7XGV
 
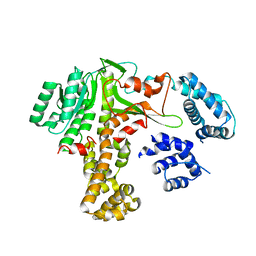 | | Legionella glucosyltransferase | | Descriptor: | Lgt2 | | Authors: | Chen, T.T, Ouyang, S.Y. | | Deposit date: | 2022-04-06 | | Release date: | 2023-04-12 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Structural Basis for the Action Mechanism of Legionella Glycosyltransferase.
Small Struct, 2023
|
|
7XGX
 
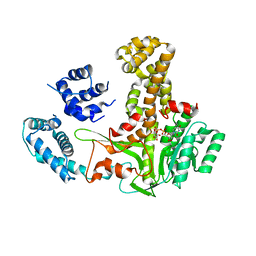 | | Glucosyltransferase | | Descriptor: | Lgt2, URIDINE-5'-DIPHOSPHATE-GLUCOSE | | Authors: | Chen, T.T, Ouyang, S.Y. | | Deposit date: | 2022-04-07 | | Release date: | 2023-04-12 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Structural Basis for the Action Mechanism of Legionella Glycosyltransferase.
Small Struct, 2023
|
|
5HQ6
 
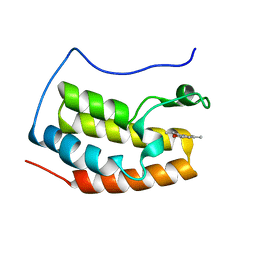 | | Crystal structure of fragment bound with Brd4 | | Descriptor: | (2R)-2,6-dimethyl-2H-1,4-benzoxazin-3(4H)-one, Bromodomain-containing protein 4 | | Authors: | Chen, T.T, Xu, Y.C. | | Deposit date: | 2016-01-21 | | Release date: | 2017-01-25 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of fragment bound with Brd4
to be published
|
|
5HQ7
 
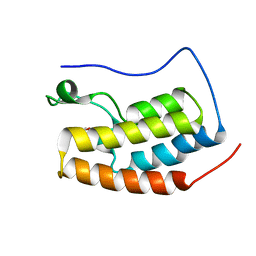 | | Crystal structure of fragment bound with Brd4 | | Descriptor: | Bromodomain-containing protein 4, N-ethyl-6,7-dimethoxyquinazolin-4-amine | | Authors: | Chen, T.T, Xu, Y.C. | | Deposit date: | 2016-01-21 | | Release date: | 2017-01-25 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of fragment bound with Brd4
to be published
|
|
5HQ5
 
 | | Crystal structure of fragment bound with Brd4 | | Descriptor: | 7-methyl-N-(o-tolyl)-[1,2,4]triazolo[4,3-a]pyrimidin-5-amine, Bromodomain-containing protein 4 | | Authors: | Chen, T.T, Xu, Y.C. | | Deposit date: | 2016-01-21 | | Release date: | 2017-01-25 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of fragment bound with Brd4
to be published
|
|
7XQL
 
 | | complex structure of LegA15 with GNP | | Descriptor: | Ankyrin repeat-containing protein, PHOSPHOAMINOPHOSPHONIC ACID-GUANYLATE ESTER | | Authors: | Chen, T.T, Lin, Y.L. | | Deposit date: | 2022-05-07 | | Release date: | 2023-05-17 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.272 Å) | | Cite: | Atypical Legionella GTPase effector hijacks host vesicular transport factor p115 to regulate host lipid droplet.
Sci Adv, 8, 2022
|
|
8K6I
 
 | | LnaB-Actin-PRUb ternary complex | | Descriptor: | Actin gamma 1, Legionella effector LnaB, PHOSPHATE ION, ... | | Authors: | Chen, T.T, Ouyang, S.Y. | | Deposit date: | 2023-07-25 | | Release date: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (3.19 Å) | | Cite: | Complex structure of Legionella effector LnaB with host Actin and PR-Ub
To Be Published
|
|
8K6F
 
