7S00
 
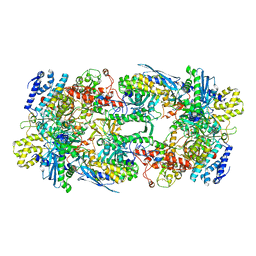 | | X-ray structure of the phage AR9 non-virion RNA polymerase core | | Descriptor: | DNA-directed RNA polymerase, DNA-directed RNA polymerase beta subunit, DNA-directed RNA polymerase beta' subunit, ... | | Authors: | Leiman, P.G, Sokolova, M.L, Fraser, A. | | Deposit date: | 2021-08-28 | | Release date: | 2022-07-06 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structural basis of template strand deoxyuridine promoter recognition by a viral RNA polymerase.
Nat Commun, 13, 2022
|
|
4W7K
 
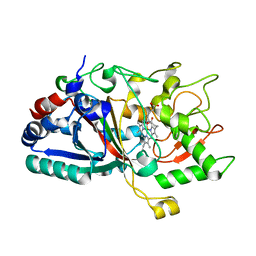 | |
2KCZ
 
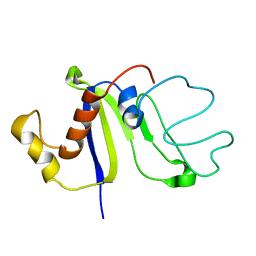 | | Solution NMR structure of the C-terminal domain of protein DR_A0006 from Deinococcus radiodurans. Northeast Structural Genomics Consortium Target DrR147D | | Descriptor: | Uncharacterized protein DR_A0006 | | Authors: | Mills, J.L, Ghosh, A, Garcia, E, Wang, H, Ciccosanti, C, Xiao, R, Nair, R, Everett, J.K, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-12-31 | | Release date: | 2009-01-20 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of the C-Terminal Domain of Protein DR_A0006 from Deinococcus radiodurans. Northeast Structural Genomics Consortium Target DrR147D.
To be Published
|
|
2KEL
 
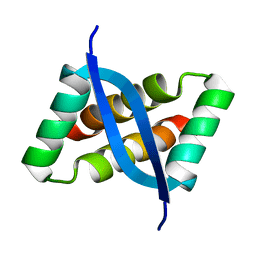 | | Structure of the transcription regulator SvtR from the hyperthermophilic archaeal virus SIRV1 | | Descriptor: | Uncharacterized protein 56B | | Authors: | Guilliere, F, Kessler, A, Peixeiro, N, Sezonov, G, Prangishvili, D, Delepierre, M, Guijarro, J.I. | | Deposit date: | 2009-01-30 | | Release date: | 2009-06-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure, function, and targets of the transcriptional regulator SvtR from the hyperthermophilic archaeal virus SIRV1.
J.Biol.Chem., 284, 2009
|
|
2KHQ
 
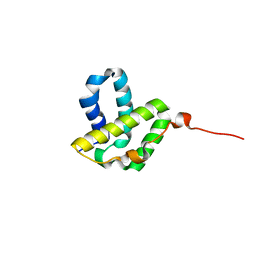 | | Solution NMR structure of a phage integrase SSP1947 fragment 59-159 from Staphylococcus saprophyticus, Northeast Structural Genomics Consortium Target SyR103B | | Descriptor: | Integrase | | Authors: | Eletsky, A, Mills, J.L, Hua, J, Belote, R.L, Ciccosanti, C, Jiang, M, Nair, R, Rost, B, Acton, T.B, Xiao, R, Swapna, G.V.T, Everett, J.K, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-04-10 | | Release date: | 2009-04-28 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of a phage integrase SSP1947 fragment 59-159 from
Staphylococcus saprophyticus
To be Published
|
|
7SMZ
 
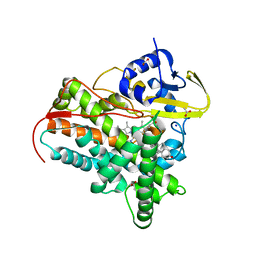 | | X-ray crystal structure of CYP142A3 from Mycobacterium Marinum in complex with 4-cholesten-3-one | | Descriptor: | (8ALPHA,9BETA)-CHOLEST-4-EN-3-ONE, ACETATE ION, Cytochrome P450 142A3, ... | | Authors: | Ghith, A, Bruning, J.B, Bell, S.G. | | Deposit date: | 2021-10-27 | | Release date: | 2022-09-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | The Structures of the Steroid Binding CYP142 Cytochrome P450 Enzymes from Mycobacterium ulcerans and Mycobacterium marinum.
Acs Infect Dis., 8, 2022
|
|
2KC3
 
