5ZIX
 
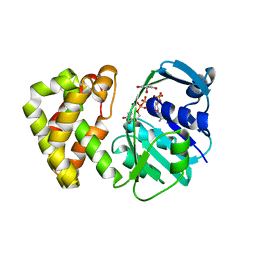 | |
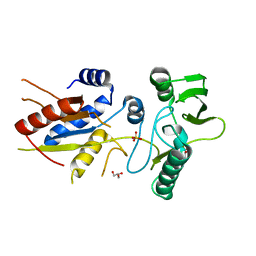
5XNW
 
 | |
5ZIK
 
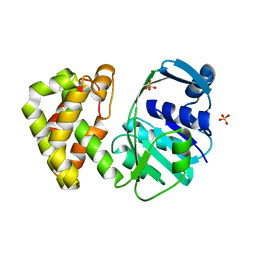 | | Crystal structure of Ketopantoate reductase from Pseudomonas aeruginosa | | Descriptor: | GLYCEROL, Probable 2-dehydropantoate 2-reductase, SULFATE ION | | Authors: | Khanppnavar, B, Datta, S. | | Deposit date: | 2018-03-16 | | Release date: | 2019-03-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Genome-wide survey and crystallographic analysis suggests a role for both horizontal gene transfer and duplication in pantothenate biosynthesis pathways.
Biochim Biophys Acta Gen Subj, 1863, 2019
|
|
8PNV
 
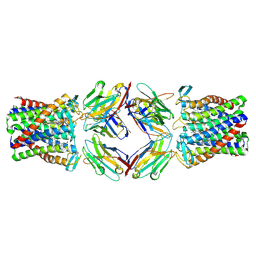 | | Cryo-EM structure of styrene oxide isomerase | | Descriptor: | Nanobody, PROTOPORPHYRIN IX CONTAINING FE, Styrene oxide isomerase | | Authors: | Khanppnavar, B, Korkhov, B, Li, X. | | Deposit date: | 2023-07-02 | | Release date: | 2024-04-03 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (2.048 Å) | | Cite: | Structural basis of the Meinwald rearrangement catalysed by styrene oxide isomerase.
Nat.Chem., 2024
|
|
8BUZ
 
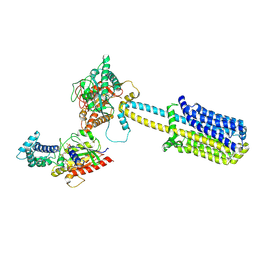 | | Structure of Adenylyl cyclase 8 bound to stimulatory G-protein, Ca2+/Calmodulin, Forskolin and MANT-GTP | | Descriptor: | 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, Adenylate cyclase type 8, FORSKOLIN, ... | | Authors: | Khanppnavar, B, Korkhov, V.M, Mehta, V. | | Deposit date: | 2022-12-01 | | Release date: | 2023-12-13 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Regulatory sites of CaM-sensitive adenylyl cyclase AC8 revealed by cryo-EM and structural proteomics.
Embo Rep., 25, 2024
|
|
8BV5
 
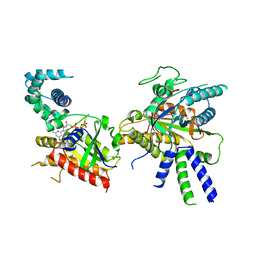 | | Focus refinement of soluble domain of Adenylyl cyclase 8 bound to stimulatory G protein, Forskolin, ATPalphaS, and Ca2+/Calmodulin in lipid nanodisc conditions | | Descriptor: | 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, Adenylate cyclase type 8, FORSKOLIN, ... | | Authors: | Khanppnavar, B, Korkhov, V.M. | | Deposit date: | 2023-01-04 | | Release date: | 2024-01-17 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.54 Å) | | Cite: | Regulatory sites of CaM-sensitive adenylyl cyclase AC8 revealed by cryo-EM and structural proteomics.
Embo Rep., 25, 2024
|
|
8PNU
 
