6N9P
 
 | |
7KXO
 
 | | BTK1 SOAKED WITH COMPOUND 24 | | Descriptor: | 3-tert-butyl-N-[(1R)-6-{2-[5-methyl-1-(oxan-4-yl)-1H-pyrazol-4-yl]-3H-imidazo[4,5-b]pyridin-7-yl}-1,2,3,4-tetrahydronaphthalen-1-yl]-1,2,4-oxadiazole-5-carboxamide, DIMETHYL SULFOXIDE, Isoform BTK-C of Tyrosine-protein kinase BTK | | Authors: | Viacava Follis, A. | | Deposit date: | 2020-12-04 | | Release date: | 2021-05-19 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Discovery of potent and selective reversible Bruton's tyrosine kinase inhibitors.
Bioorg.Med.Chem., 40, 2021
|
|
7KXN
 
 | | BTK1 SOAKED WITH COMPOUND 26 | | Descriptor: | 3-tert-butyl-N-[(1S)-6-{2-[5-methyl-1-(propan-2-yl)-1H-pyrazol-4-yl]-1H-imidazo[4,5-b]pyridin-7-yl}-1,2,3,4-tetrahydronaphthalen-1-yl]-1,2,4-oxadiazole-5-carboxamide, DIMETHYL SULFOXIDE, Isoform BTK-C of Tyrosine-protein kinase BTK | | Authors: | Viacava Follis, A. | | Deposit date: | 2020-12-04 | | Release date: | 2021-05-19 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.34 Å) | | Cite: | Discovery of potent and selective reversible Bruton's tyrosine kinase inhibitors.
Bioorg.Med.Chem., 40, 2021
|
|
7KXQ
 
 | | BTK1 SOAKED WITH COMPOUND 30 | | Descriptor: | 3-tert-butyl-N-[(5R)-2-{2-[3,5-dimethyl-1-(propan-2-yl)-1H-pyrazol-4-yl]-3H-imidazo[4,5-b]pyridin-7-yl}-6,7,8,9-tetrahydro-5H-benzo[7]annulen-5-yl]-1,2,4-oxadiazole-5-carboxamide, DIMETHYL SULFOXIDE, Isoform BTK-C of Tyrosine-protein kinase BTK | | Authors: | Viacava Follis, A. | | Deposit date: | 2020-12-04 | | Release date: | 2021-05-19 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Discovery of potent and selective reversible Bruton's tyrosine kinase inhibitors.
Bioorg.Med.Chem., 40, 2021
|
|
7KXP
 
 | | BTK1 SOAKED WITH COMPOUND 25 | | Descriptor: | 3-tert-butyl-N-[(1S)-6-{2-[3-methyl-1-(oxan-4-yl)-1H-pyrazol-4-yl]-1H-imidazo[4,5-b]pyridin-7-yl}-1,2,3,4-tetrahydronaphthalen-1-yl]-1,2,4-oxadiazole-5-carboxamide, DIMETHYL SULFOXIDE, Tyrosine-protein kinase BTK | | Authors: | Viacava Follis, A. | | Deposit date: | 2020-12-04 | | Release date: | 2021-05-19 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Discovery of potent and selective reversible Bruton's tyrosine kinase inhibitors.
Bioorg.Med.Chem., 40, 2021
|
|
4XUZ
 
 | | Structure of CTX-M-15 bound to RPX-7009 at 1.5 A | | Descriptor: | 1,2-ETHANEDIOL, Beta-lactamase, CHLORIDE ION, ... | | Authors: | Clifton, M.C, Gardberg, A. | | Deposit date: | 2015-01-26 | | Release date: | 2015-04-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Discovery of a Cyclic Boronic Acid beta-Lactamase Inhibitor (RPX7009) with Utility vs Class A Serine Carbapenemases.
J.Med.Chem., 58, 2015
|
|
5P9H
 
 | | BTK1 COCRYSTALLIZED WITH RN983 | | Descriptor: | 6-~{tert}-butyl-8-fluoranyl-2-[3-(hydroxymethyl)-4-[1-methyl-5-[[5-(1-methylpiperidin-4-yl)pyridin-2-yl]amino]-6-oxidanylidene-pyridazin-3-yl]pyridin-2-yl]phthalazin-1-one, Tyrosine-protein kinase BTK | | Authors: | Gardberg, A.S. | | Deposit date: | 2016-09-20 | | Release date: | 2017-05-24 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Ability of Bruton's Tyrosine Kinase Inhibitors to Sequester Y551 and Prevent Phosphorylation Determines Potency for Inhibition of Fc Receptor but not B-Cell Receptor Signaling.
Mol. Pharmacol., 91, 2017
|
|
5P9J
 
