8CZ5
 
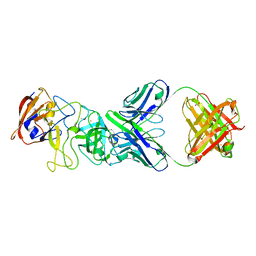 | | N11 P domain 2D9 Fab P complex | | Descriptor: | ACETATE ION, CADMIUM ION, Fab 2D9 Heavy Chain, ... | | Authors: | Leuthold, M, Kilic, T, Hansman, G. | | Deposit date: | 2022-05-24 | | Release date: | 2022-10-19 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.645 Å) | | Cite: | Structural Basis for Rabbit Hemorrhagic Disease Virus Antibody Specificity.
J.Virol., 96, 2022
|
|
8TBQ
 
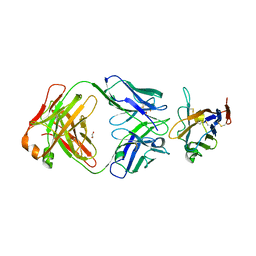 | | The crystal structure of human VISTA extra cellular domain in complex with Fab fragment of pH-selective anti-VISTA antibody | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, SNS-101 Fab heavy chain, ... | | Authors: | van der Horst, E.H, Mukherjee, A, Thisted, T. | | Deposit date: | 2023-06-29 | | Release date: | 2024-03-20 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.593 Å) | | Cite: | VISTA checkpoint inhibition by pH-selective antibody SNS-101 with optimized safety and pharmacokinetic profiles enhances PD-1 response.
Nat Commun, 15, 2024
|
|
7SGG
 
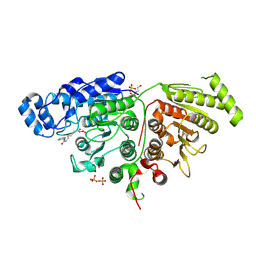 | |
7SGK
 
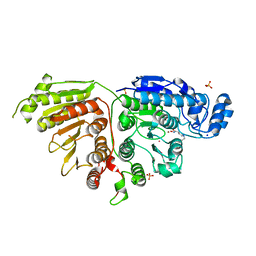 | |
9JJ7
 
 | |
9JHX
 
 | | Versatile Aromatic Prenyltransferase auraA mutant-Y207A in complex with DMSPP | | Descriptor: | DIMETHYLALLYL S-THIOLODIPHOSPHATE, Versatile Aromatic Prenyltransferase auraA | | Authors: | Li, D, Zhang, Y, Wang, W, Wang, P. | | Deposit date: | 2024-09-10 | | Release date: | 2025-02-05 | | Method: | ELECTRON MICROSCOPY (2.86 Å) | | Cite: | Characterization and structural analysis of a versatile aromatic prenyltransferase for imidazole-containing diketopiperazines.
Nat Commun, 16, 2025
|
|
7WX2
 
 | | CBP-BrD complexed with NEO2734 | | Descriptor: | 1,3-dimethyl-5-[2-(oxan-4-yl)-3-[2-(trifluoromethyloxy)ethyl]benzimidazol-5-yl]pyridin-2-one, CREB-binding protein | | Authors: | Zeng, L, Lei, J.D. | | Deposit date: | 2022-02-14 | | Release date: | 2022-06-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.24 Å) | | Cite: | Targeting CDCP1 gene transcription coactivated by BRD4 and CBP/p300 in castration-resistant prostate cancer.
Oncogene, 41, 2022
|
|
7WWZ
 
 | | BRD4-BD1 complexed with NEO2734 | | Descriptor: | 1,3-dimethyl-5-[2-(oxan-4-yl)-3-[2-(trifluoromethyloxy)ethyl]benzimidazol-5-yl]pyridin-2-one, Isoform C of Bromodomain-containing protein 4 | | Authors: | Zeng, L, Lei, J.D. | | Deposit date: | 2022-02-14 | | Release date: | 2022-06-08 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.16 Å) | | Cite: | Targeting CDCP1 gene transcription coactivated by BRD4 and CBP/p300 in castration-resistant prostate cancer.
Oncogene, 41, 2022
|
|
6ILM
 
