4P8Y
 
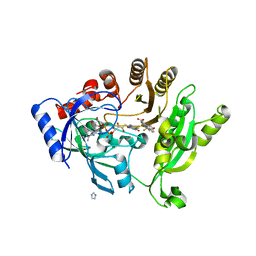 | | Crystal structure of M. tuberculosis DprE1 in complex with the non-covalent inhibitor Ty21c | | Descriptor: | 3-[(4-methoxybenzyl)amino]-6-(trifluoromethyl)quinoxaline-2-carboxylic acid, FLAVIN-ADENINE DINUCLEOTIDE, IMIDAZOLE, ... | | Authors: | Neres, J, Pojer, F, Cole, S.T. | | Deposit date: | 2014-04-01 | | Release date: | 2014-12-10 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | 2-Carboxyquinoxalines Kill Mycobacterium tuberculosis through Noncovalent Inhibition of DprE1.
Acs Chem.Biol., 10, 2015
|
|
3CXZ
 
 | |
5J5R
 
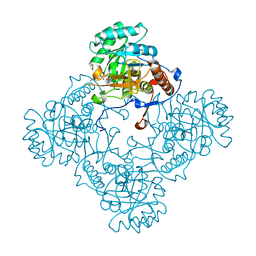 | | M. thermoresistible GuaB2 delta-CBS in complex with inhibitor VCC234718 | | Descriptor: | INOSINIC ACID, Inosine-5'-monophosphate dehydrogenase,Inosine-5'-monophosphate dehydrogenase, cyclohexyl{4-[(isoquinolin-5-yl)sulfonyl]piperazin-1-yl}methanone | | Authors: | Pacitto, A, Ascher, D.B, Blundell, T.L. | | Deposit date: | 2016-04-03 | | Release date: | 2016-10-19 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The Inosine Monophosphate Dehydrogenase, GuaB2, Is a Vulnerable New Bactericidal Drug Target for Tuberculosis.
ACS Infect Dis, 3, 2017
|
|
2BMX
 
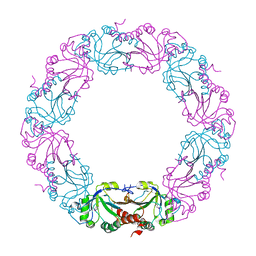 | | Mycobacterium tuberculosis AhpC | | Descriptor: | ALKYL HYDROPEROXIDASE C | | Authors: | Guimaraes, B.G, Alzari, P.M. | | Deposit date: | 2005-03-16 | | Release date: | 2005-05-10 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure and Mechanism of the Alkyl Hydroperoxidase Ahpc, a Key Element of the Mycobacterium Tuberculosis Defense System Against Oxidative Stress.
J.Biol.Chem., 280, 2005
|
|
2CKD
 
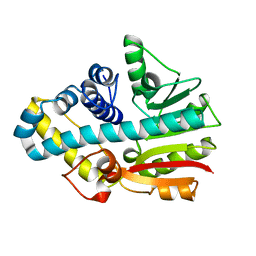 | | Crystal structure of ML2640 from Mycobacterium leprae | | Descriptor: | PUTATIVE S-ADENOSYL-L-METHIONINE-DEPENDENT METHYLTRANSFERASE ML2640 | | Authors: | Grana, M, Buschiazzo, A, Wehenkel, A, Haouz, A, Alzari, P.M. | | Deposit date: | 2006-04-17 | | Release date: | 2007-05-29 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The Crystal Structure of M. Leprae Ml2640C Defines a Large Family of Putative S-Adenosylmethionine- Dependent Methyltransferases in Mycobacteria.
Protein Sci., 16, 2007
|
|
2G9W
 
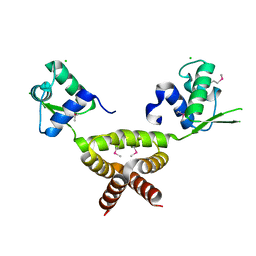 | | Crystal Structure of Rv1846c, a Putative Transcriptional Regulatory Protein of Mycobacterium Tuberculosis | | Descriptor: | CHLORIDE ION, conserved hypothetical protein | | Authors: | Saul, F.A, Haouz, A, Fiez-Vandal, C, Shepard, W, Alzari, P.M. | | Deposit date: | 2006-03-07 | | Release date: | 2007-03-13 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Genome-wide regulon and crystal structure of BlaI (Rv1846c) from Mycobacterium tuberculosis
Mol.Microbiol., 71, 2009
|
|
2UYQ
 
