7S7V
 
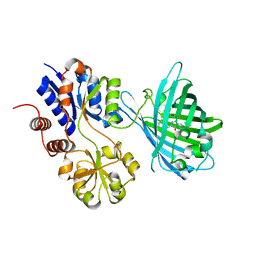 | | Crystal structure of iNicSnFR3a Fluorescent Nicotine Sensor | | Descriptor: | iNicSnFR 3.0 Fluorescent Nicotine Sensor | | Authors: | Fan, C, Shivange, A.V, Looger, L.L, Lester, H.A, Rees, D.C. | | Deposit date: | 2021-09-17 | | Release date: | 2021-10-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Correction: Fluorescence activation mechanism and imaging of drug permeation with new sensors for smoking-cessation ligands.
Elife, 11, 2022
|
|
7S7T
 
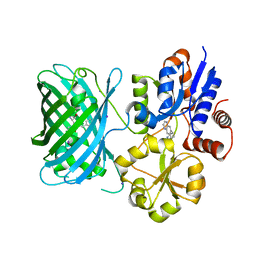 | | iNicSnFR3a Nicotine Sensor comprising Periplasmic Binding sequence plus Fluorescent Sequence with varenicline bound | | Descriptor: | IODIDE ION, VARENICLINE, iNicSnFR 3.0 Fluorescent Nicotine Sensor | | Authors: | Fan, C, Shivange, A.V, Looger, L.L, Lester, H.A, Rees, D.C. | | Deposit date: | 2021-09-17 | | Release date: | 2021-10-13 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Correction: Fluorescence activation mechanism and imaging of drug permeation with new sensors for smoking-cessation ligands.
Elife, 11, 2022
|
|
6V1R
 
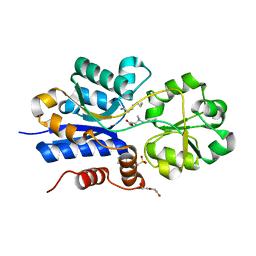 | | Crystal structure of iAChSnFR Fluorescent Acetylcholine Sensor precursor binding protein | | Descriptor: | 2-[N-CYCLOHEXYLAMINO]ETHANE SULFONIC ACID, ACETYLCHOLINE, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Fan, C, Borden, P.M, Looger, L.L, Lester, H.A, Rees, D.C. | | Deposit date: | 2019-11-21 | | Release date: | 2020-11-25 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | A genetically encoded fluorescent sensor for in vivo acetylcholine detection
To Be Published
|
|
7N59
 
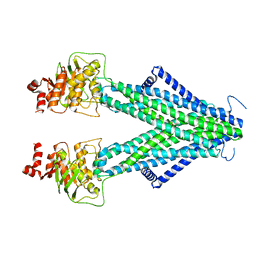 | | Structure of AtAtm3 in the inward-facing conformation with GSSG bound | | Descriptor: | ABC transporter B family member 25, mitochondrial, OXIDIZED GLUTATHIONE DISULFIDE | | Authors: | Fan, C, Rees, D.C. | | Deposit date: | 2021-06-05 | | Release date: | 2022-04-13 | | Last modified: | 2022-04-20 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Glutathione binding to the plant At Atm3 transporter and implications for the conformational coupling of ABC transporters.
Elife, 11, 2022
|
|
2MIN
 
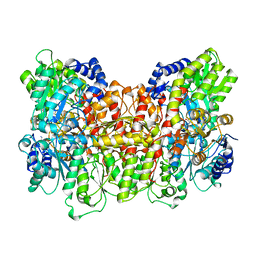 | | NITROGENASE MOFE PROTEIN FROM AZOTOBACTER VINELANDII, OXIDIZED STATE | | Descriptor: | 3-HYDROXY-3-CARBOXY-ADIPIC ACID, CALCIUM ION, FE(8)-S(7) CLUSTER, ... | | Authors: | Peters, J.W, Stowell, M.H.B, Soltis, S.M, Day, M.W, Kim, J, Rees, D.C. | | Deposit date: | 1996-12-20 | | Release date: | 1997-04-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Redox-dependent structural changes in the nitrogenase P-cluster.
Biochemistry, 36, 1997
|
|
6WA8
 
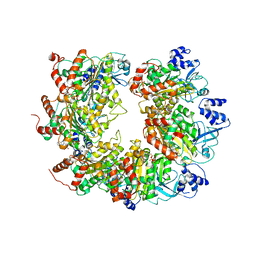 | |
6VQU
 
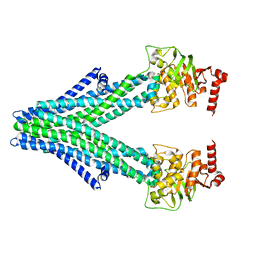 | | Structure of a bacterial Atm1-family ABC exporter | | Descriptor: | ATM1-type heavy metal exporter | | Authors: | Fan, C, Rees, D.C. | | Deposit date: | 2020-02-05 | | Release date: | 2020-07-29 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (3.88 Å) | | Cite: | A structural framework for unidirectional transport by a bacterial ABC exporter.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
6VQT
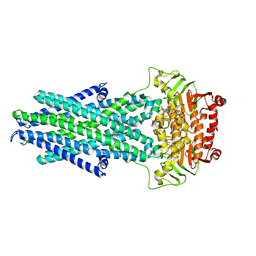 
 | |
1AMF
 