 | | Cryo-EM structure of Echovirus 6 complexed with its uncoating receptor FcRn at PH 7.4 | | Descriptor: | Beta-2-microglobulin, Capsid protein VP1, Capsid protein VP2, ... | | Authors: | Gao, G.F, Liu, S, Zhao, X, Peng, R. | | Deposit date: | 2018-10-19 | | Release date: | 2019-05-15 | | Last modified: | 2025-06-25 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Human Neonatal Fc Receptor Is the Cellular Uncoating Receptor for Enterovirus B.
Cell, 177, 2019
|
|
6P5A
 
 | | Drosophila P element transposase strand transfer complex | | Descriptor: | DNA (44-MER), DNA (5'-D(P*AP*GP*GP*TP*GP*GP*TP*CP*CP*CP*GP*TP*CP*GP*G)-3'), DNA (5'-D(P*CP*GP*AP*AP*CP*TP*AP*TP*A)-3'), ... | | Authors: | Kellogg, E.H, Nogales, E, Ghanim, G, Rio, D.C. | | Deposit date: | 2019-05-30 | | Release date: | 2019-10-30 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structure of a P element transposase-DNA complex reveals unusual DNA structures and GTP-DNA contacts.
Nat.Struct.Mol.Biol., 26, 2019
|
|
6ILP
 
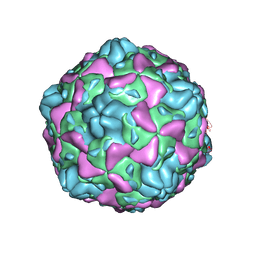 | | Cryo-EM structure of full Echovirus 6 particle at PH 7.4 | | Descriptor: | Capsid protein VP1, Capsid protein VP2, Capsid protein VP3, ... | | Authors: | Gao, G.F, Liu, S, Zhao, X, Peng, R. | | Deposit date: | 2018-10-19 | | Release date: | 2019-05-15 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Human Neonatal Fc Receptor Is the Cellular Uncoating Receptor for Enterovirus B.
Cell, 177, 2019
|
|
6ILN
 
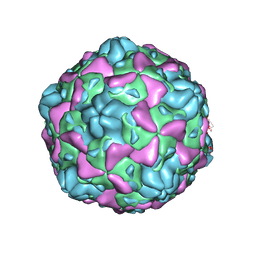 | | Cryo-EM structure of full Echovirus 6 particle at PH 5.5 | | Descriptor: | Capsid protein VP1, Capsid protein VP2, Capsid protein VP3, ... | | Authors: | Gao, G.F, Liu, S, Zhao, X, Peng, R. | | Deposit date: | 2018-10-19 | | Release date: | 2019-05-15 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Human Neonatal Fc Receptor Is the Cellular Uncoating Receptor for Enterovirus B.
Cell, 177, 2019
|
|
6PE2
 
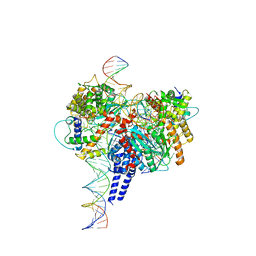 | | Drosophila P element transposase strand transfer complex | | Descriptor: | DNA (27-MER), DNA (5'-D(P*CP*GP*AP*AP*CP*TP*AP*TP*A)-3'), DNA (56-MER), ... | | Authors: | Kellogg, E.H, Nogales, E, Ghanim, G, Rio, D.C. | | Deposit date: | 2019-06-19 | | Release date: | 2019-10-30 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Structure of a P element transposase-DNA complex reveals unusual DNA structures and GTP-DNA contacts.
Nat.Struct.Mol.Biol., 26, 2019
|
|
8Y9E
 
