2E0M
 
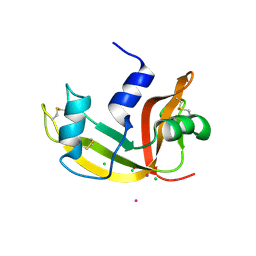 | | Mutant Human Ribonuclease 1 (T24L, Q28L, R31L, R32L) | | Descriptor: | CADMIUM ION, CHLORIDE ION, Ribonuclease | | Authors: | Yamada, H, Tamada, T, Kosaka, M, Kuroki, R. | | Deposit date: | 2006-10-10 | | Release date: | 2007-08-28 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | 'Crystal lattice engineering,' an approach to engineer protein crystal contacts by creating intermolecular symmetry: crystallization and structure determination of a mutant human RNase 1 with a hydrophobic interface of leucines
Protein Sci., 16, 2007
|
|
6INH
 
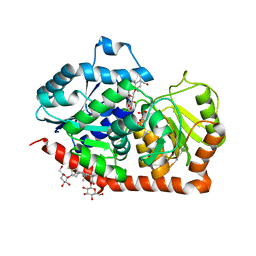 | | A glycosyltransferase with UDP and the substrate | | Descriptor: | 1-O-[(8alpha,9beta,10alpha,13alpha)-13-(beta-D-glucopyranosyloxy)-18-oxokaur-16-en-18-yl]-beta-D-glucopyranose, GLYCEROL, UDP-glycosyltransferase 76G1, ... | | Authors: | Zhu, X. | | Deposit date: | 2018-10-25 | | Release date: | 2019-07-31 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Hydrophobic recognition allows the glycosyltransferase UGT76G1 to catalyze its substrate in two orientations.
Nat Commun, 10, 2019
|
|
2YV0
 
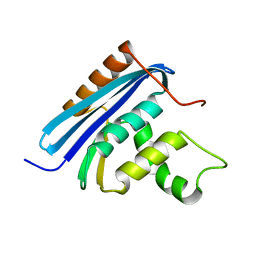 | | Structural and Thermodynamic Analyses of E. coli ribonuclease HI Variant with Quintuple Thermostabilizing Mutations | | Descriptor: | Ribonuclease HI | | Authors: | Haruki, M, Motegi, T, Tadokoro, T, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2007-04-06 | | Release date: | 2008-03-18 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural and thermodynamic analyses of Escherichia coli RNase HI variant with quintuple thermostabilizing mutations.
Febs J., 274, 2007
|
|
6E7D
 
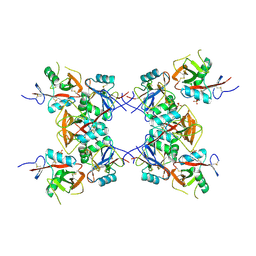 | | Structure of the inhibitory NKR-P1B receptor bound to the host-encoded ligand, Clr-b | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, C-type lectin domain family 2 member D, Killer cell lectin-like receptor subfamily B member 1B allele B, ... | | Authors: | Balaji, G.R, Rossjohn, J, Berry, R. | | Deposit date: | 2018-07-26 | | Release date: | 2018-10-24 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Recognition of host Clr-b by the inhibitory NKR-P1B receptor provides a basis for missing-self recognition.
Nat Commun, 9, 2018
|
|
6INI
 
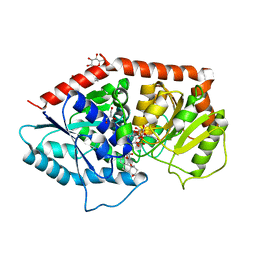 | | a glycosyltransferase complex with UDP and the product | | Descriptor: | (8alpha,9beta,10alpha,13alpha)-13-{[beta-D-glucopyranosyl-(1->2)-[beta-D-glucopyranosyl-(1->3)]-beta-D-glucopyranosyl]oxy}kaur-16-en-18-oic acid, 1-O-[(8alpha,9beta,10alpha,13alpha)-13-(beta-D-glucopyranosyloxy)-18-oxokaur-16-en-18-yl]-beta-D-glucopyranose, GLYCEROL, ... | | Authors: | Zhu, X, Yang, T, Naismith, J.H. | | Deposit date: | 2018-10-25 | | Release date: | 2019-07-31 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Hydrophobic recognition allows the glycosyltransferase UGT76G1 to catalyze its substrate in two orientations.
Nat Commun, 10, 2019
|
|
5BUP
 