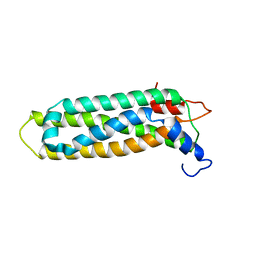 | |
2KK5
 
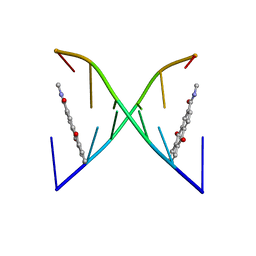 | | High Fidelity Base Pairing at the 3'-Terminus | | Descriptor: | 5'-D(*AP*CP*GP*CP*GP*(2AU))-3', 9,10-dioxo-9,10-dihydroanthracene-2-carboxamide | | Authors: | Patra, A, Richert, C. | | Deposit date: | 2009-06-15 | | Release date: | 2009-07-21 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | High fidelity base pairing at the 3'-terminus.
J.Am.Chem.Soc., 131, 2009
|
|
2KER
 | | alpha-amylase inhibitor Parvulustat (Z-2685) from Streptomyces parvulus | | Descriptor: | Alpha-amylase inhibitor Z-2685 | | Authors: | Rehm, S, Han, S, Hassani, I, Sokocevic, A, Jonker, H.R.A, Engels, J.W, Schwalbe, H. | | Deposit date: | 2009-02-02 | | Release date: | 2009-02-17 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | The high resolution NMR structure of parvulustat (Z-2685) from Streptomyces parvulus FH-1641: comparison with tendamistat from Streptomyces tendae 4158
Chembiochem, 10, 2009
|
|
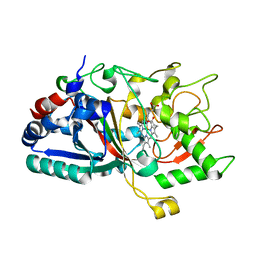
4W7M
 
 | |
2KH1
 
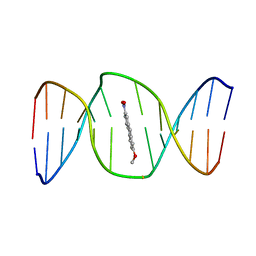 | | 2-Hydroxy-7-nitrofluorene covalently linked into a 13mer DNA duplex - solution structure of the face-up orientation | | Descriptor: | 5'-D(*CP*GP*AP*CP*GP*TP*(3DR)P*TP*GP*CP*AP*GP*C)-3', 5'-D(*GP*CP*TP*GP*CP*AP*(3DR)P*AP*CP*GP*TP*CP*G)-3', 7-nitro-9H-fluoren-2-ol | | Authors: | Dallmann, A, Pfaffe, M, Muegge, C, Mahrwald, R, Kovalenko, S.A, Ernsting, N.P. | | Deposit date: | 2009-03-23 | | Release date: | 2009-12-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Local THz Time Domain Spectroscopy of Duplex DNA via Fluorescence of an Embedded Probe.
J.Phys.Chem.B, 113, 2009
|
|
4WCG
 
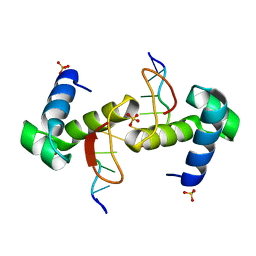 | | The binding mode of Cyprinid Herpesvirus3 ORF112-Zalpha to Z-DNA | | Descriptor: | DNA (5'-D(P*CP*GP*CP*GP*CP*G)-3'), ORF112, SULFATE ION | | Authors: | Kus, K, Athanasiadis, A. | | Deposit date: | 2014-09-04 | | Release date: | 2015-11-18 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The Structure of the Cyprinid herpesvirus 3 ORF112-Z alpha Z-DNA Complex Reveals a Mechanism of Nucleic Acids Recognition Conserved with E3L, a Poxvirus Inhibitor of Interferon Response.
J.Biol.Chem., 290, 2015
|
|
2KJA
 