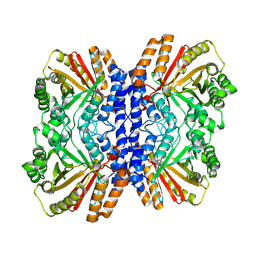 | |
8Y7T
 
 | | Crystal structure of SARS-CoV-2 main protease in complex with C2 | | Descriptor: | 3C-like proteinase nsp5, 6-(iminomethyl)-4-(2-pyridin-2-ylethyl)-2-[4-(trifluoromethyl)phenyl]-1,2,4-triazine-3,5-dione | | Authors: | Zeng, R, Deng, X.Y, Yang, Z.Y, Wang, K, Jiang, Y.Y, Lei, J. | | Deposit date: | 2024-02-05 | | Release date: | 2025-02-05 | | Last modified: | 2025-04-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A deep learning model for structure-based bioactivity optimization and its application in the bioactivity optimization of a SARS-CoV-2 main protease inhibitor.
Eur.J.Med.Chem., 291, 2025
|
|
8Y9D
 
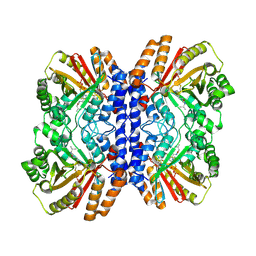 | | Versatile Aromatic Prenyltransferase auraA in complex with DMAPP and cyclo-(L-Val-L-His) | | Descriptor: | (3~{S},6~{S})-3-(1~{H}-imidazol-4-ylmethyl)-6-propan-2-yl-piperazine-2,5-dione, DIMETHYLALLYL S-THIOLODIPHOSPHATE, Versatile Aromatic Prenyltransferase auraA | | Authors: | Li, D, Zhang, Y, Wang, W, Wang, P. | | Deposit date: | 2024-02-06 | | Release date: | 2025-02-05 | | Method: | ELECTRON MICROSCOPY (2.45 Å) | | Cite: | Characterization and structural analysis of a versatile aromatic prenyltransferase for imidazole-containing diketopiperazines.
Nat Commun, 16, 2025
|
|
8Y7U
 
 | | Crystal structure of SARS-CoV-2 main protease in complex with C5 | | Descriptor: | 2-(3-fluoro-4-(trifluoromethyl)phenyl)-6-(iminomethyl)-4-(2-oxo-2-(pyridin-2-yl)ethyl)-1,2,4-triazine-3,5(2H,4H)-dione, 3C-like proteinase nsp5 | | Authors: | Zeng, R, Deng, X.Y, Yang, Z.Y, Wang, K, Jiang, Y.Y, Lei, J. | | Deposit date: | 2024-02-05 | | Release date: | 2025-02-05 | | Last modified: | 2025-04-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | A deep learning model for structure-based bioactivity optimization and its application in the bioactivity optimization of a SARS-CoV-2 main protease inhibitor.
Eur.J.Med.Chem., 291, 2025
|
|
8Y9G
 
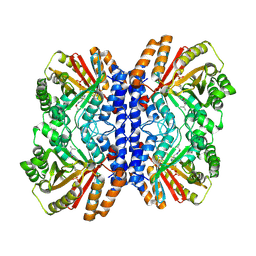 | | Versatile Aromatic Prenyltransferase auraA in complex with DMSPP and cyclo-(L-Val-DH-His) | | Descriptor: | (3~{Z},6~{S})-3-(1~{H}-imidazol-4-ylmethylidene)-6-propan-2-yl-piperazine-2,5-dione, DIMETHYLALLYL S-THIOLODIPHOSPHATE, Versatile Aromatic Prenyltransferase auraA | | Authors: | Li, D, Zhang, Y, Wang, W, Wang, P. | | Deposit date: | 2024-02-06 | | Release date: | 2025-02-05 | | Method: | ELECTRON MICROSCOPY (2.67 Å) | | Cite: | Characterization and structural analysis of a versatile aromatic prenyltransferase for imidazole-containing diketopiperazines.
Nat Commun, 16, 2025
|
|
5ER2
 
