7XMP
 
 | |
5E68
 
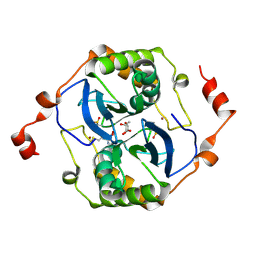 | | High resolution crystal structure of LuxS - Quorum sensor molecular complex from Salmonella typhi at 1.58 Angstroms | | Descriptor: | (2R,4S)-2-methyl-2,3,3,4-tetrahydroxytetrahydrofuran, METHIONINE, S-ribosylhomocysteine lyase, ... | | Authors: | Perumal, P, Raina, R, Arockiasamy, A, SundaraBaalaji, N. | | Deposit date: | 2015-10-09 | | Release date: | 2016-10-12 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | High resolution crystal structure of LuxS - Quorum sensor molecular complex from Salmonella typhi at 1.58 Angstroms
To Be Published
|
|
7W5D
 
 | |
5IQY
 
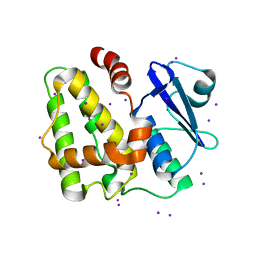 | | Structure of apo-Dehydroascorbate Reductase from Pennisetum Glaucum phased by Iodide-SAD method | | Descriptor: | Dehydroascorbate reductase, IODIDE ION | | Authors: | Das, B.K, Kumar, A, Manidola, P, Arockiasamy, A. | | Deposit date: | 2016-03-11 | | Release date: | 2016-05-04 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Non-native ligands define the active site of Pennisetum glaucum (L.) R. Br dehydroascorbate reductase
Biochem.Biophys.Res.Commun., 473, 2016
|
|
1NL3
 
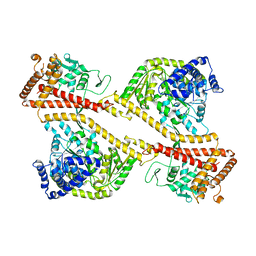 | | CRYSTAL STRUCTURE OF THE SECA PROTEIN TRANSLOCATION ATPASE FROM MYCOBACTERIUM TUBERCULOSIS in APO FORM | | Descriptor: | PREPROTEIN TRANSLOCASE SECA 1 SUBUNIT | | Authors: | Sharma, V, Arockiasamy, A, Ronning, D.R, Savva, C.G, Holzenburg, A, Braunstein, M, Jacobs Jr, W.R, Sacchettini, J.C, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2003-01-06 | | Release date: | 2003-03-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structure of M. tuberculosis SecA, A Preprotein Translocating ATPase
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
1NKT
 
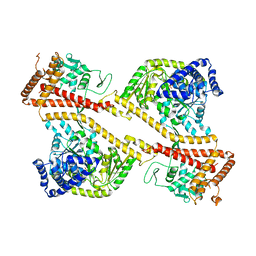 | | CRYSTAL STRUCTURE OF THE SECA PROTEIN TRANSLOCATION ATPASE FROM MYCOBACTERIUM TUBERCULOSIS COMPLEX WITH ADPBS | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, MAGNESIUM ION, Preprotein translocase secA 1 subunit | | Authors: | Sharma, V, Arockiasamy, A, Ronning, D.R, Savva, C.G, Holzenburg, A, Braunstein, M, Jacobs Jr, W.R, Sacchettini, J.C, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2003-01-03 | | Release date: | 2003-03-04 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.601 Å) | | Cite: | Crystal Structure of M. tuberculosis SecA, A Preprotein Translocating ATPase
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
4ZPU
 
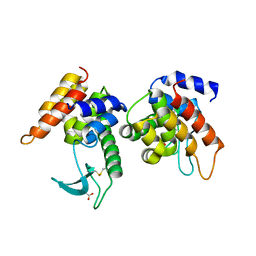 | |
3HDE
 
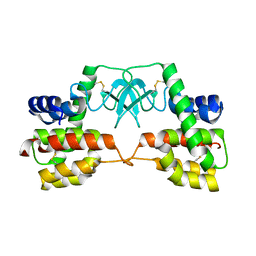 | | Crystal structure of full-length endolysin R21 from phage 21 | | Descriptor: | Lysozyme | | Authors: | Sun, Q, Arockiasamy, A, McKee, E, Caronna, E, Sacchettini, J.C. | | Deposit date: | 2009-05-07 | | Release date: | 2009-11-03 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Regulation of a muralytic enzyme by dynamic membrane topology.
Nat.Struct.Mol.Biol., 16, 2009
|
|
5EVO
 
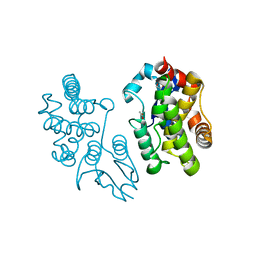 | | Structure of Dehydroascrobate Reductase from Pennisetum Americanum in complex with two non-native ligands, Acetate in the G-site and Glycerol in the H-site | | Descriptor: | ACETATE ION, Dehydroascorbate reductase, GLYCEROL | | Authors: | Kumar, A.O, Das, B.K, Arockiasamy, A. | | Deposit date: | 2015-11-20 | | Release date: | 2016-05-04 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | Non-native ligands define the active site of Pennisetum glaucum (L.) R. Br dehydroascorbate reductase.
Biochem.Biophys.Res.Commun., 473, 2016
|
|
7F8S
 
