1SKU
 
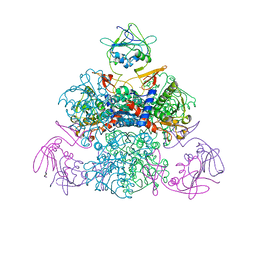 | | E. coli Aspartate Transcarbamylase 240's Loop Mutant (K244N) | | Descriptor: | Aspartate carbamoyltransferase catalytic chain, Aspartate carbamoyltransferase regulatory chain, MALONATE ION, ... | | Authors: | Alam, N, Stieglitz, K.A, Caban, M.D, Gourinath, S, Tsuruta, H, Kantrowitz, E.R. | | Deposit date: | 2004-03-05 | | Release date: | 2004-03-30 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | 240s Loop Interactions Stabilize the T State of Escherichia coli Aspartate Transcarbamoylase.
J.Biol.Chem., 279, 2004
|
|
2NXQ
 
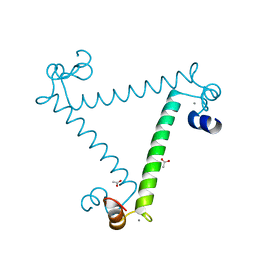 | | Crystal structure of calcium binding protein 1 from Entamoeba histolytica: a novel arrangement of EF hand motifs | | Descriptor: | ACETATE ION, CALCIUM ION, Calcium-binding protein | | Authors: | Kumar, S, Padhan, N, Alam, N, Gourinath, S. | | Deposit date: | 2006-11-18 | | Release date: | 2007-08-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of calcium binding protein-1 from Entamoeba histolytica: A novel arrangement of EF hand motifs.
Proteins, 68, 2007
|
|
2PQM
 
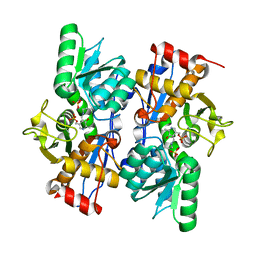 | |
8IZH
 
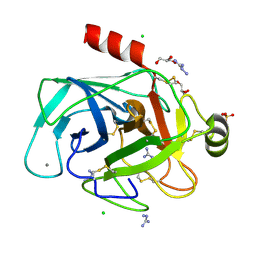 | | Crystal structure of trypsin-aminoguanidine complex at 2.30 Angstroms resolution | | Descriptor: | AMINOGUANIDINE, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Ahmad, M.S, Kalam, N, Akbar, Z, Rasheed, S, Choudhary, M.I. | | Deposit date: | 2023-04-07 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis for the binding of famotidine, cimetidine, guanidine, and pimagedine with serine protease.
Biochem.Biophys.Res.Commun., 733, 2024
|
|
8IZI
 
 | | Crystal structure of trypsin-cimetidine complex at 2.10 Angstroms resolution | | Descriptor: | 2-cyano-1-methyl-3-[2-[(5-methyl-1~{H}-imidazol-4-yl)methylsulfanyl]ethyl]guanidine, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Ahmad, M.S, Kalam, N, Akbar, Z, Rasheed, S, Choudhary, M.I. | | Deposit date: | 2023-04-07 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural basis for the binding of famotidine, cimetidine, guanidine, and pimagedine with serine protease.
Biochem.Biophys.Res.Commun., 733, 2024
|
|
8IYV
 
 | | Crystal structure of trypsin-famotidine complex at 2.10 Angstroms resolution | | Descriptor: | CALCIUM ION, CHLORIDE ION, Cationic trypsin, ... | | Authors: | Ahmad, M.S, Kalam, N, Akbar, Z, Rasheed, S, Choudhary, M.I. | | Deposit date: | 2023-04-06 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural basis for the binding of famotidine, cimetidine, guanidine, and pimagedine with serine protease.
Biochem.Biophys.Res.Commun., 733, 2024
|
|
8IZK
 