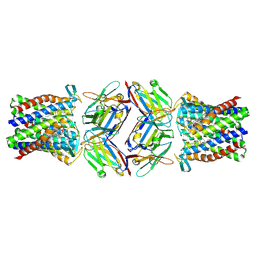 | |
7ZH0
 
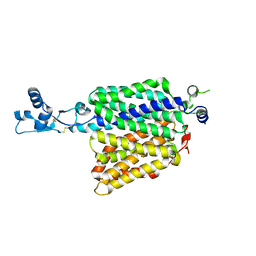 | |
7ZH6
 
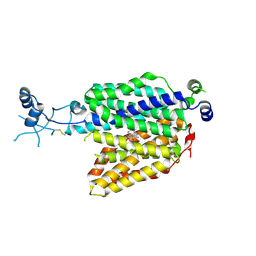 | |
7ZHA
 
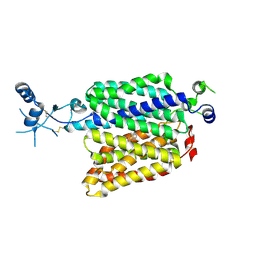 | |
6K1R
 
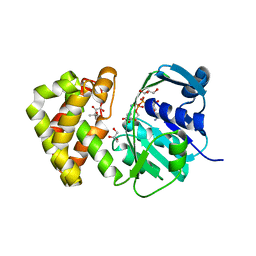 | |
7YZ9
 
 | | Structure of catalytic domain of Rv1625c bound to nanobody NB4 | | Descriptor: | 3'-O-(N-METHYLANTHRANILOYL)-GUANOSINE-5'-TRIPHOSPHATE, Adenylate cyclase, GLYCEROL, ... | | Authors: | Khanppnavar, B, Mehta, V.J, Iype, T, Korkhov, V.M. | | Deposit date: | 2022-02-19 | | Release date: | 2022-08-31 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structure of Mycobacterium tuberculosis Cya, an evolutionary ancestor of the mammalian membrane adenylyl cyclases.
Elife, 11, 2022
|
|
7YZK
 
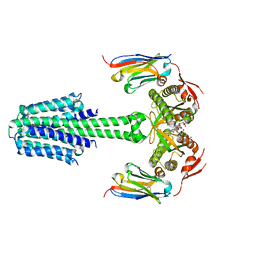 | | Structure of Mycobacterium tuberculosis adenylyl cyclase Rv1625c / Cya | | Descriptor: | 3'-O-(N-METHYLANTHRANILOYL)-GUANOSINE-5'-TRIPHOSPHATE, Adenylate cyclase, MANGANESE (II) ION, ... | | Authors: | Mehta, V, Khanppnavar, B, Korkhov, V.M. | | Deposit date: | 2022-02-20 | | Release date: | 2022-08-31 | | Method: | ELECTRON MICROSCOPY (3.57 Å) | | Cite: | Structure of Mycobacterium tuberculosis Cya, an evolutionary ancestor of the mammalian membrane adenylyl cyclases.
Elife, 11, 2022
|
|
7YZI
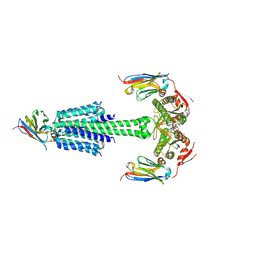 
 | | Structure of Mycobacterium tuberculosis adenylyl cyclase Rv1625c / Cya | | Descriptor: | 3'-O-(N-METHYLANTHRANILOYL)-GUANOSINE-5'-TRIPHOSPHATE, Adenylate cyclase, MANGANESE (II) ION, ... | | Authors: | Mehta, V, Khanppnavar, B, Korkhov, V.M. | | Deposit date: | 2022-02-20 | | Release date: | 2022-10-05 | | Method: | ELECTRON MICROSCOPY (3.83 Å) | | Cite: | Structure of Mycobacterium tuberculosis Cya, an evolutionary ancestor of the mammalian membrane adenylyl cyclases.
Elife, 11, 2022
|
|
7PD4
 
 | |
7PD8
 
 | |
7PDH
 
 | |
7PDD
 
 | |
7PDG
 
 | |
7PDE
 
 | |
7PDF
 
 | |
8QOJ
 
 | |
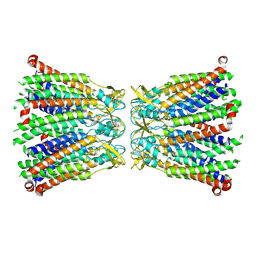
8R7P
 | |
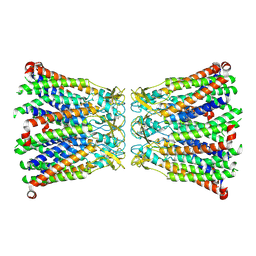
8R7Q
 | |