 | | BTK1 COCRYSTALLIZED WITH IBRUTINIB | | Descriptor: | 1-[(3~{R})-3-[4-azanyl-3-(4-phenoxyphenyl)pyrazolo[3,4-d]pyrimidin-1-yl]piperidin-1-yl]propan-1-one, Tyrosine-protein kinase BTK | | Authors: | Gardberg, A.S. | | Deposit date: | 2016-09-20 | | Release date: | 2017-05-24 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | Ability of Bruton's Tyrosine Kinase Inhibitors to Sequester Y551 and Prevent Phosphorylation Determines Potency for Inhibition of Fc Receptor but not B-Cell Receptor Signaling.
Mol. Pharmacol., 91, 2017
|
|
5P9L
 
 | | BTK1 IN COMPLEX WITH CC 292 | | Descriptor: | Tyrosine-protein kinase BTK, ~{N}-[3-[[5-fluoranyl-2-[[4-(2-methoxyethoxy)phenyl]amino]pyrimidin-4-yl]amino]phenyl]propanamide | | Authors: | Gardberg, A.S. | | Deposit date: | 2016-09-20 | | Release date: | 2017-05-24 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Ability of Bruton's Tyrosine Kinase Inhibitors to Sequester Y551 and Prevent Phosphorylation Determines Potency for Inhibition of Fc Receptor but not B-Cell Receptor Signaling.
Mol. Pharmacol., 91, 2017
|
|
5P9F
 
 | | BTK IN COMPLEX WITH GDC-0834 | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, N-{3-[6-({4-[(2R)-1,4-dimethyl-3-oxopiperazin-2-yl]phenyl}amino)-4-methyl-5-oxo-4,5-dihydropyrazin-2-yl]-2-methylphenyl }-4,5,6,7-tetrahydro-1-benzothiophene-2-carboxamide, ... | | Authors: | Gardberg, A.S. | | Deposit date: | 2016-09-20 | | Release date: | 2017-05-24 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | Ability of Bruton's Tyrosine Kinase Inhibitors to Sequester Y551 and Prevent Phosphorylation Determines Potency for Inhibition of Fc Receptor but not B-Cell Receptor Signaling.
Mol. Pharmacol., 91, 2017
|
|
5P9G
 
 | | Structure of BTK with RN486 | | Descriptor: | 6-cyclopropyl-8-fluoranyl-2-[2-(hydroxymethyl)-3-[1-methyl-5-[[5-(4-methylpiperazin-1-yl)pyridin-2-yl]amino]-6-oxidanylidene-pyridin-3-yl]phenyl]isoquinolin-1-one, Tyrosine-protein kinase BTK | | Authors: | Gardberg, A.S. | | Deposit date: | 2016-09-20 | | Release date: | 2017-05-24 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Ability of Bruton's Tyrosine Kinase Inhibitors to Sequester Y551 and Prevent Phosphorylation Determines Potency for Inhibition of Fc Receptor but not B-Cell Receptor Signaling.
Mol. Pharmacol., 91, 2017
|
|
5P9M
 
 | | BTK1 BINDS COVALENTLY TO HY-15771 ONO-4059 | | Descriptor: | 6-azanyl-9-[(3~{R})-1-[(~{E})-but-2-enoyl]pyrrolidin-3-yl]-7-(4-phenoxyphenyl)purin-8-one, DIMETHYL SULFOXIDE, Tyrosine-protein kinase BTK | | Authors: | Gardberg, A.S. | | Deposit date: | 2016-09-20 | | Release date: | 2017-05-24 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.41 Å) | | Cite: | Ability of Bruton's Tyrosine Kinase Inhibitors to Sequester Y551 and Prevent Phosphorylation Determines Potency for Inhibition of Fc Receptor but not B-Cell Receptor Signaling.
Mol. Pharmacol., 91, 2017
|
|
5P9I
 