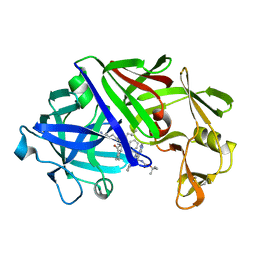 | | High-resolution X-ray diffraction study of the complex between endothiapepsin and an oligopeptide inhibitor. the analysis of the inhibitor binding and description of the rigid body shift in the enzyme | | Descriptor: | 6-ammonio-N-{[(2R,3R)-3-{[N-(tert-butoxycarbonyl)-L-phenylalanyl-3-(1H-imidazol-3-ium-4-yl)-L-alanyl]amino}-4-cyclohexyl-2-hydroxybutyl](2-methylpropyl)carbamoyl}-L-norleucyl-L-phenylalanine, ENDOTHIAPEPSIN | | Authors: | Sali, A, Veerapandian, B, Cooper, J.B, Foundling, S.I, Hoover, D.J, Blundell, T.L. | | Deposit date: | 1991-01-02 | | Release date: | 1991-04-15 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | High-resolution X-ray diffraction study of the complex between endothiapepsin and an oligopeptide inhibitor: the analysis of the inhibitor binding and description of the rigid body shift in the enzyme.
EMBO J., 8, 1989
|
|
8K0D
 
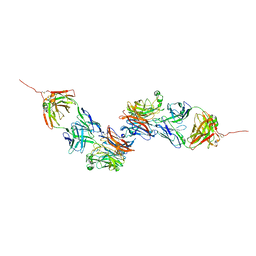 | |
8K0C
 
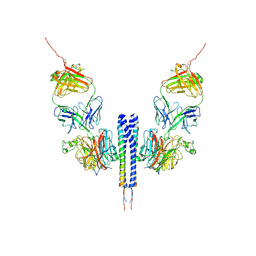 | |
6ILK
 
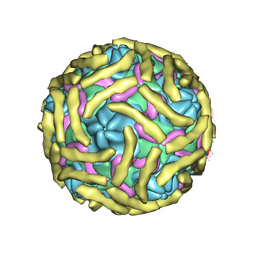 | | Cryo-EM structure of Echovirus 6 complexed with its attachment receptor CD55 at PH 7.4 | | Descriptor: | Capsid protein VP1, Capsid protein VP2, Capsid protein VP3, ... | | Authors: | Gao, G.F, Liu, S, Zhao, X, Peng, R. | | Deposit date: | 2018-10-18 | | Release date: | 2019-05-15 | | Last modified: | 2025-06-18 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Human Neonatal Fc Receptor Is the Cellular Uncoating Receptor for Enterovirus B.
Cell, 177, 2019
|
|
8EN4
 
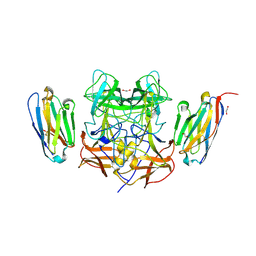 | | Structure of GII.4 norovirus in complex with Nanobody 53 | | Descriptor: | 1,2-ETHANEDIOL, Nanobody 53, VP1 | | Authors: | Kher, G, Sabin, C, Koromyslova, A, Pancera, M, Hansman, G. | | Deposit date: | 2022-09-28 | | Release date: | 2023-03-15 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Direct Blockade of the Norovirus Histo-Blood Group Antigen Binding Pocket by Nanobodies.
J.Virol., 97, 2023
|
|
8EN5
 
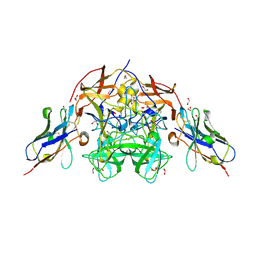 | | Structure of GII.4 norovirus in complex with Nanobody 56 | | Descriptor: | 1,2-ETHANEDIOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, GII.4 P domain, ... | | Authors: | Kher, G, Sabin, C, Pancera, M, Koromyslova, A, Hansman, G. | | Deposit date: | 2022-09-28 | | Release date: | 2023-03-15 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Direct Blockade of the Norovirus Histo-Blood Group Antigen Binding Pocket by Nanobodies.
J.Virol., 97, 2023
|
|
8EMZ
 
 | | Structure of GII.17 norovirus in complex with Nanobody 2 | | Descriptor: | 1,2-ETHANEDIOL, GII.17 P domain, Nanobody 2 | | Authors: | Kher, G, Sabin, C, Pancera, M, Koromyslova, A, Hansman, G. | | Deposit date: | 2022-09-28 | | Release date: | 2023-03-15 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Direct Blockade of the Norovirus Histo-Blood Group Antigen Binding Pocket by Nanobodies.
J.Virol., 97, 2023
|
|
