1CVR
 
 | |
1IBJ
 
 | | Crystal structure of cystathionine beta-lyase from Arabidopsis thaliana | | Descriptor: | CARBONATE ION, CYSTATHIONINE BETA-LYASE, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Breitinger, U, Clausen, T, Messerschmidt, A. | | Deposit date: | 2001-03-28 | | Release date: | 2001-04-04 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The three-dimensional structure of cystathionine beta-lyase from Arabidopsis and its substrate specificity
Plant Physiol., 126, 2001
|
|
3F6U
 
 | | Crystal structure of human Activated Protein C (APC) complexed with PPACK | | Descriptor: | CALCIUM ION, D-phenylalanyl-N-[(2S,3S)-6-{[amino(iminio)methyl]amino}-1-chloro-2-hydroxyhexan-3-yl]-L-prolinamide, SODIUM ION, ... | | Authors: | Schmidt, A.E, Padmanabhan, K, Underwood, M.C, Bode, W, Mather, T, Bajaj, S.P. | | Deposit date: | 2008-11-06 | | Release date: | 2008-11-25 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Thermodynamic linkage between the S1 site, the Na+ site, and the Ca2+ site in the protease domain of human activated protein C (APC).
J.Biol.Chem., 277, 2002
|
|
3GSS
 
 | | HUMAN GLUTATHIONE S-TRANSFERASE P1-1 IN COMPLEX WITH ETHACRYNIC ACID-GLUTATHIONE CONJUGATE | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ETHACRYNIC ACID, GLUTATHIONE, ... | | Authors: | Oakley, A.J, Rossjohn, J, Parker, M.W. | | Deposit date: | 1996-10-29 | | Release date: | 1997-11-12 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The three-dimensional structure of the human Pi class glutathione transferase P1-1 in complex with the inhibitor ethacrynic acid and its glutathione conjugate.
Biochemistry, 36, 1997
|
|
2PVB
 
 | |
2RIS
 
 | |
1NPR
 
 | | CRYSTAL STRUCTURE OF AQUIFEX AEOLICUS NUSG IN C222(1) | | Descriptor: | Transcription antitermination protein nusG | | Authors: | Knowlton, J.R, Bubunenko, M, Andrykovitch, M, Guo, W, Routzhan, K.M, Waugh, D.S, Court, D.L, Ji, X. | | Deposit date: | 2003-01-18 | | Release date: | 2003-03-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | A Spring-Loaded State of NusG in Its Functional Cycle Is Suggested by X-ray Crystallography and Supported by
Site-Directed Mutants
Biochemistry, 42, 2003
|
|
1NPP
 
 | | CRYSTAL STRUCTURE OF AQUIFEX AEOLICUS NUSG IN P2(1) | | Descriptor: | ISOPROPYL ALCOHOL, Transcription antitermination protein nusG | | Authors: | Knowlton, J.R, Bubunenko, M, Andrykovitch, M, Guo, W, Routzahn, K.M, Waugh, D.S, Court, D.L, Ji, X. | | Deposit date: | 2003-01-18 | | Release date: | 2003-03-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A Spring-Loaded State of NusG in Its Functional Cycle Is Suggested by X-ray Crystallography and Supported by
Site-Directed Mutants
Biochemistry, 42, 2003
|
|
1MCV
 
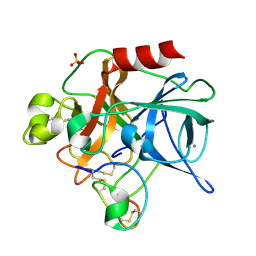 | | Crystal Structure Analysis of a Hybrid Squash Inhibitor in Complex with Porcine Pancreatic Elastase | | Descriptor: | CALCIUM ION, Elastase 1, HEI-TOE I, ... | | Authors: | Ay, J, Hilpert, K, Krauss, N, Schneider-Mergener, J, Hoehne, W. | | Deposit date: | 2002-08-06 | | Release date: | 2003-02-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure of a hybrid squash inhibitor in complex with porcine pancreatic elastase at 1.8 A resolution.
Acta Crystallogr.,Sect.D, 59, 2003
|
|
6P5G
 
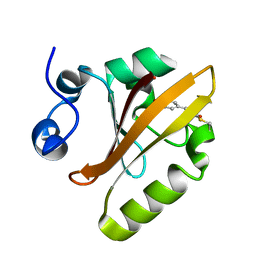 | | Photoactive Yellow Protein PYP Dark Full | | Descriptor: | Photoactive yellow protein | | Authors: | Pandey, S, Schmidt, M. | | Deposit date: | 2019-05-30 | | Release date: | 2019-09-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Time-resolved serial femtosecond crystallography at the European XFEL.
Nat.Methods, 17, 2020
|
|
6P4I
 
