6TQ5
 
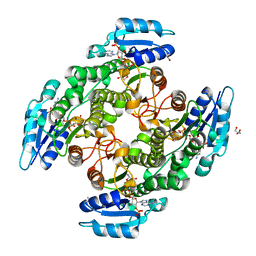 | | Alcohol dehydrogenase from Candida magnoliae DSMZ 70638 (ADHA): complex with NADP+ | | Descriptor: | Enzyme subunit, GLYCEROL, ISOPROPYL ALCOHOL, ... | | Authors: | Rovida, S, Aalbers, F.S, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2019-12-16 | | Release date: | 2020-04-08 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Approaching boiling point stability of an alcohol dehydrogenase through computationally-guided enzyme engineering.
Elife, 9, 2020
|
|
3PO7
 
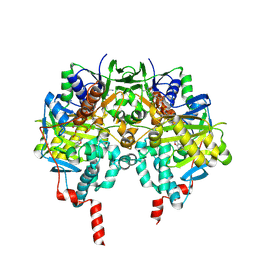 | | Human monoamine oxidase B in complex with zonisamide | | Descriptor: | 1-(1,2-benzoxazol-3-yl)methanesulfonamide, Amine oxidase [flavin-containing] B, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Binda, C, Aldeco, M, Mattevi, A, Edmondson, D.E. | | Deposit date: | 2010-11-22 | | Release date: | 2011-01-12 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Interactions of monoamine oxidases with the antiepileptic drug zonisamide: specificity of inhibition and structure of the human monoamine oxidase B complex
J.Med.Chem., 54, 2011
|
|
7ZRY
 
 | |
4F4Q
 
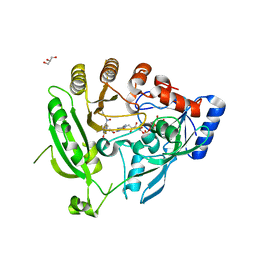 | | Crystal structure of M. smegmatis DprE1 in complex with FAD and covalently bound BTZ043 | | Descriptor: | 8-(hydroxyamino)-2-[(2S)-2-methyl-1,4-dioxa-8-azaspiro[4.5]dec-8-yl]-6-(trifluoromethyl)-4H-1,3-benzothiazin-4-one, DprE1, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Neres, J, Pojer, F, Molteni, E, Chiarelli, L, Riccardi, G, Mattevi, A, Cole, S.T, Binda, C. | | Deposit date: | 2012-05-11 | | Release date: | 2012-09-05 | | Last modified: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (2.619 Å) | | Cite: | Structural Basis for Benzothiazinone-Mediated Killing of Mycobacterium tuberculosis.
Sci Transl Med, 4, 2012
|
|
1LQT
 
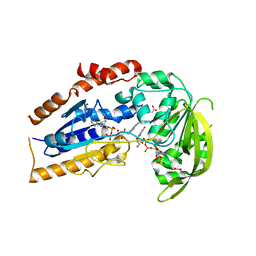 | | A covalent modification of NADP+ revealed by the atomic resolution structure of FprA, a Mycobacterium tuberculosis oxidoreductase | | Descriptor: | 4-OXO-NICOTINAMIDE-ADENINE DINUCLEOTIDE PHOSPHATE, ACETATE ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Bossi, R.T, Aliverti, A, Raimondi, D, Fischer, F, Zanetti, G, Ferrari, D, Tahallah, N, Maier, C.S, Heck, A.J.R, Rizzi, M, Mattevi, A, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2002-05-13 | | Release date: | 2002-07-31 | | Last modified: | 2024-12-25 | | Method: | X-RAY DIFFRACTION (1.05 Å) | | Cite: | A covalent modification of NADP+ revealed by the atomic resolution structure of FprA, a Mycobacterium tuberculosis oxidoreductase.
Biochemistry, 41, 2002
|
|
2UUU
 
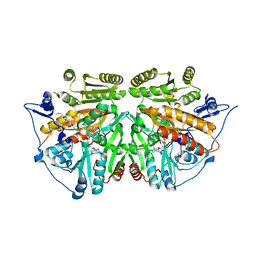 | | alkyldihydroxyacetonephosphate synthase in P212121 | | Descriptor: | ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE, FLAVIN-ADENINE DINUCLEOTIDE, HEXADECAN-1-OL | | Authors: | Razeto, A, Mattiroli, F, Carpanelli, E, Aliverti, A, Pandini, V, Coda, A, Mattevi, A. | | Deposit date: | 2007-03-07 | | Release date: | 2007-06-26 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The Crucial Step in Ether Phospholipid Biosynthesis: Structural Basis of a Noncanonical Reaction Associated with a Peroxisomal Disorder.
Structure, 15, 2007
|
|
4D03
 