 | | Crystal structure of trypsin-guanidine complex at 2.05 Angstroms resolution | | Descriptor: | CALCIUM ION, CHLORIDE ION, Cationic trypsin, ... | | Authors: | Ahmad, M.S, Kalam, N, Akbar, Z, Rasheed, S, Choudhary, M.I. | | Deposit date: | 2023-04-07 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis for the binding of famotidine, cimetidine, guanidine, and pimagedine with serine protease.
Biochem.Biophys.Res.Commun., 733, 2024
|
|
1B1U
 
 | |
5XOP
 
 | | Crystal Structure of N-terminal domain EhCaBP1 EF-2 mutant | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, CALCIUM ION, Calcium-binding protein 1 (EhCBP1), ... | | Authors: | Kumar, S, Gourinath, S. | | Deposit date: | 2017-05-30 | | Release date: | 2017-12-06 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of calcium binding protein-1 from Entamoeba histolytica: a novel arrangement of EF hand motifs.
Proteins, 68, 2007
|
|
3BM5
 
 | | Crystal structure of O-acetyl-serine sulfhydrylase from Entamoeba histolytica in complex with cysteine | | Descriptor: | CYSTEINE, Cysteine synthase, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Krishna, C, Kumar, M, Kumar, S, Gourinath, S. | | Deposit date: | 2007-12-12 | | Release date: | 2008-04-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of native O-acetyl-serine sulfhydrylase from Entamoeba histolytica and its complex with cysteine: structural evidence for cysteine binding and lack of interactions with serine acetyl transferase.
Proteins, 72, 2008
|
|
5N71
 
 | | CRYSTAL STRUCTURE OF MATURE CATHEPSIN D FROM THE TICK IXODES RICINUS (IRCD1) | | Descriptor: | DI(HYDROXYETHYL)ETHER, Putative cathepsin d, SULFATE ION | | Authors: | Brynda, J, Hanova, I, Hobizalova, R, Mares, M. | | Deposit date: | 2017-02-17 | | Release date: | 2017-12-27 | | Last modified: | 2018-03-28 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Novel Structural Mechanism of Allosteric Regulation of Aspartic Peptidases via an Evolutionarily Conserved Exosite.
Cell Chem Biol, 25, 2018
|
|
5N7Q
 
 | | CRYSTAL STRUCTURE OF MATURE CATHEPSIN D FROM THE TICK IXODES RICINUS (IRCD1) IN COMPLEX WITH THE INHIBITOR PEPSTATIN A | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, PEPSTATIN A, ... | | Authors: | Brynda, J, Hanova, I, Hobizalova, R, Mares, M. | | Deposit date: | 2017-02-21 | | Release date: | 2017-12-27 | | Last modified: | 2018-06-13 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Novel Structural Mechanism of Allosteric Regulation of Aspartic Peptidases via an Evolutionarily Conserved Exosite.
Cell Chem Biol, 25, 2018
|
|
5N70
 
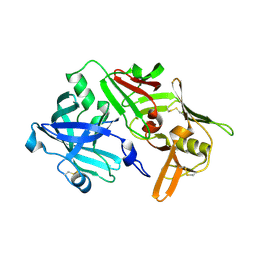 | | CRYSTAL STRUCTURE OF MATURE CATHEPSIN D FROM THE TICK IXODES RICINUS (IRCD1) IN COMPLEX WITH THE N-TERMINAL OCTAPEPTIDE OF THE PROPEPTID | | Descriptor: | ALA-PHE-ARG-ILE-PRO-LEU-THR-ARG, Putative cathepsin d | | Authors: | Brynda, J, Hanova, I, Hobizalova, R, Mares, M. | | Deposit date: | 2017-02-17 | | Release date: | 2017-12-27 | | Last modified: | 2018-03-28 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Novel Structural Mechanism of Allosteric Regulation of Aspartic Peptidases via an Evolutionarily Conserved Exosite.
Cell Chem Biol, 25, 2018
|
|
5N7N
 
