3KQV
 
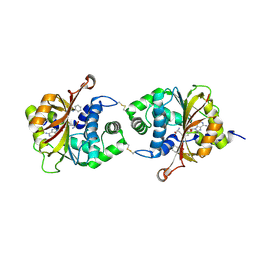 | | Crystal Structure of hPNMT in Complex AdoHcy and Formanilide | | Descriptor: | FORMANILIDE, Phenylethanolamine N-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Drinkwater, N, Martin, J.L. | | Deposit date: | 2009-11-17 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Fragment-based screening by X-ray crystallography, MS and isothermal titration calorimetry to identify PNMT (phenylethanolamine N-methyltransferase) inhibitors.
Biochem.J., 431, 2010
|
|
3KG9
 
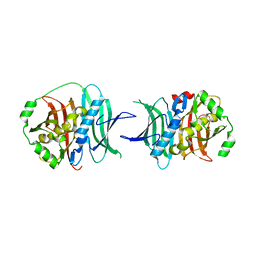 | |
6WYE
 
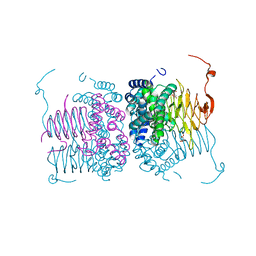 | | Crystal structure of Neisseria gonorrhoeae serine acetyltransferase (CysE) | | Descriptor: | (2S)-2-hydroxybutanedioic acid, SODIUM ION, Serine acetyltransferase | | Authors: | Hicks, J.L, Oldham, K.E, Summers, E.L, Prentice, E.J. | | Deposit date: | 2020-05-12 | | Release date: | 2021-06-09 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Serine acetyltransferase from Neisseria gonorrhoeae; structural and biochemical basis of inhibition.
Biochem.J., 479, 2022
|
|
3KLA
 
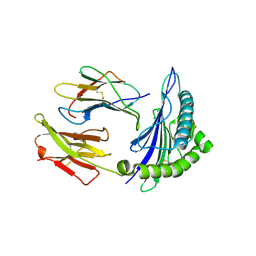 | | Ca2+ release from the endoplasmic reticulum of NY-ESO-1 specific T cells is modulated by the affinity of T cell receptor and by the use of the CD8 co-receptor | | Descriptor: | Beta-2-microglobulin, HLA class I histocompatibility antigen, A-2 alpha chain, ... | | Authors: | Chen, J.L, Morgan, A.J, Stewart-Jones, G, Shepherd, D, Bossi, G, Wooldridge, L. | | Deposit date: | 2009-11-07 | | Release date: | 2010-02-16 | | Last modified: | 2021-10-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Ca2+ Release from the Endoplasmic Reticulum of NY-ESO-1-Specific T Cells Is Modulated by the Affinity of TCR and by the Use of the CD8 Coreceptor.
J.Immunol., 184, 2010
|
|
3KPW
 
 | | Crystal Structure of hPNMT in Complex AdoHcy and 1-Aminoisoquinoline | | Descriptor: | ISOQUINOLIN-1-AMINE, Phenylethanolamine N-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Drinkwater, N, Martin, J.L. | | Deposit date: | 2009-11-17 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Fragment-based screening by X-ray crystallography, MS and isothermal titration calorimetry to identify PNMT (phenylethanolamine N-methyltransferase) inhibitors.
Biochem.J., 431, 2010
|
|
3KQP
 
 | | Crystal Structure of hPNMT in Complex AdoHcy and 6-Aminoquinoline | | Descriptor: | Phenylethanolamine N-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE, quinolin-6-amine | | Authors: | Drinkwater, N, Martin, J.L. | | Deposit date: | 2009-11-17 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Fragment-based screening by X-ray crystallography, MS and isothermal titration calorimetry to identify PNMT (phenylethanolamine N-methyltransferase) inhibitors.
Biochem.J., 431, 2010
|
|
7S6K
 
 | | J08 fragment antigen binding in complex with SARS-CoV-2-6P-Mut2 S protein (conformation 2) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, J08 fragment antigen binding heavy chain variable domain, ... | | Authors: | Ozorowski, G, Torres, J.L, Ward, A.B. | | Deposit date: | 2021-09-14 | | Release date: | 2022-05-11 | | Last modified: | 2022-05-25 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structural insights of a highly potent pan-neutralizing SARS-CoV-2 human monoclonal antibody.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7S6L
 
 | | J08 fragment antigen binding in complex with SARS-CoV-2-6P-Mut7 S protein (conformation 3) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, J08 fragment antigen binding heavy chain variable domain, ... | | Authors: | Ozorowski, G, Torres, J.L, Ward, A.B. | | Deposit date: | 2021-09-14 | | Release date: | 2022-05-11 | | Last modified: | 2022-05-25 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Structural insights of a highly potent pan-neutralizing SARS-CoV-2 human monoclonal antibody.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7S6J
 
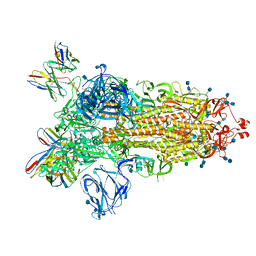 | | J08 fragment antigen binding in complex with SARS-CoV-2-6P-Mut2 S protein (conformation 1) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, J08 fragment antigen binding heavy chain variable domain, ... | | Authors: | Ozorowski, G, Torres, J.L, Ward, A.B. | | Deposit date: | 2021-09-14 | | Release date: | 2022-05-11 | | Last modified: | 2022-05-25 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structural insights of a highly potent pan-neutralizing SARS-CoV-2 human monoclonal antibody.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7S6I
 