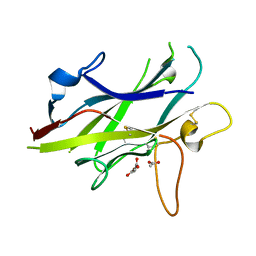 | | Crystal structure of the ZP-C domain of mouse ZP2 | | Descriptor: | ACETATE ION, Zona pellucida sperm-binding protein 2 | | Authors: | Nishimura, K, Jovine, L. | | Deposit date: | 2015-06-04 | | Release date: | 2016-01-27 | | Last modified: | 2017-09-06 | | Method: | X-RAY DIFFRACTION (2.251 Å) | | Cite: | A structured interdomain linker directs self-polymerization of human uromodulin.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
7Z6Q
 
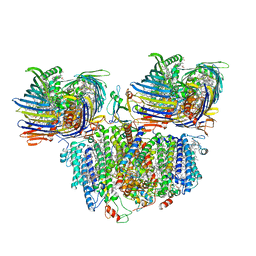 | |
2DT9
 
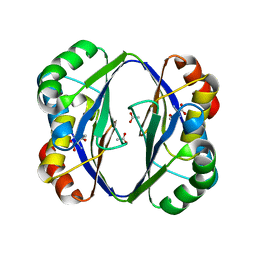 | | Crystal structure of the regulatory subunit of aspartate kinase from Thermus flavus | | Descriptor: | ACETATE ION, Aspartokinase, THREONINE | | Authors: | Yoshida, A, Tomita, T, Fushinobu, S, Kuzuyama, T, Nishiyama, M. | | Deposit date: | 2006-07-11 | | Release date: | 2007-07-17 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal structures of the regulatory subunit of Thr-sensitive aspartate kinase from Thermus thermophilus
Febs J., 276, 2009
|
|
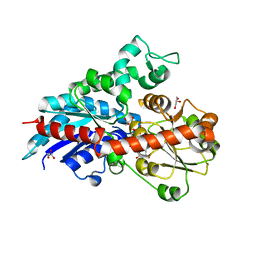
6ING
 
 | |
7ZKO
 
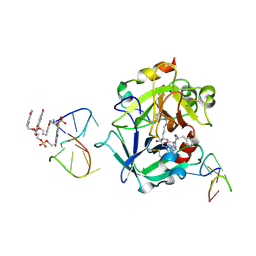 | | X-ray structure of the complex between human alpha thrombin and a pseudo-cyclic thrombin binding aptamer (TBA-NNp/DDp) - Crystal form delta | | Descriptor: | 3-[13-methyl-5,7,12,14-tetrakis(oxidanylidene)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1(15),2,4(16),8,10-pentaen-6-yl]propyl 3-[5,7,12,14-tetrakis(oxidanylidene)-13-(3-oxidanylpropyl)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1,3,8(16),9,11(15)-pentaen-6-yl]propyl hydrogen phosphate, 3-[5-[3-bis(oxidanyl)phosphanyloxypropoxy]naphthalen-1-yl]oxypropyl 3-(5-oxidanylnaphthalen-1-yl)oxypropyl hydrogen phosphate, D-phenylalanyl-N-[(2S,3S)-6-{[amino(iminio)methyl]amino}-1-chloro-2-hydroxyhexan-3-yl]-L-prolinamide, ... | | Authors: | Troisi, R, Sica, F. | | Deposit date: | 2022-04-13 | | Release date: | 2022-11-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A terminal functionalization strategy reveals unusual binding abilities of anti-thrombin anticoagulant aptamers.
Mol Ther Nucleic Acids, 30, 2022
|
|
7ZKN
 