 | | BTK1 SOAKED WITH IBRUTINIB-Rev | | Descriptor: | 1-{(3R)-3-[4-amino-3-(4-phenoxyphenyl)-1H-pyrazolo[3,4-d]pyrimidin-1-yl]piperidin-1-yl}prop-2-en-1-one, DIMETHYL SULFOXIDE, Tyrosine-protein kinase BTK | | Authors: | Gardberg, A.S. | | Deposit date: | 2016-09-20 | | Release date: | 2017-05-24 | | Last modified: | 2021-11-17 | | Method: | X-RAY DIFFRACTION (1.11 Å) | | Cite: | Ability of Bruton's Tyrosine Kinase Inhibitors to Sequester Y551 and Prevent Phosphorylation Determines Potency for Inhibition of Fc Receptor but not B-Cell Receptor Signaling.
Mol. Pharmacol., 91, 2017
|
|
5P9K
 
 | | CRYSTAL STRUCTURE OF BTK with CNX 774 | | Descriptor: | 4-[4-[[5-fluoranyl-4-[[3-(propanoylamino)phenyl]amino]pyrimidin-2-yl]amino]phenoxy]-~{N}-methyl-pyridine-2-carboxamide, Tyrosine-protein kinase BTK | | Authors: | Gardberg, A.S. | | Deposit date: | 2016-09-20 | | Release date: | 2017-05-24 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | Ability of Bruton's Tyrosine Kinase Inhibitors to Sequester Y551 and Prevent Phosphorylation Determines Potency for Inhibition of Fc Receptor but not B-Cell Receptor Signaling.
Mol. Pharmacol., 91, 2017
|
|
6DI1
 
 | | CRYSTAL STRUCTURE OF BTK IN COMPLEX WITH COVALENT FRAGMENT LIGAND | | Descriptor: | 4-amino-2-[(3S)-3-(propanoylamino)pyrrolidin-1-yl]pyrimidine-5-carboxamide, Tyrosine-protein kinase BTK | | Authors: | Jiang, X. | | Deposit date: | 2018-05-22 | | Release date: | 2018-10-17 | | Last modified: | 2018-10-24 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Discovery of a novel series of pyridine and pyrimidine carboxamides as potent and selective covalent inhibitors of Btk.
Bioorg. Med. Chem. Lett., 28, 2018
|
|
6DI0
 
 | | CRYSTAL STRUCTURE OF BTK IN COMPLEX WITH FRAGMENT LIGAND | | Descriptor: | 6-(cyclohexylamino)pyridine-3-carboxamide, Tyrosine-protein kinase BTK | | Authors: | MOCHALKIN, I. | | Deposit date: | 2018-05-22 | | Release date: | 2018-10-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Discovery of a novel series of pyridine and pyrimidine carboxamides as potent and selective covalent inhibitors of Btk.
Bioorg. Med. Chem. Lett., 28, 2018
|
|
4F83
 
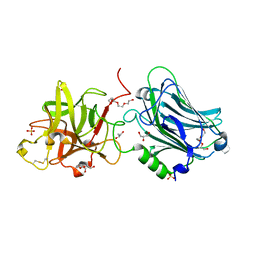 | | Crystal structure of the receptor binding domain of botulinum neurotoxin mosaic serotype C/D with a tetraethylene glycol molecule bound on the Hcn sub-domain and a sulfate ion at the putative active site | | Descriptor: | GLYCEROL, SULFATE ION, TETRAETHYLENE GLYCOL, ... | | Authors: | Zhang, Y, Buchko, G.W, Gardberg, A, Edwards, T.E, Sankaran, B, Robinson, H, Varnum, S.M, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2012-05-16 | | Release date: | 2012-06-20 | | Last modified: | 2013-06-12 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural insights into the functional role of the Hcn sub-domain of the receptor-binding domain of the botulinum neurotoxin mosaic serotype C/D.
Biochimie, 95, 2013
|
|
4ED9
 
 | |
3S6O
 
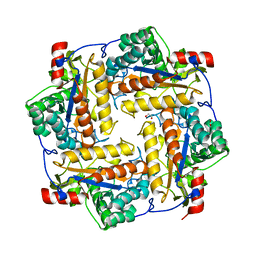 | |
3SC4
 
 | |
3SIB
 
 | |
3SIA
 
 | |
3SJS
 
 | |
3SW5
 
 | |
3TK8
 
 | | Structure of a 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase from Burkholderia pseudomallei | | Descriptor: | 1,2-ETHANEDIOL, 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase, SULFATE ION, ... | | Authors: | Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2011-08-25 | | Release date: | 2011-10-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of a 2,3,4,5-tetrahydropyridine-2,6-dicarboxylate N-succinyltransferase from Burkholderia pseudomallei
TO BE PUBLISHED
|
|
