6X0H
 | |
6X0J
 | |
6X0I
 
 | |
6X9B
 
 | |
6X9C
 
 | |
6XOZ
 
 | |
6X99
 
 | |
6XP0
 
 | |
6XP3
 
 | |
6X9A
 
 | |
6XP1
 
 | |
6X9D
 
 | |
6XP2
 
 | |
4E3X
 
 | | Crystal Structure of Mus musculus 1-pyrroline-5-carboxylate dehydrogenase cryoprotected in proline | | Descriptor: | 3,6,9,12,15,18-HEXAOXAICOSANE, Delta-1-pyrroline-5-carboxylate dehydrogenase, mitochondrial, ... | | Authors: | Tanner, J.J, Pemberton, T.A. | | Deposit date: | 2012-03-10 | | Release date: | 2012-07-25 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.24 Å) | | Cite: | Proline: Mother Nature's cryoprotectant applied to protein crystallography.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4E3W
 
 | |
4E3V
 
 | |
4E3U
 
 | |
5UAW
 
 | | Structure of apo human PYCR-1 crystallized in space group P21212 | | Descriptor: | Pyrroline-5-carboxylate reductase 1, mitochondrial, SULFATE ION | | Authors: | Tanner, J.J. | | Deposit date: | 2016-12-20 | | Release date: | 2017-03-15 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Resolving the cofactor-binding site in the proline biosynthetic enzyme human pyrroline-5-carboxylate reductase 1.
J. Biol. Chem., 292, 2017
|
|
5UAT
 
 | | Structure of human PYCR-1 complexed with NADPH | | Descriptor: | DI(HYDROXYETHYL)ETHER, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Pyrroline-5-carboxylate reductase 1, ... | | Authors: | Tanner, J.J. | | Deposit date: | 2016-12-20 | | Release date: | 2017-03-15 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Resolving the cofactor-binding site in the proline biosynthetic enzyme human pyrroline-5-carboxylate reductase 1.
J. Biol. Chem., 292, 2017
|
|
5UAU
 
 | | Structure of human PYCR-1 complexed with proline | | Descriptor: | PROLINE, Pyrroline-5-carboxylate reductase 1, mitochondrial, ... | | Authors: | Tanner, J.J. | | Deposit date: | 2016-12-20 | | Release date: | 2017-03-15 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Resolving the cofactor-binding site in the proline biosynthetic enzyme human pyrroline-5-carboxylate reductase 1.
J. Biol. Chem., 292, 2017
|
|
5UAV
 
 | | Structure of human PYCR-1 complexed with NADPH and L-tetrahydrofuroic acid | | Descriptor: | DI(HYDROXYETHYL)ETHER, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Pyrroline-5-carboxylate reductase 1, ... | | Authors: | Tanner, J.J. | | Deposit date: | 2016-12-20 | | Release date: | 2017-03-15 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Resolving the cofactor-binding site in the proline biosynthetic enzyme human pyrroline-5-carboxylate reductase 1.
J. Biol. Chem., 292, 2017
|
|
5UAX
 
 | | Structure of apo human PYCR-1 crystallized in space group C2 | | Descriptor: | CHLORIDE ION, Pyrroline-5-carboxylate reductase 1, mitochondrial | | Authors: | Tanner, J.J. | | Deposit date: | 2016-12-20 | | Release date: | 2017-03-15 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Resolving the cofactor-binding site in the proline biosynthetic enzyme human pyrroline-5-carboxylate reductase 1.
J. Biol. Chem., 292, 2017
|
|
5UR2
 
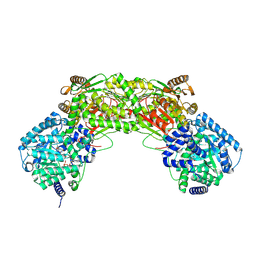 | |
5UX5
 
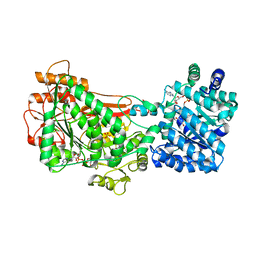 | | Structure of Proline Utilization A (PutA) from Corynebacterium freiburgense | | Descriptor: | BIFUNCTIONAL PROTEIN Proline utilization A (PutA), FLAVIN-ADENINE DINUCLEOTIDE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Tanner, J.J. | | Deposit date: | 2017-02-22 | | Release date: | 2017-04-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure and characterization of a class 3B proline utilization A: Ligand-induced dimerization and importance of the C-terminal domain for catalysis.
J. Biol. Chem., 292, 2017
|
|
1P7K
 
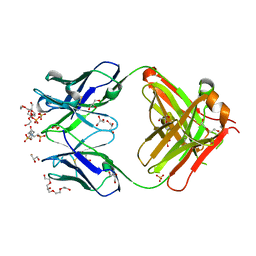 | | Crystal structure of an anti-ssDNA antigen-binding fragment (Fab) bound to 4-(2-Hydroxyethyl)piperazine-1-ethanesulfonic acid (HEPES) | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Schuermann, J.P, Henzl, M.T, Deutscher, S.L, Tanner, J.J. | | Deposit date: | 2003-05-02 | | Release date: | 2004-05-11 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structure of an anti-DNA fab complexed with a non-DNA ligand provides insights into cross-reactivity and molecular mimicry.
Proteins, 57, 2004
|
|