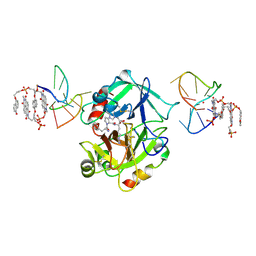 | | X-ray structure of the complex between human alpha thrombin and a pseudo-cyclic thrombin binding aptamer (TBA-NNp/DDp) - Crystal form gamma | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3-[13-methyl-5,7,12,14-tetrakis(oxidanylidene)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1(15),2,4(16),8,10-pentaen-6-yl]propyl 3-[5,7,12,14-tetrakis(oxidanylidene)-13-(3-oxidanylpropyl)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1,3,8(16),9,11(15)-pentaen-6-yl]propyl hydrogen phosphate, 3-[5-[3-bis(oxidanyl)phosphanyloxypropoxy]naphthalen-1-yl]oxypropyl 3-(5-oxidanylnaphthalen-1-yl)oxypropyl hydrogen phosphate, ... | | Authors: | Troisi, R, Sica, F. | | Deposit date: | 2022-04-13 | | Release date: | 2022-11-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.03 Å) | | Cite: | A terminal functionalization strategy reveals unusual binding abilities of anti-thrombin anticoagulant aptamers.
Mol Ther Nucleic Acids, 30, 2022
|
|
7ZKM
 
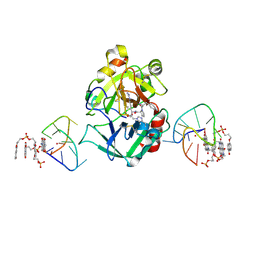 | | X-ray structure of the complex between human alpha thrombin and a pseudo-cyclic thrombin binding aptamer (TBA-NNp/DDp) - Crystal form beta | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3-[13-methyl-5,7,12,14-tetrakis(oxidanylidene)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1(15),2,4(16),8,10-pentaen-6-yl]propyl 3-[5,7,12,14-tetrakis(oxidanylidene)-13-(3-oxidanylpropyl)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1,3,8(16),9,11(15)-pentaen-6-yl]propyl hydrogen phosphate, 3-[5-[3-bis(oxidanyl)phosphanyloxypropoxy]naphthalen-1-yl]oxypropyl 3-(5-oxidanylnaphthalen-1-yl)oxypropyl hydrogen phosphate, ... | | Authors: | Troisi, R, Sica, F. | | Deposit date: | 2022-04-13 | | Release date: | 2022-11-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A terminal functionalization strategy reveals unusual binding abilities of anti-thrombin anticoagulant aptamers.
Mol Ther Nucleic Acids, 30, 2022
|
|
7ZKL
 
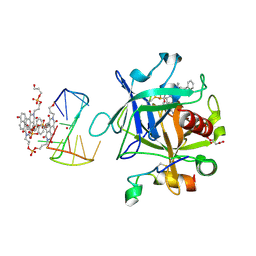 | | X-ray structure of the complex between human alpha thrombin and a pseudo-cyclic thrombin binding aptamer (TBA-NNp/DDp) - Crystal form alpha | | Descriptor: | (2S)-2-hydroxybutanedioic acid, 3-[13-methyl-5,7,12,14-tetrakis(oxidanylidene)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1(15),2,4(16),8,10-pentaen-6-yl]propyl 3-[5,7,12,14-tetrakis(oxidanylidene)-13-(3-oxidanylpropyl)-6,13-diazatetracyclo[6.6.2.0^{4,16}.0^{11,15}]hexadeca-1,3,8(16),9,11(15)-pentaen-6-yl]propyl hydrogen phosphate, 3-[5-[3-bis(oxidanyl)phosphanyloxypropoxy]naphthalen-1-yl]oxypropyl 3-(5-oxidanylnaphthalen-1-yl)oxypropyl hydrogen phosphate, ... | | Authors: | Troisi, R, Sica, F. | | Deposit date: | 2022-04-13 | | Release date: | 2022-11-30 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.18 Å) | | Cite: | A terminal functionalization strategy reveals unusual binding abilities of anti-thrombin anticoagulant aptamers.
Mol Ther Nucleic Acids, 30, 2022
|
|
3D7G
 