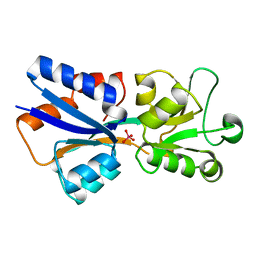 | | CRYSTAL STRUCTURE OF MODA, A MOLYBDATE TRANSPORT PROTEIN, COMPLEXED WITH MOLYBDATE | | Descriptor: | MOLYBDATE ION, MOLYBDATE TRANSPORT PROTEIN MODA | | Authors: | Hu, Y, Rech, S, Gunsalus, R.P, Rees, D.C. | | Deposit date: | 1997-06-13 | | Release date: | 1997-12-24 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structure of the molybdate binding protein ModA.
Nat.Struct.Biol., 4, 1997
|
|
1AG0
 
 | |
1B25
 
 | | FORMALDEHYDE FERREDOXIN OXIDOREDUCTASE FROM PYROCOCCUS FURIOSUS | | Descriptor: | IRON/SULFUR CLUSTER, PROTEIN (FORMALDEHYDE FERREDOXIN OXIDOREDUCTASE), TUNGSTOPTERIN | | Authors: | Hu, Y.L, Faham, S, Roy, R, Adams, M.W.W, Rees, D.C. | | Deposit date: | 1998-12-04 | | Release date: | 1999-03-24 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Formaldehyde ferredoxin oxidoreductase from Pyrococcus furiosus: the 1.85 A resolution crystal structure and its mechanistic implications.
J.Mol.Biol., 286, 1999
|
|
1B4N
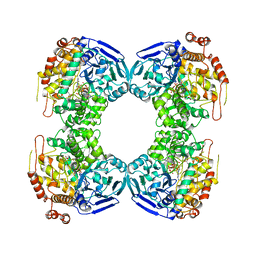 
 | | FORMALDEHYDE FERREDOXIN OXIDOREDUCTASE FROM PYROCOCCUS FURIOSUS, COMPLEXED WITH GLUTARATE | | Descriptor: | CALCIUM ION, FORMALDEHYDE FERREDOXIN OXIDOREDUCTASE, GLUTARIC ACID, ... | | Authors: | Hu, Y.L, Faham, S, Roy, R, Adams, M.W.W, Rees, D.C. | | Deposit date: | 1998-12-24 | | Release date: | 1999-05-18 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Formaldehyde ferredoxin oxidoreductase from Pyrococcus furiosus: the 1.85 A resolution crystal structure and its mechanistic implications.
J.Mol.Biol., 286, 1999
|
|
5VPW
 
 | | Nitrogenase Cp1 at pH 5 | | Descriptor: | 3-HYDROXY-3-CARBOXY-ADIPIC ACID, FE (II) ION, FE(8)-S(7) CLUSTER, ... | | Authors: | Morrison, C.N, Spatzal, T, Rees, D.C. | | Deposit date: | 2017-05-05 | | Release date: | 2017-07-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Reversible Protonated Resting State of the Nitrogenase Active Site.
J. Am. Chem. Soc., 139, 2017
|
|
5VQ4
 
 | | Nitrogenase Av1 at pH 5 | | Descriptor: | 3-HYDROXY-3-CARBOXY-ADIPIC ACID, FE (III) ION, FE(8)-S(7) CLUSTER, ... | | Authors: | Morrison, C.N, Spatzal, T, Rees, D.C. | | Deposit date: | 2017-05-07 | | Release date: | 2017-07-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Reversible Protonated Resting State of the Nitrogenase Active Site.
J. Am. Chem. Soc., 139, 2017
|
|
5VQ3
 
 | | Nitrogenase Cp1 at pH 6.5 | | Descriptor: | 3-HYDROXY-3-CARBOXY-ADIPIC ACID, FE (II) ION, FE(8)-S(7) CLUSTER, ... | | Authors: | Morrison, C.N, Spatzal, T, Rees, D.C. | | Deposit date: | 2017-05-07 | | Release date: | 2017-07-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Reversible Protonated Resting State of the Nitrogenase Active Site.
J. Am. Chem. Soc., 139, 2017
|
|
1BAR
 
 | | THREE-DIMENSIONAL STRUCTURES OF ACIDIC AND BASIC FIBROBLAST GROWTH FACTORS | | Descriptor: | ACIDIC FIBROBLAST GROWTH FACTOR | | Authors: | Zhu, X, Komiya, H, Chirino, A, Faham, S, Fox, G.M, Arakawa, T, Hsu, B.T, Rees, D.C. | | Deposit date: | 1992-09-29 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Three-dimensional structures of acidic and basic fibroblast growth factors.
Science, 251, 1991
|
|
1BEX
 