 | | LnaB-Actin-PRUb ternary complex | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Actin gamma 1, Legionella effector LnaB, ... | | Authors: | Chen, T.T, Ouyang, S.Y. | | Deposit date: | 2023-07-25 | | Release date: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (3.41 Å) | | Cite: | Complex structure of Legionella effector LnaB with host Actin and PR-Ub
To Be Published
|
|
8K6R
 
 | |
8K6V
 
 | | LnaB-Actin-PRUb ternary complex | | Descriptor: | Actin gamma 1, LnaB, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Chen, T.T, Ouyang, S.Y. | | Deposit date: | 2023-07-25 | | Release date: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Complex structure of Legionella effector LnaB with host Actin and ADPR-Ub
To Be Published
|
|
8K6E
 
 | | LnaB-Actin-PRUb ternary complex | | Descriptor: | Actin gamma 1, Legionella effector LnaB, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, ... | | Authors: | Chen, T.T, Ouyang, S.Y. | | Deposit date: | 2023-07-25 | | Release date: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (2.74 Å) | | Cite: | Complex structure of Legionella effector LnaB with host Actin and PR-Ub
To Be Published
|
|
8HCY
 
 | | Legionella glycosyltransferase | | Descriptor: | Dot/Icm secretion system substrate | | Authors: | Chen, T.T, Ouyang, S.Y. | | Deposit date: | 2022-11-03 | | Release date: | 2023-11-08 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Structural Basis for the Action Mechanism of Legionella Glycosyltransferase.
Small Struct, 2023
|
|
7WX6
 
 | | A Legionella acetyltransferase VipF | | Descriptor: | CHLORAMPHENICOL, COENZYME A, N-acetyltransferase | | Authors: | Chen, T.T, Lin, Y.L, Chen, Z, Han, A.D. | | Deposit date: | 2022-02-14 | | Release date: | 2022-09-14 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.273 Å) | | Cite: | Structural basis for the acetylation mechanism of the Legionella effector VipF.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
7WX7
 
 | | complex of a legionella acetyltransferase VipF and COA/ACO | | Descriptor: | ACETYL COENZYME *A, COENZYME A, N-acetyltransferase | | Authors: | Chen, T.T, Lin, Y.L, Zhang, S.J, Han, A.D. | | Deposit date: | 2022-02-14 | | Release date: | 2023-02-22 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.781 Å) | | Cite: | Structural basis for the acetylation mechanism of the Legionella effector VipF.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
7WX5
 
 | | a Legionella acetyltransferase effector VipF | | Descriptor: | ACETYL COENZYME *A, N-acetyltransferase | | Authors: | Chen, T.T, Lin, Y.L, Zhang, S.J, Han, A.D. | | Deposit date: | 2022-02-14 | | Release date: | 2023-02-22 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.392 Å) | | Cite: | Structural basis for the acetylation mechanism of the Legionella effector VipF.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
7EW8
 
 | | Legionella pneumophila effector AnkD | | Descriptor: | ANK_REP_REGION domain-containing protein | | Authors: | Chen, T.T, Lin, Y.L. | | Deposit date: | 2021-05-24 | | Release date: | 2022-06-01 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.594 Å) | | Cite: | Atypical Legionella GTPase effector hijacks host vesicular transport factor p115 to regulate host lipid droplet.
Sci Adv, 8, 2022
|
|
3UQP
 
 | | Crystal structure of Bace1 with its inhibitor | | Descriptor: | Beta-secretase 1, METHYL (2R)-1-[(6S,9S,12S,13S,17S,20S,23R)-9-(3-AMINO-3-OXOPROPYL)-12,23-DIBENZYL-13-HYDROXY-2,2,8,20,22-PENTAMETHYL-17-(2-METHYLPROPYL)-4,7,10,15,18,21,24-HEPTAOXO-6-(PROPAN-2-YL)-3-OXA-5,8,11,16,19,22-HEXAAZATETRACOSAN-24-YL]PYRROLIDINE-2-CARBOXYLATE, SULFATE ION | | Authors: | Chen, T.T, Chen, W.Y, Li, L, Xu, Y.C. | | Deposit date: | 2011-11-21 | | Release date: | 2012-11-21 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Cyanobacterial Peptides as a Prototype for the Design of Potent beta-Secretase Inhibitors and the Development of Selective Chemical Probes for Other Aspartic Proteases
J.Med.Chem., 55, 2012
|
|
3UQX
 