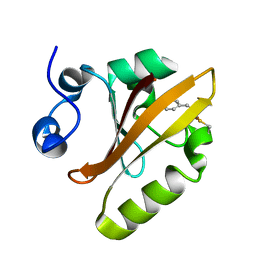 | | Photoactive Yellow Protein PYP 10ps | | Descriptor: | Photoactive yellow protein | | Authors: | Pandey, S, Schmidt, M. | | Deposit date: | 2019-05-27 | | Release date: | 2019-09-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Time-resolved serial femtosecond crystallography at the European XFEL.
Nat.Methods, 17, 2020
|
|
6P5F
 
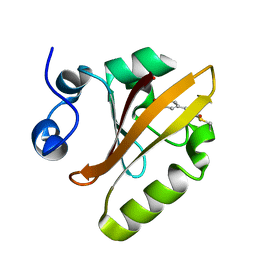 | | Photoactive Yellow Protein PYP Pure Dark | | Descriptor: | Photoactive yellow protein | | Authors: | Pandey, S, Schmidt, M. | | Deposit date: | 2019-05-30 | | Release date: | 2019-09-18 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Time-resolved serial femtosecond crystallography at the European XFEL.
Nat.Methods, 17, 2020
|
|
2RIU
 
 | | Alternative models for two crystal structures of Candida albicans 3,4-dihydroxy-2-butanone 4-phosphate synthase- alternate interpreation | | Descriptor: | 3,4-dihydroxy-2-butanone 4-phosphate synthase, RIBULOSE-5-PHOSPHATE | | Authors: | Stenkamp, R.E, Le Trong, I. | | Deposit date: | 2007-10-12 | | Release date: | 2008-01-29 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Alternative models for two crystal structures of Candida albicans 3,4-dihydroxy-2-butanone 4-phosphate synthase.
Acta Crystallogr.,Sect.D, 64, 2008
|
|
1WU1
 
 | |
1XKA
 
 | |
1XKB
 
 | |
1ZFI
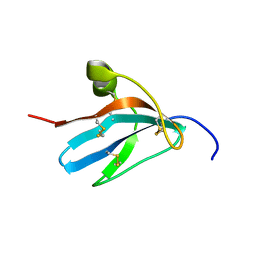 
 | | Solution structure of the leech carboxypeptidase inhibitor | | Descriptor: | Metallocarboxypeptidase inhibitor | | Authors: | Arolas, J.L, D'Silva, L, Popowicz, G.M, Aviles, F.X, Holak, T.A, Ventura, S. | | Deposit date: | 2005-04-20 | | Release date: | 2005-09-13 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | NMR structural characterization and computational predictions of the major intermediate in oxidative folding of leech carboxypeptidase inhibitor
STRUCTURE, 13, 2005
|
|
1PPC
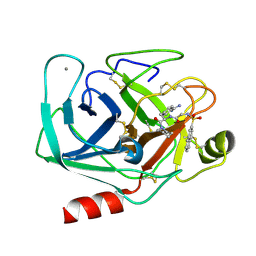 
 | | GEOMETRY OF BINDING OF THE BENZAMIDINE-AND ARGININE-BASED INHIBITORS N-ALPHA-(2-NAPHTHYL-SULPHONYL-GLYCYL)-DL-P-AMIDINOPHENYLALANYL-PIPERIDINE (NAPAP) AND (2R,4R)-4-METHYL-1-[N-ALPHA-(3-METHYL-1,2,3,4-TETRAHYDRO-8-QUINOLINESULPHONYL)-L-ARGINYL]-2-PIPERIDINE CARBOXYLIC ACID (MQPA) TO HUMAN ALPHA-THROMBIN: X-RAY CRYSTALLOGRAPHIC DETERMINATION OF THE NAPAP-TRYPSIN COMPLEX AND MODELING OF NAPAP-THROMBIN AND MQPA-THROMBIN | | Descriptor: | 1-[N-(naphthalen-2-ylsulfonyl)glycyl-4-carbamimidoyl-D-phenylalanyl]piperidine, CALCIUM ION, TRYPSIN | | Authors: | Bode, W, Turk, D. | | Deposit date: | 1991-10-24 | | Release date: | 1994-01-31 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Geometry of binding of the benzamidine- and arginine-based inhibitors N alpha-(2-naphthyl-sulphonyl-glycyl)-DL-p-amidinophenylalanyl-pipe ridine (NAPAP) and (2R,4R)-4-methyl-1-[N alpha-(3-methyl-1,2,3,4-tetrahydr quinolinesulphonyl)-L-arginyl]-2-piperidine carboxylic acid (MQPA) to human alpha-thrombin.X-ray crystallographic determination of the NAPAP-trypsin complex and modeling of NAPAP-thrombin and MQPA-thrombin.
Eur.J.Biochem., 193, 1990
|
|
1ZFL
 