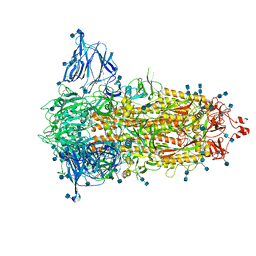 | | SARS-CoV-2-6P-Mut2 S protein | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein | | Authors: | Ozorowski, G, Torres, J.L, Ward, A.B. | | Deposit date: | 2021-09-14 | | Release date: | 2022-05-11 | | Last modified: | 2022-05-25 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structural insights of a highly potent pan-neutralizing SARS-CoV-2 human monoclonal antibody.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
6WW6
 
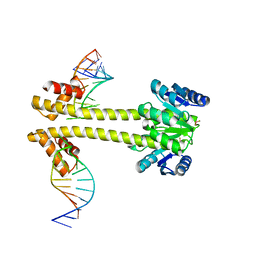 | | Crystal structure of EutV bound to RNA | | Descriptor: | BERYLLIUM TRIFLUORIDE ION, Response regulator, eutP P2 | | Authors: | Ataide, S.F, Walshe, J.L. | | Deposit date: | 2020-05-07 | | Release date: | 2021-11-24 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | Structural characterization of the ANTAR antiterminator domain bound to RNA.
Nucleic Acids Res., 50, 2022
|
|
3KQT
 
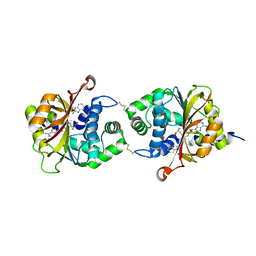 | |
3KT2
 
 | | Crystal Structure of N88D mutant HIV-1 Protease | | Descriptor: | Protease | | Authors: | Bihani, S.C, Das, A, Prashar, V, Ferrer, J.L, Hosur, M.V. | | Deposit date: | 2009-11-24 | | Release date: | 2010-02-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.651 Å) | | Cite: | Resistance mechanism revealed by crystal structures of unliganded nelfinavir-resistant HIV-1 protease non-active site mutants N88D and N88S.
Biochem.Biophys.Res.Commun., 389, 2009
|
|
7S3A
 
 | |
7S3B
 
 | |
3KG7
 
 | |
7S3C
 
 | |
3KPU
 
 | | Crystal Structure of hPNMT in Complex AdoHcy and 4-quinolinol | | Descriptor: | Phenylethanolamine N-methyltransferase, S-ADENOSYL-L-HOMOCYSTEINE, quinolin-4-ol | | Authors: | Drinkwater, N, Martin, J.L. | | Deposit date: | 2009-11-17 | | Release date: | 2010-09-29 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Fragment-based screening by X-ray crystallography, MS and isothermal titration calorimetry to identify PNMT (phenylethanolamine N-methyltransferase) inhibitors.
Biochem.J., 431, 2010
|
|
3KR0
 
 | |
3KT5
 
 | | Crystal Structure of N88S mutant HIV-1 Protease | | Descriptor: | Protease | | Authors: | Bihani, S.C, Das, A, Prashar, V, Ferrer, J.L, Hosur, M.V. | | Deposit date: | 2009-11-24 | | Release date: | 2010-02-16 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (1.801 Å) | | Cite: | Resistance mechanism revealed by crystal structures of unliganded nelfinavir-resistant HIV-1 protease non-active site mutants N88D and N88S.
Biochem.Biophys.Res.Commun., 389, 2009
|
|
6XLW
 
 | | Crystal structure of U2AF65 bound to AdML splice site sequence | | Descriptor: | DI(HYDROXYETHYL)ETHER, DNA/RNA (5'-R(P*UP*UP*(UD)P*UP*U)-D(P*(BRU))-R(P*CP*C)-3'), GLYCEROL, ... | | Authors: | Maji, D, Jenkins, J.L, Kielkopf, C.L. | | Deposit date: | 2020-06-29 | | Release date: | 2020-10-07 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Representative cancer-associated U2AF2 mutations alter RNA interactions and splicing.
J.Biol.Chem., 295, 2020
|
|
6XH1
 | |
6XYU
 
 | | Update of AChE from Drosophila Melanogaster complex with tacrine derivative 9-(3-iodobenzylamino)-1,2,3,4-tetrahydroacridine | | Descriptor: | 9-(3-IODOBENZYLAMINO)-1,2,3,4-TETRAHYDROACRIDINE, Acetylcholinesterase, IODIDE ION, ... | | Authors: | Nachon, F, Sussman, J.L. | | Deposit date: | 2020-01-31 | | Release date: | 2020-03-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | A Second Look at the Crystal Structures ofDrosophila melanogasterAcetylcholinesterase in Complex with Tacrine Derivatives Provides Insights Concerning Catalytic Intermediates and the Design of Specific Insecticides.
Molecules, 25, 2020
|
|
3KR2
 
 | |
6XH0
 
 | |