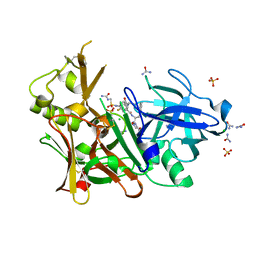 | | Crystal structure of BACE1 with its inhibitor | | Descriptor: | Beta-secretase 1, CHLORIDE ION, N-[(1R)-1-(4-fluorophenyl)ethyl]-N'-[(2S,3S)-3-hydroxy-4-{4-[(1S)-1-hydroxyethyl]-1H-1,2,3-triazol-1-yl}-1-phenylbutan-2-yl]-5-[methyl(methylsulfonyl)amino]benzene-1,3-dicarboxamide, ... | | Authors: | Chen, T.T, Chen, W.Y, Xu, Y.C. | | Deposit date: | 2011-11-21 | | Release date: | 2012-11-21 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Flexibility of the Flap in the Active Site of BACE1 as Revealed by Crystal Structures and MD simulations
to be published
|
|
4FCO
 
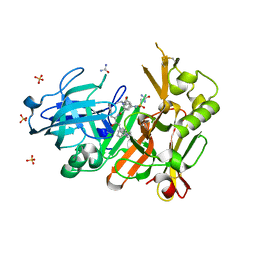 | | Crystal structure of bace1 with its inhibitor | | Descriptor: | Beta-secretase 1, N-[(2S,3R)-4-{[2-(1-benzylpiperidin-4-yl)ethyl]amino}-3-hydroxy-1-phenylbutan-2-yl]-5-[methyl(methylsulfonyl)amino]-N'-[(1R)-1-phenylethyl]benzene-1,3-dicarboxamide, SULFATE ION, ... | | Authors: | Chen, T.T, Chen, W.Y, Li, L, Xu, Y.C. | | Deposit date: | 2012-05-25 | | Release date: | 2013-05-29 | | Last modified: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Flexibility of the Flap in the Active Site of BACE1 as Revealed by Crystal Structures and MD simulations
To be Published, 2012
|
|
3SHY
 
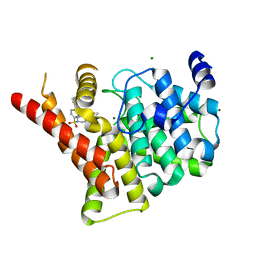 | | Crystal structure of the PDE5A1 catalytic domain in complex with novel inhibitors | | Descriptor: | 6-ethyl-5-fluoro-2-{5-[(4-methylpiperazin-1-yl)sulfonyl]-2-propoxyphenyl}pyrimidin-4(3H)-one, MAGNESIUM ION, ZINC ION, ... | | Authors: | Chen, T.T, Chen, T, Xu, Y.C. | | Deposit date: | 2011-06-17 | | Release date: | 2011-08-24 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.647 Å) | | Cite: | Utilization of Halogen Bond in Lead Optimization: A Case Study of Rational Design of Potent Phosphodiesterase Type 5 (PDE5) Inhibitors.
J.Med.Chem., 54, 2011
|
|
3SIE
 