 | | Solution structure of III-A, the major intermediate in the oxidative folding of leech carboxypeptidase inhibitor | | Descriptor: | Metallocarboxypeptidase inhibitor | | Authors: | Arolas, J.L, D'Silva, L, Popowicz, G.M, Aviles, F.X, Holak, T.A, Ventura, S. | | Deposit date: | 2005-04-20 | | Release date: | 2005-09-13 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | NMR structural characterization and computational predictions of the major intermediate in oxidative folding of leech carboxypeptidase inhibitor
STRUCTURE, 13, 2005
|
|
3CTI
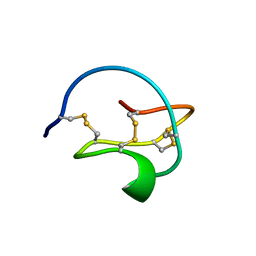 
 | |
1IAB
 
 | | CRYSTAL STRUCTURES, SPECTROSCOPIC FEATURES, AND CATALYTIC PROPERTIES OF COBALT(II), COPPER(II), NICKEL(II), AND MERCURY(II) DERIVATIVES OF THE ZINC ENDOPEPTIDASE ASTACIN. A CORRELATION OF STRUCTURE AND PROTEOLYTIC ACTIVITY | | Descriptor: | ASTACIN, COBALT (II) ION | | Authors: | Gomis-Rueth, F.-X, Stoecker, W, Bode, W. | | Deposit date: | 1994-05-09 | | Release date: | 1994-08-31 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Crystal structures, spectroscopic features, and catalytic properties of cobalt(II), copper(II), nickel(II), and mercury(II) derivatives of the zinc endopeptidase astacin. A correlation of structure and proteolytic activity.
J.Biol.Chem., 269, 1994
|
|
1IAA
 
 | | CRYSTAL STRUCTURES, SPECTROSCOPIC FEATURES, AND CATALYTIC PROPERTIES OF COBALT(II), COPPER(II), NICKEL(II), AND MERCURY(II) DERIVATIVES OF THE ZINC ENDOPEPTIDASE ASTACIN. A CORRELATION OF STRUCTURE AND PROTEOLYTIC ACTIVITY | | Descriptor: | ASTACIN, COPPER (II) ION | | Authors: | Gomis-Rueth, F.-X, Stoecker, W, Bode, W. | | Deposit date: | 1994-05-09 | | Release date: | 1994-08-31 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structures, spectroscopic features, and catalytic properties of cobalt(II), copper(II), nickel(II), and mercury(II) derivatives of the zinc endopeptidase astacin. A correlation of structure and proteolytic activity.
J.Biol.Chem., 269, 1994
|
|
1IAE
 
 | | CRYSTAL STRUCTURES, SPECTROSCOPIC FEATURES, AND CATALYTIC PROPERTIES OF COBALT(II), COPPER(II), NICKEL(II), AND MERCURY(II) DERIVATIVES OF THE ZINC ENDOPEPTIDASE ASTACIN. A CORRELATION OF STRUCTURE AND PROTEOLYTIC ACTIVITY | | Descriptor: | ASTACIN, NICKEL (II) ION | | Authors: | Grams, F, Stoecker, W, Bode, W. | | Deposit date: | 1994-05-09 | | Release date: | 1994-08-31 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Crystal structures, spectroscopic features, and catalytic properties of cobalt(II), copper(II), nickel(II), and mercury(II) derivatives of the zinc endopeptidase astacin. A correlation of structure and proteolytic activity.
J.Biol.Chem., 269, 1994
|
|
1BQY
 
 | | Plasminogen activator (TSV-PA) from snake venom | | Descriptor: | L-alpha-glutamyl-N-{(1S)-4-{[amino(iminio)methyl]amino}-1-[(1S)-2-chloro-1-hydroxyethyl]butyl}glycinamide, PLASMINOGEN ACTIVATOR | | Authors: | Parry, M.A.A, Bode, W. | | Deposit date: | 1998-08-20 | | Release date: | 1999-08-20 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The crystal structure of the novel snake venom plasminogen activator TSV-PA: a prototype structure for snake venom serine proteinases.
Structure, 6, 1998
|
|
1BQB
 | | AUREOLYSIN, STAPHYLOCOCCUS AUREUS METALLOPROTEINASE | | Descriptor: | CALCIUM ION, PROTEIN (AUREOLYSIN), ZINC ION | | Authors: | Medrano, F.J, Banbula, A, Potempa, J, Travis, J, Bode, W. | | Deposit date: | 1998-07-14 | | Release date: | 1999-01-13 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Amino-acid sequence and three-dimensional structure of the Staphylococcus aureus metalloproteinase at 1.72 A resolution.
Structure, 6, 1998
|
|
