 | | Solution-state NMR structure of prion protein mutant V210I at neutral pH | | Descriptor: | Major prion protein | | Authors: | Biljan, I, Ilc, G, Giachin, G, Legname, G, Plavec, J. | | Deposit date: | 2012-06-26 | | Release date: | 2012-09-19 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural Rearrangements at Physiological pH: Nuclear Magnetic Resonance Insights from the V210I Human Prion Protein Mutant.
Biochemistry, 51, 2012
|
|
2KUN
 
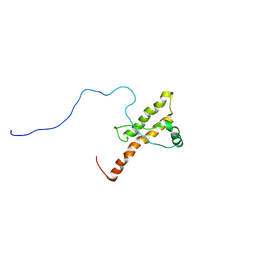 | | Three dimensional structure of HuPrP(90-231 M129 Q212P) | | Descriptor: | Major prion protein | | Authors: | Ilc, G, Giachin, G, Jaremko, M, Jaremko, L, Zhukov, I, Plavec, J, Legname, G, Benetti, F. | | Deposit date: | 2010-02-23 | | Release date: | 2010-08-25 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the human prion protein with the pathological Q212P mutation reveals unique structural features.
Plos One, 5, 2010
|
|
2LEJ
 
 | | human prion protein mutant HuPrP(90-231, M129, V210I) | | Descriptor: | Major prion protein | | Authors: | Biljan, I, Ilc, G, Giachin, G, Raspadori, A, Zhukov, I, Plavec, J, Legname, G. | | Deposit date: | 2011-06-16 | | Release date: | 2011-08-17 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Toward the Molecular Basis of Inherited Prion Diseases: NMR Structure of the Human Prion Protein with V210I Mutation.
J.Mol.Biol., 412, 2011
|
|
8PUM
 
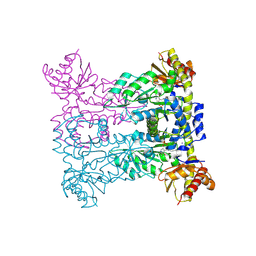 | |
8PUS
 
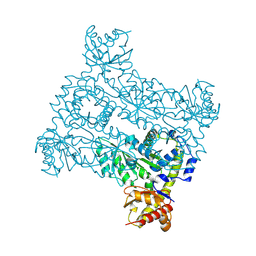 | |
6HEQ
 
 | | Prion nanobody 484 | | Descriptor: | Prion nanobody 484 | | Authors: | Soror, S.H, Abskharon, R.N, Wohlkonig, A. | | Deposit date: | 2018-08-20 | | Release date: | 2019-12-04 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.23 Å) | | Cite: | Structural evidence for the critical role of the prion protein hydrophobic region in forming an infectious prion.
Plos Pathog., 15, 2019
|
|
6HER
 
 | | Mouse prion protein in complex with Nanobody 484 | | Descriptor: | GLYCEROL, Major prion protein, Nanobody 484, ... | | Authors: | Soror, S.H, Abskharon, R, Wohlkonig, A. | | Deposit date: | 2018-08-20 | | Release date: | 2019-12-04 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.199 Å) | | Cite: | Structural evidence for the critical role of the prion protein hydrophobic region in forming an infectious prion.
Plos Pathog., 15, 2019
|
|
6HHD
 | | Mouse Prion Protein in complex with Nanobody 484 | | Descriptor: | CHLORIDE ION, GLYCEROL, Major prion protein, ... | | Authors: | Soror, S, Abskharon, R, Wohlkonig, A. | | Deposit date: | 2018-08-28 | | Release date: | 2019-12-11 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.102 Å) | | Cite: | Structural evidence for the critical role of the prion protein hydrophobic region in forming an infectious prion.
Plos Pathog., 15, 2019
|
|
7QFB
 
 | | Crystal structure of Protein Phosphatase 1 in complex with PP1-binding peptide from PTG | | Descriptor: | GLYCEROL, MANGANESE (II) ION, Protein phosphatase 1 regulatory subunit 3C, ... | | Authors: | Semrau, M.S, Storici, P, Lolli, G. | | Deposit date: | 2021-12-05 | | Release date: | 2022-11-02 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Molecular architecture of the glycogen- committed PP1/PTG holoenzyme.
Nat Commun, 13, 2022
|
|
7QFA
 
 | |
7QF7
 
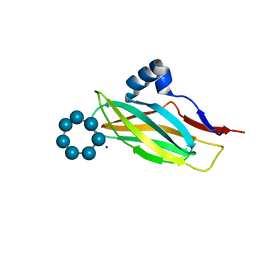 | |
7QM2
 
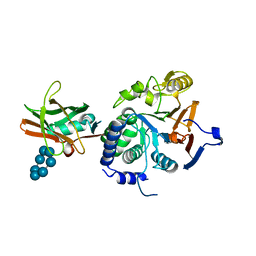 | |
2LFT
 
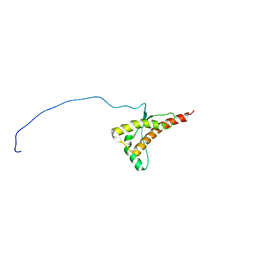 | | Human prion protein with E219K protective polymorphism | | Descriptor: | Major prion protein | | Authors: | Biljan, I, Ilc, G, Giacin, G, Zhukov, I, Plavec, J, Legname, G. | | Deposit date: | 2011-07-14 | | Release date: | 2012-06-27 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural basis for the protective effect of the human prion protein carrying the dominant-negative E219K polymorphism.
Biochem.J., 446, 2012
|
|
2LSB
 
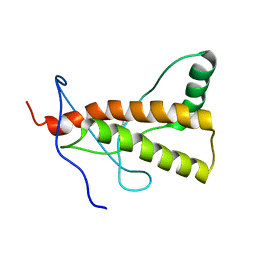 | | Solution-state NMR structure of the human prion protein | | Descriptor: | Major prion protein | | Authors: | Biljan, I, Ilc, G, Giancin, G, Zhukov, I, Plavec, J, Legname, G. | | Deposit date: | 2012-04-26 | | Release date: | 2012-06-27 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Structural basis for the protective effect of the human prion protein carrying the dominant-negative E219K polymorphism.
Biochem.J., 446, 2012
|
|
8PVZ
 
 | |
8PW0
 
 | |
8PVT
 
 | |