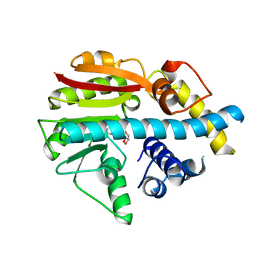 | | Crystal structure of ML2640c from Mycobacterium leprae in complex with S-adenosylmethionine | | Descriptor: | HYPOTHETICAL PROTEIN ML2640, S-ADENOSYLMETHIONINE | | Authors: | Grana, M, Buschiazzo, A, Wehenkel, A, Haouz, A, Miras, I, Shepard, W, Alzari, P.M. | | Deposit date: | 2007-04-11 | | Release date: | 2007-08-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Crystal Structure of M. Leprae Ml2640C Defines a Large Family of Putative S-Adenosylmethionine- Dependent Methyltransferases in Mycobacteria.
Protein Sci., 16, 2007
|
|
2WZV
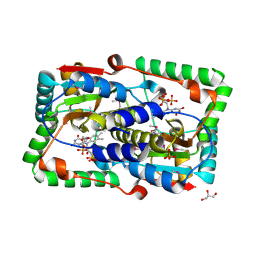 
 | | Crystal structure of the FMN-dependent nitroreductase NfnB from Mycobacterium smegmatis | | Descriptor: | FLAVIN MONONUCLEOTIDE, GLYCEROL, NFNB PROTEIN, ... | | Authors: | Bellinzoni, M, Manina, G, Riccardi, G, Alzari, P.M. | | Deposit date: | 2009-12-03 | | Release date: | 2010-07-14 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Biological and Structural Characterization of the Mycobacterium Smegmatis Nitroreductase Nfnb, and its Role in Benzothiazinone Resistance
Mol.Microbiol., 77, 2010
|
|
2UYO
 
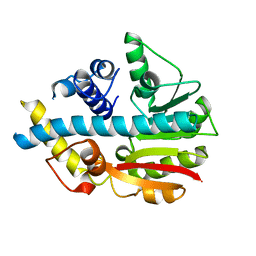 | | Crystal structure of ML2640c from Mycobacterium leprae in an hexagonal crystal form | | Descriptor: | HYPOTHETICAL PROTEIN ML2640 | | Authors: | Grana, M, Buschiazzo, A, Wehenkel, A, Haouz, A, Miras, I, Shepard, W, Alzari, P.M. | | Deposit date: | 2007-04-11 | | Release date: | 2007-08-07 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The Crystal Structure of M. Leprae Ml2640C Defines a Large Family of Putative S-Adenosylmethionine- Dependent Methyltransferases in Mycobacteria.
Protein Sci., 16, 2007
|
|
2WZW
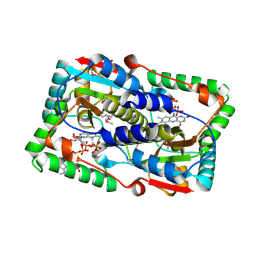 
 | | Crystal structure of the FMN-dependent nitroreductase NfnB from Mycobacterium smegmatis in complex with NADPH | | Descriptor: | FLAVIN MONONUCLEOTIDE, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, NFNB PROTEIN, ... | | Authors: | Bellinzoni, M, Manina, G, Riccardi, G, Alzari, P.M. | | Deposit date: | 2009-12-03 | | Release date: | 2010-07-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Biological and Structural Characterization of the Mycobacterium Smegmatis Nitroreductase Nfnb, and its Role in Benzothiazinone Resistance
Mol.Microbiol., 77, 2010
|
|
6TOB
 
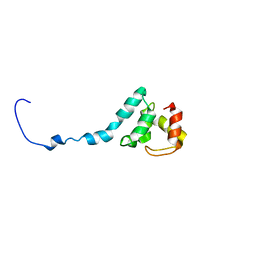 | |
4XWP
 
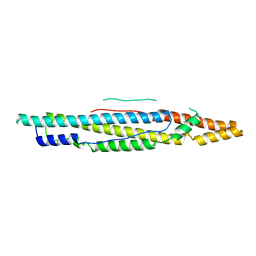 | |
4XY3
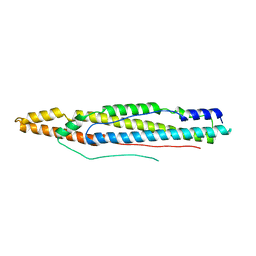 
 | | Structure of ESX-1 secreted protein EspB | | Descriptor: | ESX-1 secretion-associated protein EspB | | Authors: | Wagner, J.M, Korotkov, K.V. | | Deposit date: | 2015-02-01 | | Release date: | 2015-02-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.04 Å) | | Cite: | Structure of EspB, a secreted substrate of the ESX-1 secretion system of Mycobacterium tuberculosis.
J.Struct.Biol., 191, 2015
|
|
4XXN
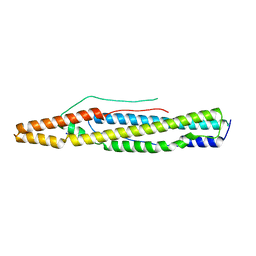 
 | | Structure of PE-PPE domains of ESX-1 secreted protein EspB, I222 | | Descriptor: | CHLORIDE ION, ESX-1 secretion-associated protein EspB, SODIUM ION | | Authors: | Korotkov, K.V. | | Deposit date: | 2015-01-30 | | Release date: | 2015-02-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Structure of EspB, a secreted substrate of the ESX-1 secretion system of Mycobacterium tuberculosis.
J.Struct.Biol., 191, 2015
|
|
4XXX
 