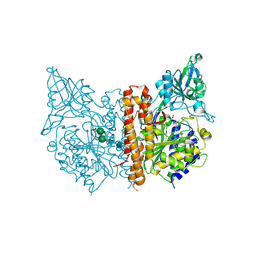 | |
3D7F
 
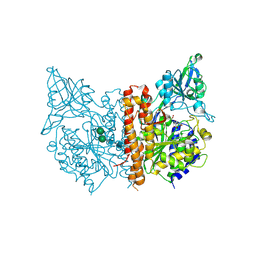 | |
5ZFU
 
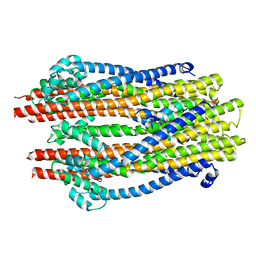 | | Structure of the ExbB/ExbD hexameric complex (ExbB6ExbD3TM) | | Descriptor: | 22-mer peptide from Biopolymer transport protein ExbD, Biopolymer transport protein ExbB | | Authors: | Yonekura, K, Yamashita, Y, Matsuoka, R, Maki-Yonekura, S. | | Deposit date: | 2018-03-07 | | Release date: | 2018-05-09 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (6.7 Å) | | Cite: | Hexameric and pentameric complexes of the ExbBD energizer in the Ton system.
Elife, 7, 2018
|
|
7WJT
 
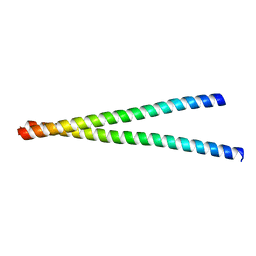 | |
6T7F
 
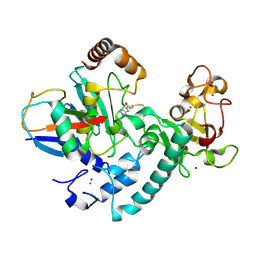 | | RCR E3 ligase E2-Ubiquitin transthiolation intermediate | | Descriptor: | 3,3-bis(sulfanyl)-~{N}-(1~{H}-1,2,3-triazol-4-ylmethyl)propanamide, E3 ubiquitin-protein ligase MYCBP2, Polyubiquitin-C, ... | | Authors: | Mabbitt, P.D, Virdee, S. | | Deposit date: | 2019-10-21 | | Release date: | 2020-08-05 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.58 Å) | | Cite: | Structural basis for RING-Cys-Relay E3 ligase activity and its role in axon integrity.
Nat.Chem.Biol., 16, 2020
|
|
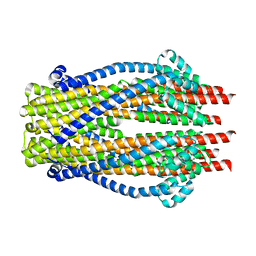
5ZFP
 | |
5ZFV
 
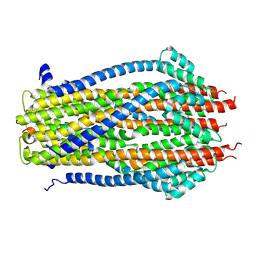 | | Structure of the ExbB/ExbD pentameric complex (ExbB5ExbD1TM) | | Descriptor: | 22-mer peptide from Biopolymer transport protein ExbD, Biopolymer transport protein ExbB | | Authors: | Yonekura, K, Yamashita, Y, Matsuoka, R, Maki-Yonekura, S. | | Deposit date: | 2018-03-07 | | Release date: | 2018-05-09 | | Method: | ELECTRON MICROSCOPY (7.1 Å) | | Cite: | Hexameric and pentameric complexes of the ExbBD energizer in the Ton system.
Elife, 7, 2018
|
|
7Y3F
 