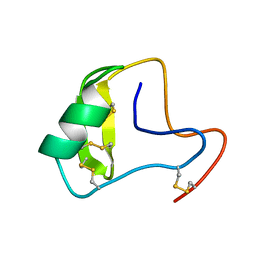 | | Chemical shift assignments, constraints, and coordinates for CN5 scorpion toxin | | Descriptor: | Beta-toxin Cn5 | | Authors: | Prochnicka-Chalufour, A, Corzo, G, Garcia, B.I, Possani, L.D, Delepierre, M. | | Deposit date: | 2009-05-26 | | Release date: | 2009-10-06 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Cn5, a crustacean toxin found in the venom of the scorpions Centruroides noxius and Centruroides suffusus suffusus.
Biochim.Biophys.Acta, 1794, 2009
|
|
2K5X
 
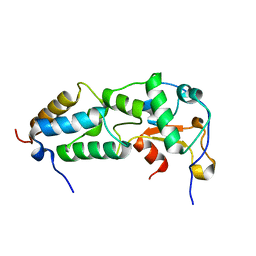 | |
2K7E
 | | NMR structure of the human tRNALys3 bound to the HIV genome Loop I | | Descriptor: | RNA (5'-R(*CP*UP*(SUR)P*UP*UP*AP*AP*(PSU)P*CP*UP*GP*C)-3'), RNA (5'-R(*GP*CP*GP*GP*UP*GP*UP*AP*AP*AP*AP*G)-3') | | Authors: | Bilbille, Y, Vendeix, F.P.A, Guenther, R, Malkiewicz, A, Ariza, X, Vilarrasa, J, Agris, P. | | Deposit date: | 2008-08-09 | | Release date: | 2008-10-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The structure of the human tRNALys3 anticodon bound to the HIV genome is stabilized by modified nucleosides and adjacent mismatch base pairs.
Nucleic Acids Res., 37, 2009
|
|
2KCT
 
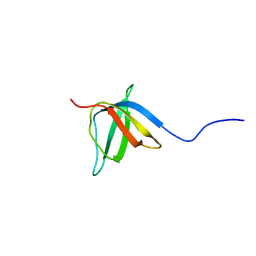 | | Solution nmr structure of the ob-fold domain of heme chaperone ccme from desulfovibrio vulgaris. northeast structural genomics target dvr115g. | | Descriptor: | Cytochrome c-type biogenesis protein CcmE | | Authors: | Aramini, J.M, Rossi, P, Lee, H, Lemak, A, Wang, H, Foote, E.L, Jiang, M, Xiao, R, Nair, R, Swapna, G.V.T, Acton, T.B, Rost, B, Everett, J.K, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-12-29 | | Release date: | 2009-02-10 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution nmr structure of the ob-fold domain of heme chaperone
ccme from desulfovibrio vulgaris. northeast structural genomics target dvr115g.
To be Published
|
|
2K5N
 
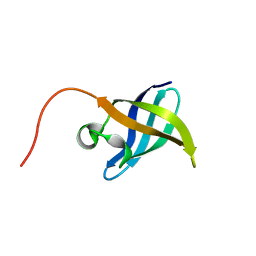 | | Solution NMR Structure of the N-Terminal Domain of Protein ECA1580 from Erwinia carotovora, Northeast Structural Genomics Consortium Target EwR156A | | Descriptor: | Putative cold-shock protein | | Authors: | Mills, J.L, Eletsky, A, Zhang, Q, Lee, D, Jiang, M, Ciccosanti, C, Xiao, R, Lui, J, Everett, J.K, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-30 | | Release date: | 2008-08-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of the Putative Cold Shock Protein from Erwinia carotovora: Northeast Structural Genomics Consortium Target EwR156a
To be Published
|
|
2K9C
 
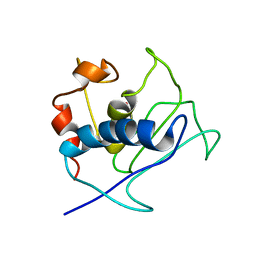 | | Paramagnetic shifts in solid-state NMR of Proteins to elicit structural information | | Descriptor: | COBALT (II) ION, Macrophage metalloelastase | | Authors: | Balayssac, S, Bertini, I, Bhaumik, A, Lelli, M, Luchinat, C. | | Deposit date: | 2008-10-08 | | Release date: | 2008-11-18 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Paramagnetic shifts in solid-state NMR of proteins to elicit structural information
Proc.Natl.Acad.Sci.Usa, 105, 2008
|
|
2KLH
 