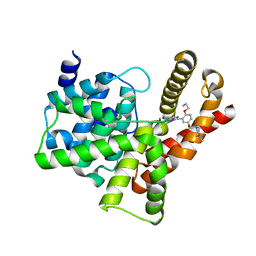 | | Crystal structure of the PDE5A1 catalytic domain in complex with novel inhibitors | | Descriptor: | 5-bromo-6-ethyl-2-{5-[(4-methylpiperazin-1-yl)sulfonyl]-2-propoxyphenyl}pyrimidin-4(3H)-one, cGMP-specific 3',5'-cyclic phosphodiesterase | | Authors: | Chen, T.T, Chen, T, Xu, Y.C. | | Deposit date: | 2011-06-17 | | Release date: | 2011-08-24 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Utilization of Halogen Bond in Lead Optimization: A Case Study of Rational Design of Potent Phosphodiesterase Type 5 (PDE5) Inhibitors.
J.Med.Chem., 54, 2011
|
|
3SHZ
 
 | | Crystal structure of the PDE5A1 catalytic domain in complex with novel inhibitors | | Descriptor: | 5-chloro-6-ethyl-2-{5-[(4-methylpiperazin-1-yl)sulfonyl]-2-propoxyphenyl}pyrimidin-4(3H)-one, MAGNESIUM ION, ZINC ION, ... | | Authors: | Chen, T.T, Chen, T, Xu, Y.C. | | Deposit date: | 2011-06-17 | | Release date: | 2011-08-24 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.449 Å) | | Cite: | Utilization of Halogen Bond in Lead Optimization: A Case Study of Rational Design of Potent Phosphodiesterase Type 5 (PDE5) Inhibitors.
J.Med.Chem., 54, 2011
|
|
3UQR
 
 | | Crystal structure of BACE1 with its inhibitor | | Descriptor: | Beta-secretase 1, METHYL (2S)-1-[(2R,5S,8S,12S,13S)-2,13-DIBENZYL-12-HYDROXY-3,5-DIMETHYL-15-(3-[METHYL(METHYLSULFONYL)AMINO]-5-{[(1R)-1-PHENYLETHYL]CARBAMOYL}PHENYL)-8-(2-METHYLPROPYL)-4,7,10,15-TETRAOXO-3,6,9,14-TETRAAZAPENTADECAN-1-OYL]PYRROLIDINE-2-CARBOXYLATE | | Authors: | Chen, T.T, Chen, W.Y, Li, L, Xu, Y.C. | | Deposit date: | 2011-11-21 | | Release date: | 2012-11-21 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (3.056 Å) | | Cite: | Cyanobacterial Peptides as a Prototype for the Design of Potent beta-Secretase Inhibitors and the Development of Selective Chemical Probes for Other Aspartic Proteases
J.Med.Chem., 55, 2012
|
|
3UQW
 
 | | Crystal structure of BACE1 with its inhibitor | | Descriptor: | Beta-secretase 1, SULFATE ION, ethyl 1-{(2S,3S)-3-[(3-{[(1R)-1-(4-fluorophenyl)ethyl]carbamoyl}-5-[methyl(methylsulfonyl)amino]benzoyl)amino]-2-hydroxy-4-phenylbutyl}-1H-pyrazole-4-carboxylate | | Authors: | Chen, T.T, Chen, W.Y, Xu, Y.C. | | Deposit date: | 2011-11-21 | | Release date: | 2012-11-21 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Flexibility of the Flap in the Active Site of BACE1 as Revealed by Crystal Structures and MD simulations
to be published
|
|
3UQU
 
 | | Crystal structure of BACE1 with its inhibitor | | Descriptor: | Beta-secretase 1, CHLORIDE ION, N-[(1R)-1-(4-fluorophenyl)ethyl]-N'-[(2S,3S)-3-hydroxy-1-phenyl-4-(1H-pyrazol-1-yl)butan-2-yl]-5-[methyl(methylsulfonyl)amino]benzene-1,3-dicarboxamide, ... | | Authors: | Chen, T.T, Chen, W.Y, Xu, Y.C. | | Deposit date: | 2011-11-21 | | Release date: | 2012-11-21 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Flexibility of the Flap in the Active Site of BACE1 as Revealed by Crystal Structures and MD simulations
to be published
|
|