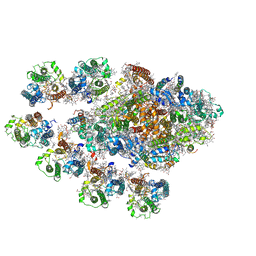 | | Structure of the Anabaena PSI-monomer-IsiA supercomplex | | Descriptor: | 1,2-DIPALMITOYL-PHOSPHATIDYL-GLYCEROLE, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, BETA-CAROTENE, ... | | Authors: | Nagao, R, Kato, K, Hamaguchi, T, Kawakami, K, Yonekura, K, Shen, J.R. | | Deposit date: | 2022-06-10 | | Release date: | 2023-03-01 | | Last modified: | 2024-05-08 | | Method: | ELECTRON MICROSCOPY (2.62 Å) | | Cite: | Structure of a monomeric photosystem I core associated with iron-stress-induced-A proteins from Anabaena sp. PCC 7120.
Nat Commun, 14, 2023
|
|
3D7H
 
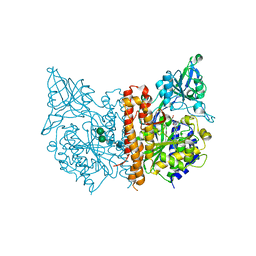 | |
4L3P
 
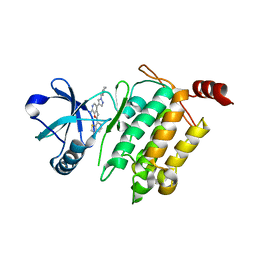 | | Crystal Structure of 2-(1-benzothiophen-7-yl)-4-[1-(piperidin-4-yl)-1H-pyrazol-4-yl]furo[2,3-c]pyridin-7-amine bound to TAK1-TAB1 | | Descriptor: | 2-(1-benzothiophen-7-yl)-4-[1-(piperidin-4-yl)-1H-pyrazol-4-yl]furo[2,3-c]pyridin-7-amine, Mitogen-activated protein kinase kinase kinase 7, TGF-beta-activated kinase 1 and MAP3K7-binding protein 1 chimera | | Authors: | Wang, J, Hornberger, K.R, Crew, A.P, Steinbacher, S, Maskos, K, Moertl, M. | | Deposit date: | 2013-06-06 | | Release date: | 2013-06-19 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.68 Å) | | Cite: | Discovery and optimization of 7-aminofuro[2,3-c]pyridine inhibitors of TAK1.
Bioorg.Med.Chem.Lett., 23, 2013
|
|
6INF
 
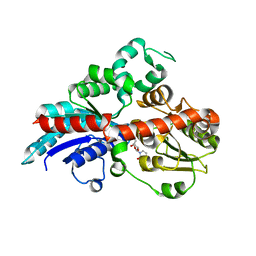 | | a glycosyltransferase complex with UDP | | Descriptor: | UDP-glycosyltransferase 76G1, URIDINE-5'-DIPHOSPHATE | | Authors: | Zhu, X, Yang, T, Naismith, J.H. | | Deposit date: | 2018-10-25 | | Release date: | 2019-07-31 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Hydrophobic recognition allows the glycosyltransferase UGT76G1 to catalyze its substrate in two orientations.
Nat Commun, 10, 2019
|
|
3D7D
 
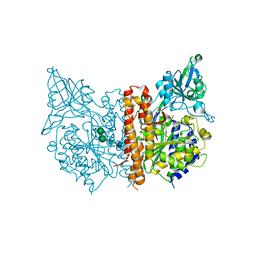 | |