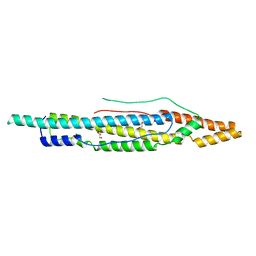 | | Structure of PE-PPE domains of ESX-1 secreted protein EspB, C2221 | | Descriptor: | CHLORIDE ION, ESX-1 secretion-associated protein EspB, GLYCEROL | | Authors: | Piton, J, Pojer, F, Korotkov, K.V. | | Deposit date: | 2015-01-30 | | Release date: | 2015-02-18 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure of EspB, a secreted substrate of the ESX-1 secretion system of Mycobacterium tuberculosis.
J.Struct.Biol., 191, 2015
|
|
4ZDJ
 
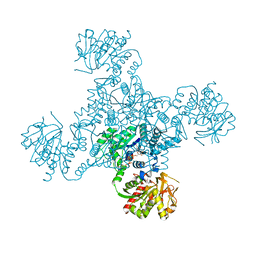 | | Crystal structure of the M. tuberculosis CTP synthase PyrG in complex with two UTP molecules | | Descriptor: | CTP synthase, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Bellinzoni, M, Barilone, N, Alzari, P.M. | | Deposit date: | 2015-04-17 | | Release date: | 2015-07-01 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Thiophenecarboxamide Derivatives Activated by EthA Kill Mycobacterium tuberculosis by Inhibiting the CTP Synthetase PyrG.
Chem.Biol., 22, 2015
|
|
4ZDI
 
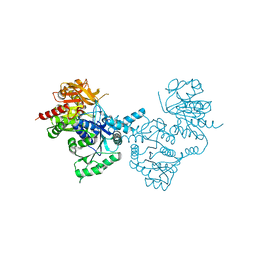 | |
4ZDK
 
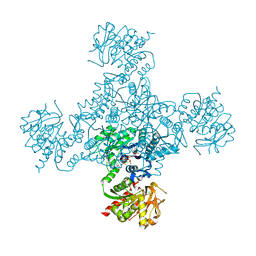 | | Crystal structure of the M. tuberculosis CTP synthase PyrG in complex with UTP, AMP-PCP and oxonorleucine | | Descriptor: | 5-OXO-L-NORLEUCINE, CTP synthase, MAGNESIUM ION, ... | | Authors: | Bellinzoni, M, Barilone, N, Alzari, P.M. | | Deposit date: | 2015-04-17 | | Release date: | 2015-07-01 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.49 Å) | | Cite: | Thiophenecarboxamide Derivatives Activated by EthA Kill Mycobacterium tuberculosis by Inhibiting the CTP Synthetase PyrG.
Chem.Biol., 22, 2015
|
|
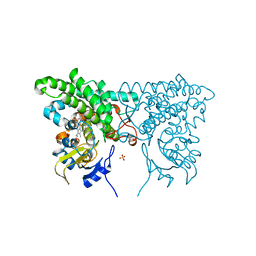
3CY0
 
 | |
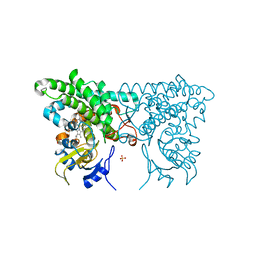
3CXY
 
 | |
3CY1
 
 | |
3CXV
 
 | |
3CXX
 
 | |
4BYY
 
 | | Apo GlxR | | Descriptor: | GLYCEROL, PHOSPHATE ION, TRANSCRIPTIONAL REGULATOR, ... | | Authors: | Pohl, E, Cann, M.J, Townsend, P.D, Money, V.A. | | Deposit date: | 2013-07-21 | | Release date: | 2014-07-30 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | The Crystal Structures of Apo and Camp-Bound Glxr from Corynebacterium Glutamicum Reveal Structural and Dynamic Changes Upon Camp Binding in Crp/Fnr Family Transcription Factors.
Plos One, 9, 2014
|
|
4CYD
 
 | | GlxR bound to cAMP | | Descriptor: | ADENOSINE-3',5'-CYCLIC-MONOPHOSPHATE, GLYCEROL, PROBABLE EXPRESSION TAG, ... | | Authors: | Townsend, P.D, Bott, M, Cann, M.J, Pohl, E. | | Deposit date: | 2014-04-10 | | Release date: | 2014-12-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | The Crystal Structures of Apo and Camp-Bound Glxr from Corynebacterium Glutamicum Reveal Structural and Dynamic Changes Upon Camp Binding in Crp/Fnr Family Transcription Factors.
Plos One, 9, 2014
|
|