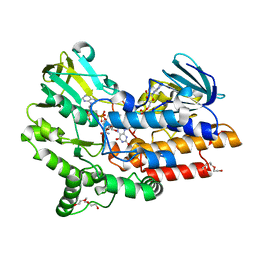 | | Structure of the Cys65Asp mutant of phenylacetone monooxygenase: oxidised state | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, HEXAETHYLENE GLYCOL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Brondani, P.B, Dudek, H.M, Martinoli, C, Mattevi, A, Fraaije, M.W. | | Deposit date: | 2014-04-24 | | Release date: | 2014-12-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Finding the Switch: Turning a Baeyer-Villiger Monooxygenase Into a Nadph Oxidase.
J.Am.Chem.Soc., 136, 2014
|
|
4D04
 
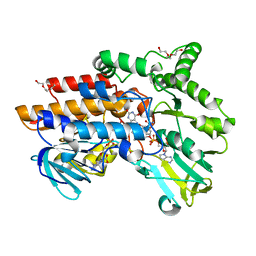 | | Structure of the Cys65Asp mutant of phenylacetone monooxygenase: reduced state | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, HEXAETHYLENE GLYCOL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Brondani, P.B, Dudek, H.M, Martinoli, C, Mattevi, A, Fraaije, M.W. | | Deposit date: | 2014-04-24 | | Release date: | 2014-12-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Finding the Switch: Turning a Baeyer-Villiger Monooxygenase Into a Nadph Oxidase.
J.Am.Chem.Soc., 136, 2014
|
|
4CZZ
 
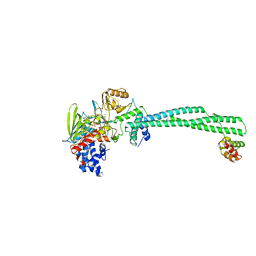 | | Histone demethylase LSD1(KDM1A)-CoREST3 Complex | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, LYSINE-SPECIFIC HISTONE DEMETHYLASE 1A, REST COREPRESSOR 3 | | Authors: | Barrios, A.P, Gomez, A.V, Saez, J.E, Ciossani, G, Toffolo, E, Battaglioli, E, Mattevi, A, Andres, M.E. | | Deposit date: | 2014-04-23 | | Release date: | 2014-06-11 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Differential Properties of Transcriptional Complexes Formed by the Corest Family.
Mol.Cell.Biol., 34, 2014
|
|
6YT2
 
 | | Crystal Structure of human monoamine oxidase B in complex with Diphenylene iodonium (DPI) | | Descriptor: | Amine oxidase [flavin-containing] B, FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, ... | | Authors: | Iacovino, L.G, Reis, J, Mai, A, Mattevi, A, Binda, C. | | Deposit date: | 2020-04-23 | | Release date: | 2020-06-10 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Diphenylene Iodonium Is a Noncovalent MAO Inhibitor: A Biochemical and Structural Analysis.
Chemmedchem, 15, 2020
|
|
2VGB
 
 | | HUMAN ERYTHROCYTE PYRUVATE KINASE | | Descriptor: | 1,6-di-O-phosphono-beta-D-fructofuranose, 2-PHOSPHOGLYCOLIC ACID, MANGANESE (II) ION, ... | | Authors: | Valentini, G, Chiarelli, L, Fortin, R, Dolzan, M, Galizzi, A, Abraham, D.J, Wang, C, Bianchi, P, Zanella, A, Mattevi, A. | | Deposit date: | 2007-11-12 | | Release date: | 2007-11-20 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.73 Å) | | Cite: | Structure and Function of Human Erythrocyte Pyruvate Kinase. Molecular Basis of Nonspherocytic Hemolytic Anemia.
J.Biol.Chem., 277, 2002
|
|
2VGI
 
 | | HUMAN ERYTHROCYTE PYRUVATE KINASE: R486W MUTANT | | Descriptor: | 1,6-di-O-phosphono-beta-D-fructofuranose, 2-PHOSPHOGLYCOLIC ACID, MANGANESE (II) ION, ... | | Authors: | Valentini, G, Chiarelli, L, Fortin, R, Dolzan, M, Galizzi, A, Abraham, D.J, Wang, C, Bianchi, P, Zanella, A, Mattevi, A. | | Deposit date: | 2007-11-13 | | Release date: | 2007-11-20 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | Structure and Function of Human Erythrocyte Pyruvate Kinase. Molecular Basis of Nonspherocytic Hemolytic Anemia.
J.Biol.Chem., 277, 2002
|
|
9EL8
 
 | | LSD1-CoREST in complex with T105 1S2R enantiomer | | Descriptor: | 3-[(1R,2S)-2-(cyclobutylamino)cyclopropyl]-N-(5-methyl-1,3,4-thiadiazol-2-yl)benzamide, Lysine-specific histone demethylase 1A, REST corepressor 1, ... | | Authors: | Caroli, J, Mattevi, A. | | Deposit date: | 2024-12-04 | | Release date: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (2.91 Å) | | Cite: | Distal drug resistance mutations promote covalent inhibitor-adduct Grob fragmentation in LSD1
to be published
|
|
9ELA
 