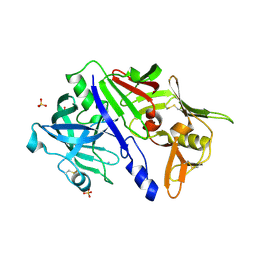 | | CRYSTAL STRUCTURE OF CATHEPSIN D ZYMOGEN FROM THE TICK IXODES RICINUS (IRCD1) | | Descriptor: | AMMONIUM ION, Putative cathepsin d, SULFATE ION | | Authors: | Brynda, J, Hanova, I, Hobizalova, R, Mares, M. | | Deposit date: | 2017-02-21 | | Release date: | 2017-12-27 | | Last modified: | 2018-03-28 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Novel Structural Mechanism of Allosteric Regulation of Aspartic Peptidases via an Evolutionarily Conserved Exosite.
Cell Chem Biol, 25, 2018
|
|
8CDE
 
 | |
8CDD
 
 | |
8CCX
 
 | | Human SOD1 in complex with S-XL6 cross-linker | | Descriptor: | COPPER (II) ION, DIMETHYL SULFOXIDE, SULFATE ION, ... | | Authors: | Antonyuk, S.V, Hossain, A, Agar, J.N, Hasnain, S.S. | | Deposit date: | 2023-01-27 | | Release date: | 2023-12-13 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.665 Å) | | Cite: | Evaluating protein cross-linking as a therapeutic strategy to stabilize SOD1 variants in a mouse model of familial ALS.
Plos Biol., 22, 2024
|
|
4LQQ
 
 | |
4LQP
 
 | |
4LQS
 
 | | Crystal structure of the Cbk1-Mob2 kinase-coactivator complex | | Descriptor: | CBK1 kinase activator protein MOB2, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, Serine/threonine-protein kinase CBK1 | | Authors: | Gogl, G, Remenyi, A. | | Deposit date: | 2013-07-19 | | Release date: | 2014-07-30 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | The Structure of an NDR/LATS Kinase-Mob Complex Reveals a Novel Kinase-Coactivator System and Substrate Docking Mechanism.
Plos Biol., 13, 2015
|
|
4HZD
 
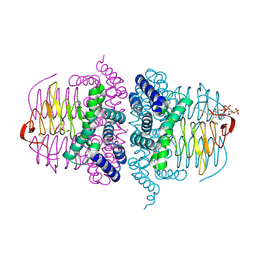 | |
8Q6M
 
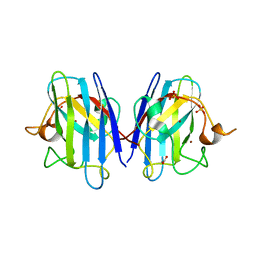 | | Human SOD1 low dose data collecton | | Descriptor: | ACETATE ION, COPPER (II) ION, SULFATE ION, ... | | Authors: | Antonyuk, S.V, Hossain, A, Agar, J.N, Hasnain, S.S. | | Deposit date: | 2023-08-14 | | Release date: | 2023-12-13 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Evaluating protein cross-linking as a therapeutic strategy to stabilize SOD1 variants in a mouse model of familial ALS.
Plos Biol., 22, 2024
|
|
4HZC
 
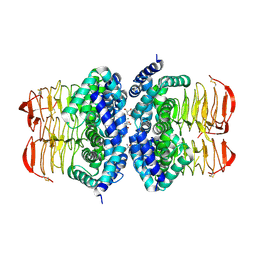 | |
1XJW
 
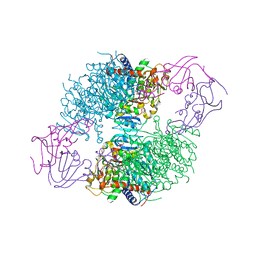 | | The Structure of E. coli Aspartate Transcarbamoylase Q137A Mutant in The R-State | | Descriptor: | Aspartate carbamoyltransferase catalytic chain, Aspartate carbamoyltransferase regulatory chain, N-(PHOSPHONACETYL)-L-ASPARTIC ACID, ... | | Authors: | Stieglitz, K.A, Alam, N, Xia, J, Gourinath, S, Tsuruta, H, Kantrowitz, E.R. | | Deposit date: | 2004-09-25 | | Release date: | 2005-05-24 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.71 Å) | | Cite: | A Single Amino Acid Substitution in the Active Site of Escherichia coli Aspartate Transcarbamoylase Prevents the Allosteric Transition.
J.Mol.Biol., 349, 2005
|
|