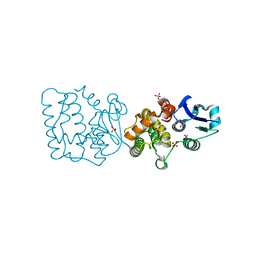 | | Pennisetum glaucum (Pearl millet) dehydroascorbate reductase (DHAR) with catalytic cysteine (Cy20) in sulphenic and sulfinic acid forms. | | Descriptor: | Dehydroascorbate reductase, SULFATE ION | | Authors: | Das, B.K, Kumar, A, Sreeshma, N.S, Arockiasamy, A. | | Deposit date: | 2021-07-02 | | Release date: | 2022-01-19 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Comparative kinetic analysis of ascorbate (Vitamin-C) recycling dehydroascorbate reductases from plants and humans.
Biochem.Biophys.Res.Commun., 591, 2021
|
|
7DK1
 
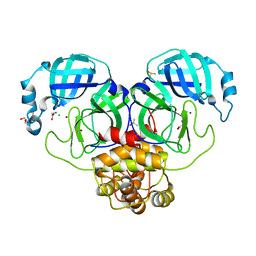 | | Crystal structure of Zinc bound SARS-CoV-2 main protease | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 3C-like proteinase, CHLORIDE ION, ... | | Authors: | Sonkar, K.S, Panchariya, L, Kuila, S, Khan, W.A, Arockiasamy, A. | | Deposit date: | 2020-11-22 | | Release date: | 2021-06-30 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.902 Å) | | Cite: | Zinc 2+ ion inhibits SARS-CoV-2 main protease and viral replication in vitro.
Chem.Commun.(Camb.), 57, 2021
|
|
5GTA
 
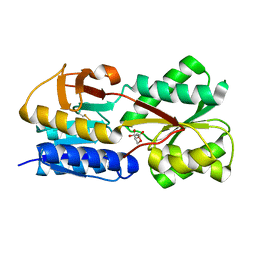 | | 3D Crystal Structure of LsrB Bound to Furanosyl diester (R)-THMF, from Salmonella typhi | | Descriptor: | (2R,4S)-2-methyl-2,3,3,4-tetrahydroxytetrahydrofuran, Autoinducer 2-binding protein LsrB | | Authors: | Gopinath, S, Perumal, P, Rahul, R, Arockiasamy, A, SundaraBaalaji, N. | | Deposit date: | 2016-08-19 | | Release date: | 2017-08-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.998 Å) | | Cite: | 3D Crystal Structure of LsrB Bound to Furanosyl diester (R)-THMF, from Salmonella typhi
To Be Published
|
|
3NSG
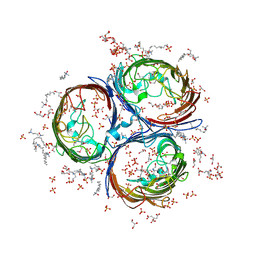 
 | | Crystal Structure of OmpF, an Outer Membrane Protein from Salmonella typhi | | Descriptor: | CITRATE ANION, GLYCEROL, L(+)-TARTARIC ACID, ... | | Authors: | Balasubramaniam, D, Arockiasamy, A, Sharma, A, Krishnaswamy, S. | | Deposit date: | 2010-07-01 | | Release date: | 2011-07-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | Asymmetric pore occupancy in crystal structure of OmpF porin from Salmonella typhi
J.Struct.Biol., 178, 2012
|
|
3UPG
 
 | |
3UU2
 
 | |
3OSX
 
 | |
7D9L
 
 | |
7F8R
 
 | |
7DKP
 
 | |
7DKR
 
 | |
7EZZ
 
 | | Crystal structure of Salmonella typhi outer membrane phospholipase (OMPLA) dimer with bound calcium | | Descriptor: | CALCIUM ION, DODECANE, Phospholipase A1, ... | | Authors: | Perumal, P, Raina, R, Sreeshma, N.S, Arockiasamy, A, Sundarabaalaji, N. | | Deposit date: | 2021-06-02 | | Release date: | 2021-06-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.76 Å) | | Cite: | Crystal structure of Salmonella typhi outer membrane phospholipase (OMPLA) dimer with bound calcium
To Be Published
|
|
3ONR
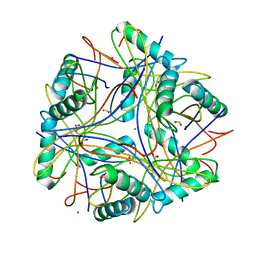 
 | | Crystal structure of the calcium chelating immunodominant antigen, calcium dodecin (Rv0379),from Mycobacterium tuberculosis with a novel calcium-binding site | | Descriptor: | CALCIUM ION, FORMIC ACID, PLATINUM (II) ION, ... | | Authors: | Arulandu, A, Sacchettini, J.C. | | Deposit date: | 2010-08-30 | | Release date: | 2011-03-16 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of calcium dodecin (Rv0379), from Mycobacterium tuberculosis with a unique calcium-binding site.
Protein Sci., 20, 2011
|
|
1XJU
 
 | |
1XJT
 
 | |
4O7K
 
 | | Crystal structure of Oncogenic Suppression Activity Protein - A Plasmid Fertility Inhibition Factor | | Descriptor: | CHLORIDE ION, ISOPROPYL ALCOHOL, PHOSPHATE ION, ... | | Authors: | Maindola, P, Goyal, P, Arulandu, A. | | Deposit date: | 2013-12-25 | | Release date: | 2014-11-05 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.748 Å) | | Cite: | Multiple enzymatic activities of ParB/Srx superfamily mediate sexual conflict among conjugative plasmids
Nat Commun, 5, 2014
|
|