 | | LSD1-CoREST in complex with T108, long soaking | | Descriptor: | Lysine-specific histone demethylase 1A, REST corepressor 1, [(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methyl (2R,3S,4S)-2,3,4-trihydroxy-5-[(1R,3R,3aS,13R)-1-hydroxy-10,11-dimethyl-4,6-dioxo-3-[3-(phenylcarbamoyl)phenyl]-2,3,5,6-tetrahydro-1H-benzo[g]pyrrolo[2,1-e]pteridin-8(4H)-yl]pentyl dihydrogen diphosphate | | Authors: | Caroli, J, Mattevi, A. | | Deposit date: | 2024-12-04 | | Release date: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Distal drug resistance mutations promote covalent inhibitor-adduct Grob fragmentation in LSD1
to be published
|
|
9EL7
 
 | | LSD1-CoREST in complex with T105 enantiomer (1R,2S) | | Descriptor: | Lysine-specific histone demethylase 1A, REST corepressor 1, [(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methyl (2S,3S,4R)-5-[(4aS)-7,8-dimethyl-4a-[(1R)-1-{3-[(5-methyl-1,3,4-thiadiazol-2-yl)carbamoyl]phenyl}-3-oxopropyl]-2,4-dioxo-3,4,4a,5-tetrahydrobenzo[g]pteridin-10(2H)-yl]-2,3,4-trihydroxypentyl dihydrogen diphosphate (non-preferred name) | | Authors: | Caroli, J, Mattevi, A. | | Deposit date: | 2024-12-04 | | Release date: | 2025-02-05 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | Distal drug resistance mutations promote covalent inhibitor-adduct Grob fragmentation in LSD1
to be published
|
|
7YWU
 
 | | Eugenol oxidase from rhodococcus jostii: mutant S81H, A423M, H434Y, S394V, I445D, S518P | | Descriptor: | 2-methoxy-4-(prop-2-en-1-yl)phenol, FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, ... | | Authors: | Alvigini, L, Mattevi, A. | | Deposit date: | 2022-02-14 | | Release date: | 2022-11-16 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure- and computational-aided engineering of an oxidase to produce isoeugenol from a lignin-derived compound.
Nat Commun, 13, 2022
|
|
7YWV
 
 | | Eugenol oxidase from rhodococcus jostii: mutant S81H, D151E, A423M, H434Y, S394V, Q425S, I445D, S518P | | Descriptor: | 2-methoxy-4-(prop-2-en-1-yl)phenol, CALCIUM ION, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Alvigini, L, Mattevi, A. | | Deposit date: | 2022-02-14 | | Release date: | 2022-11-16 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure- and computational-aided engineering of an oxidase to produce isoeugenol from a lignin-derived compound.
Nat Commun, 13, 2022
|
|
2V5Z
 
 | | Structure of human MAO B in complex with the selective inhibitor safinamide | | Descriptor: | (S)-(+)-2-[4-(FLUOROBENZYLOXY-BENZYLAMINO)PROPIONAMIDE], Amine oxidase [flavin-containing] B, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Binda, C, Wang, J, Pisani, L, Caccia, C, Carotti, A, Salvati, P, Edmondson, D.E, Mattevi, A. | | Deposit date: | 2007-07-12 | | Release date: | 2007-10-16 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structures of human monoamine oxidase B complexes with selective noncovalent inhibitors: safinamide and coumarin analogs.
J.Med.Chem., 50, 2007
|
|
2V60
 
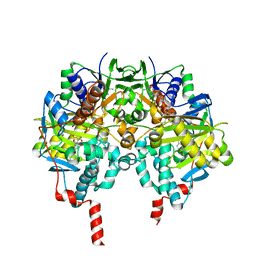 | | Structure of human MAO B in complex with the selective inhibitor 7-(3- chlorobenzyloxy)-4-carboxaldehyde-coumarin | | Descriptor: | 7-[(3-CHLOROBENZYL)OXY]-2-OXO-2H-CHROMENE-4-CARBALDEHYDE, AMINE OXIDASE (FLAVIN-CONTAINING) B, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Binda, C, Wang, J, Pisani, L, Caccia, C, Carotti, A, Salvati, P, Edmondson, D.E, Mattevi, A. | | Deposit date: | 2007-07-12 | | Release date: | 2007-10-16 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of Human Monoamine Oxidase B Complexes with Selective Noncovalent Inhibitors: Safinamide and Coumarin Analogs.
J.Med.Chem., 50, 2007
|
|
2V61
 