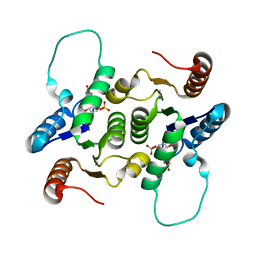 | | NMR Structure of RCL in complex with GMP | | Descriptor: | GUANOSINE-5'-MONOPHOSPHATE, c-Myc-responsive protein Rcl | | Authors: | Padilla, A, Yang, Y, Labesse, G, Zhang, C, Kaminski, P.A. | | Deposit date: | 2009-07-02 | | Release date: | 2009-10-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural characterization of the mammalian deoxynucleotide N-hydrolase Rcl and its stabilizing interactions with two inhibitors
J.Mol.Biol., 394, 2009
|
|
2KNF
 
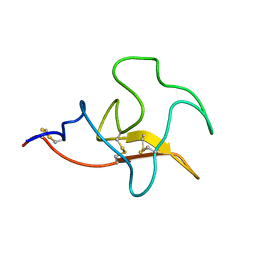 | | Solution structure and functional characterization of human plasminogen kringle 5 | | Descriptor: | Plasminogen | | Authors: | Battistel, M.D, Grishaev, A, An, S.A, Castellino, F.J, Llinas, M. | | Deposit date: | 2009-08-21 | | Release date: | 2009-10-27 | | Last modified: | 2021-10-13 | | Method: | SOLUTION NMR | | Cite: | Solution structure and functional characterization of human plasminogen kringle 5.
Biochemistry, 48, 2009
|
|
4UUQ
 
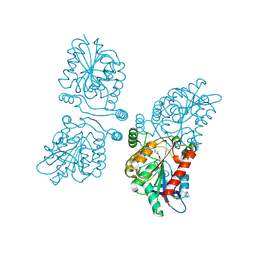 | | Crystal structure of human mono-glyceride lipase in complex with SAR127303 | | Descriptor: | 4-({[(4-chlorophenyl)sulfonyl]amino}methyl)piperidine-1-carboxylic acid, MONOGLYCERIDE LIPASE | | Authors: | Griebel, G, Pichat, P, Beeske, S, Leroy, T, Redon, N, Francon, D, Bert, L, Even, L, Lopez-Grancha, M, Tolstykh, T, Sun, F, Yu, Q, Brittain, S, Arlt, H, He, T, Zhang, B, Wiederschain, D, Bertrand, T, Houtman, J, Rak, A, Vallee, F, Michot, N, Auge, F, Menet, V, Bergis, O.E, George, P, Avenet, P, Mikol, V, Didier, M, Escoubet, J. | | Deposit date: | 2014-07-30 | | Release date: | 2015-01-21 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.36 Å) | | Cite: | Selective Blockade of the Hydrolysis of the Endocannabinoid 2-Arachidonoylglycerol Impairs Learning and Memory Performance While Producing Antinociceptive Activity in Rodents.
Sci.Rep., 5, 2015
|
|
2KAA
 
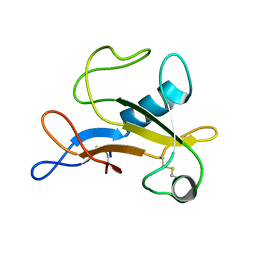 | |
2KD7
 
 | | Solution NMR structure of F5/8 type C-terminal domain of a putative chitobiase from Bacteroides thetaiotaomicron. Northeast Structural Genomics Consortium target BtR324B | | Descriptor: | Putative chitobiase | | Authors: | Eletsky, A, Mills, J.L, Lee, H, Lee, D, Jiang, M, Ciccosanti, C, Xiao, R, Nair, R, Everett, J.K, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Prestegard, J.H, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-01-04 | | Release date: | 2009-02-03 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of F5/8 type C-terminal domain of a putative chitobiase from Bacteroides thetaiotaomicron.
To be Published
|
|
1A9C
 
 | | GTP CYCLOHYDROLASE I (C110S MUTANT) IN COMPLEX WITH GTP | | Descriptor: | GTP CYCLOHYDROLASE I, GUANOSINE-5'-TRIPHOSPHATE | | Authors: | Auerbach, G, Nar, H, Bracher, A, Bacher, A, Huber, R. | | Deposit date: | 1998-04-04 | | Release date: | 1999-05-11 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | GTP Cyclohydrolase I in Complex with GTP at 2.1 A Resolution
To be Published
|
|
2KFQ
 
 | | NMR Structure of FP1 | | Descriptor: | FP1 | | Authors: | Araki, M, Tamura, A. | | Deposit date: | 2009-02-26 | | Release date: | 2009-03-31 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solubility-dependent structural formation of a 25-residue, natively unfolded protein, induced by addition of a seven-residue peptide fragment
Febs J., 276, 2009
|
|