 | | STRUCTURE OF RUTHENIUM-MODIFIED PSEUDOMONAS AERUGINOSA AZURIN | | Descriptor: | AZURIN, COPPER (II) ION, RUTHEMIUM BIS(2,2'-BIPYRIDINE)-2-IMIDAZOLE | | Authors: | Faham, S, Day, M.W, Rees, D.C. | | Deposit date: | 1998-05-18 | | Release date: | 1998-10-14 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of ruthenium-modified Pseudomonas aeruginosa azurin and [Ru(2,2'-bipyridine)2(imidazole)2]SO4 x 10H2O.
Acta Crystallogr.,Sect.D, 55, 1999
|
|
1BAS
 
 | |
1BQ8
 
 | | Rubredoxin (Methionine Mutant) from Pyrococcus Furiosus | | Descriptor: | FE (III) ION, PROTEIN (RUBREDOXIN) | | Authors: | Bau, R, Rees, D.C, Kurtz, D.M, Scott, R.A, Huang, H, Adams, M.W.W, Eidsness, M.K. | | Deposit date: | 1998-08-22 | | Release date: | 1998-08-26 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Crystal Structure of Rubredoxin from Pyrococcus Furiosus at 0.95 Angstroms Resolution, and the structures of N-terminal methionine and formylmethionine variants of Pf Rd. Contributions of N-terminal interactions to thermostability
J.BIOL.INORG.CHEM., 3, 1998
|
|
1BVB
 
 | | HEME-PACKING MOTIFS REVEALED BY THE CRYSTAL STRUCTURE OF CYTOCHROME C554 FROM NITROSOMONAS EUROPAEA | | Descriptor: | CYTOCHROME C-554, PHOSPHATE ION, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Iverson, T.M, Arciero, D.M, Hsu, B.T, Logan, M.S.P, Hooper, A.B, Rees, D.C. | | Deposit date: | 1998-09-16 | | Release date: | 1999-05-18 | | Last modified: | 2019-08-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Heme packing motifs revealed by the crystal structure of the tetra-heme cytochrome c554 from Nitrosomonas europaea.
Nat.Struct.Biol., 5, 1998
|
|
1BRF
 
 | | Rubredoxin (Wild Type) from Pyrococcus Furiosus | | Descriptor: | FE (III) ION, PROTEIN (RUBREDOXIN) | | Authors: | Bau, R, Rees, D.C, Kurtz, D.M, Scott, R.A, Huang, H, Adams, M.W.W, Eidsness, M.K. | | Deposit date: | 1998-08-24 | | Release date: | 1998-09-02 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (0.95 Å) | | Cite: | Crystal Structure of Rubredoxin from Pyrococcus Furiosus at 0.95 Angstroms Resolution, and the structures of N-terminal methionine and formylmethionine variants of Pf Rd. Contributions of N-terminal interactions to thermostability
J.BIOL.INORG.CHEM., 3, 1998
|
|
1BQ9
 
 | | Rubredoxin (Formyl Methionine Mutant) from Pyrococcus Furiosus | | Descriptor: | FE (III) ION, PROTEIN (RUBREDOXIN) | | Authors: | Bau, R, Rees, D.C, Kurtz, D.M, Scott, R.A, Huang, H, Adams, M.W.W, Eidsness, M.K. | | Deposit date: | 1998-08-22 | | Release date: | 1998-08-26 | | Last modified: | 2022-12-21 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Crystal Structure of Rubredoxin from Pyrococcus Furiosus at 0.95 Angstroms Resolution, and the structures of N-terminal methionine and formylmethionine variants of Pf Rd. Contributions of N-terminal interactions to thermostability
J.BIOL.INORG.CHEM., 3, 1998
|
|
1CAD
 
 | | X-RAY CRYSTAL STRUCTURES OF THE OXIDIZED AND REDUCED FORMS OF THE RUBREDOXIN FROM THE MARINE HYPERTHERMOPHILIC ARCHAEBACTERIUM PYROCOCCUS FURIOSUS | | Descriptor: | FE (III) ION, RUBREDOXIN | | Authors: | Day, M.W, Hsu, B.T, Joshua-Tor, L, Park, J.B, Zhou, Z.H, Adams, M.W.W, Rees, D.C. | | Deposit date: | 1992-05-18 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray crystal structures of the oxidized and reduced forms of the rubredoxin from the marine hyperthermophilic archaebacterium Pyrococcus furiosus.
Protein Sci., 1, 1992
|
|
1CAA
 
 | | X-RAY CRYSTAL STRUCTURES OF THE OXIDIZED AND REDUCED FORMS OF THE RUBREDOXIN FROM THE MARINE HYPERTHERMOPHILIC ARCHAEBACTERIUM PYROCOCCUS FURIOSUS | | Descriptor: | FE (III) ION, RUBREDOXIN | | Authors: | Day, M.W, Hsu, B.T, Joshua-Tor, L, Park, J.B, Zhou, Z.H, Adams, M.W.W, Rees, D.C. | | Deposit date: | 1992-05-18 | | Release date: | 1993-10-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-ray crystal structures of the oxidized and reduced forms of the rubredoxin from the marine hyperthermophilic archaebacterium Pyrococcus furiosus.
Protein Sci., 1, 1992
|
|
8TC3
 
 | |