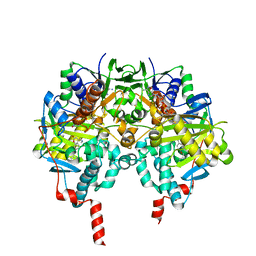 | | Structure of human MAO B in complex with the selective inhibitor 7-(3- chlorobenzyloxy)-4-(methylamino)methyl-coumarin | | Descriptor: | 7-[(3-CHLOROBENZYL)OXY]-4-[(METHYLAMINO)METHYL]-2H-CHROMEN-2-ONE, AMINE OXIDASE (FLAVIN-CONTAINING) B, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Binda, C, Wang, J, Pisani, L, Caccia, C, Carotti, A, Salvati, P, Edmondson, D.E, Mattevi, A. | | Deposit date: | 2007-07-13 | | Release date: | 2007-10-16 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structures of Human Monoamine Oxidase B Complexes with Selective Noncovalent Inhibitors: Safinamide and Coumarin Analogs.
J.Med.Chem., 50, 2007
|
|
1LLZ
 
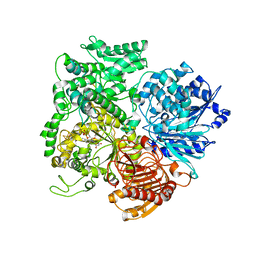 | | Structural studies on the synchronization of catalytic centers in glutamate synthase: reduced enzyme | | Descriptor: | FE3-S4 CLUSTER, FLAVIN MONONUCLEOTIDE, Ferredoxin-dependent glutamate synthase | | Authors: | van den Heuvel, R.H, Ferrari, D, Bossi, R.T, Ravasio, S, Curti, B, Vanoni, M.A, Florencio, F.J, Mattevi, A. | | Deposit date: | 2002-04-30 | | Release date: | 2002-07-31 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural studies on the synchronization of catalytic centers in glutamate synthase
J.BIOL.CHEM., 277, 2002
|
|
2V1D
 
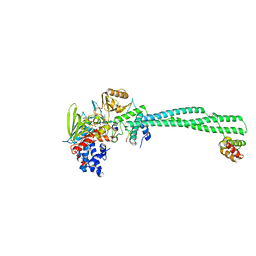 | | Structural basis of LSD1-CoREST selectivity in histone H3 recognition | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, HISTONE H3.1T, LYSINE-SPECIFIC HISTONE DEMETHYLASE 1, ... | | Authors: | Forneris, F, Binda, C, Adamo, A, Battaglioli, E, Mattevi, A. | | Deposit date: | 2007-05-23 | | Release date: | 2007-05-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural Basis of Lsd1-Corest Selectivity in Histone H3 Recognition.
J.Biol.Chem., 282, 2007
|
|
4OVI
 
 | | Phenylacetone monooxygenase: oxidised enzyme in complex with APADP | | Descriptor: | 3-ACETYLPYRIDINE ADENINE DINUCLEOTIDE, ACETATE ION, CHLORIDE ION, ... | | Authors: | Martinoli, C, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2013-09-19 | | Release date: | 2014-05-28 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Beyond the Protein Matrix: Probing Cofactor Variants in a Baeyer-Villiger Oxygenation Reaction.
Acs Catalysis, 3, 2013
|
|
5AE2
 
 | | Ether Lipid-Generating Enzyme AGPS in complex with inhibitor 1e | | Descriptor: | 3-(2-fluorophenyl)-2-methyl-N-((2-oxo-2,3-dihydro-1H-benzo[d]imidazol-5-yl)methyl)acrylamide, ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE, PEROXISOMAL, ... | | Authors: | Piano, V, Benjamin, D.I, Valente, S, Nenci, S, Marrocco, B, Mai, A, Aliverti, A, Nomura, D.K, Mattevi, A. | | Deposit date: | 2015-08-25 | | Release date: | 2015-09-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery of Inhibitors for the Ether Lipid-Generating Enzyme Agps as Anti-Cancer Agents.
Acs Chem.Biol., 10, 2015
|
|
5AE3
 
 | | Ether Lipid-Generating Enzyme AGPS in complex with antimycin A | | Descriptor: | ALKYLDIHYDROXYACETONEPHOSPHATE SYNTHASE, PEROXISOMAL, FLAVIN-ADENINE DINUCLEOTIDE, ... | | Authors: | Piano, V, Benjamin, D.I, Valente, S, Nenci, S, Marrocco, B, Mai, A, Aliverti, A, Nomura, D.K, Mattevi, A. | | Deposit date: | 2015-08-25 | | Release date: | 2015-09-09 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.18 Å) | | Cite: | Discovery of Inhibitors for the Ether Lipid-Generating Enzyme Agps as Anti-Cancer Agents.
Acs Chem.Biol., 10, 2015
|
|
