3G5G
 
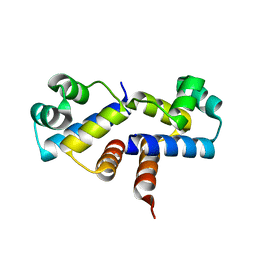 | | Crystal Structure of the Wild-Type Restriction-Modification Controller Protein C.Esp1396I | | Descriptor: | Regulatory protein | | Authors: | Ball, N.J, McGeehan, J.E, Thresh, S.J, Streeter, S.D, Kneale, G.G. | | Deposit date: | 2009-02-05 | | Release date: | 2009-08-25 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of the restriction-modification controller protein C.Esp1396I.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
8E0Y
 
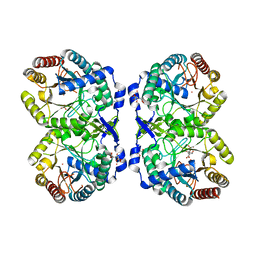 | | DAHP (3-deoxy-D-arabinoheptulosonate-7-phosphate) Synthase complexed with DAHP oxime, Pr(III), and Pi in unbound:(bound)2:other Conformations | | Descriptor: | ACETATE ION, CITRATE ANION, DAHP Oxime, ... | | Authors: | Berti, P.J, Junop, M.S, Grainger, R. | | Deposit date: | 2022-08-09 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Role of Half-of-Sites Reactivity and Inter-Subunit Communications in DAHP Synthase Catalysis and Regulation.
Biochemistry, 61, 2022
|
|
8E0Z
 
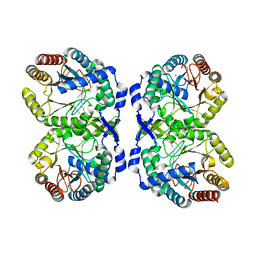 | |
8E0V
 
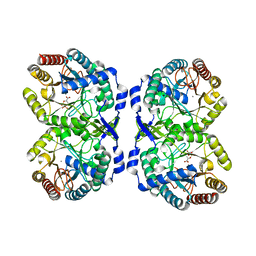 | | DAHP (3-deoxy-D-arabinoheptulosonate-7-phosphate) Synthase complexed with Mn(II), PEP, and Pi in unbound:(bound)2:other Conformations | | Descriptor: | MANGANESE (II) ION, PHOSPHATE ION, PHOSPHOENOLPYRUVATE, ... | | Authors: | Berti, P.J, Junop, M.S, Grainger, R. | | Deposit date: | 2022-08-09 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Role of Half-of-Sites Reactivity and Inter-Subunit Communications in DAHP Synthase Catalysis and Regulation.
Biochemistry, 61, 2022
|
|
8E0U
 
 | |
8E0X
 
 | | DAHP (3-deoxy-D-arabinoheptulosonate-7-phosphate) Synthase complexed with Mn(II), PEP, and Pi in unbound:(bound)2:other Conformations | | Descriptor: | MANGANESE (II) ION, PHOSPHATE ION, PHOSPHOENOLPYRUVATE, ... | | Authors: | Berti, P.J, Junop, M.S, Grainger, R. | | Deposit date: | 2022-08-09 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Role of Half-of-Sites Reactivity and Inter-Subunit Communications in DAHP Synthase Catalysis and Regulation.
Biochemistry, 61, 2022
|
|
8E0T
 
 | |
8E0S
 
 | | DAHP (3-deoxy-D-arabinoheptulosonate-7-phosphate) Synthase complexed with DAHP Oxime in unbound:(bound)2:unbound conformations | | Descriptor: | DAHP Oxime, Phospho-2-dehydro-3-deoxyheptonate aldolase, Phe-sensitive, ... | | Authors: | Berti, P.J, Junop, M.S, Grainger, R. | | Deposit date: | 2022-08-09 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Role of Half-of-Sites Reactivity and Inter-Subunit Communications in DAHP Synthase Catalysis and Regulation.
Biochemistry, 61, 2022
|
|
7Y6A
 
 | | Crystal structure of Chicken Egg Lysozyme | | Descriptor: | Lysozyme C, NITRATE ION | | Authors: | DeMirci, H. | | Deposit date: | 2022-06-18 | | Release date: | 2023-04-19 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Cryogenic X-ray crystallographic studies of biomacromolecules at Turkish Light Source " Turkish DeLight ".
Turk J Biol, 47, 2023
|
|
4F8D
 
 | |
4FN3
 
 | |
4FBI
 
 | |
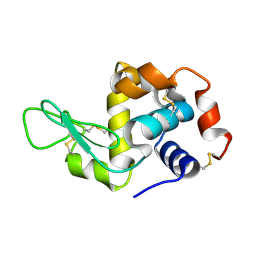
8H3W
 
 | |
5M1H
 
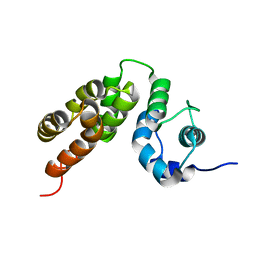 | |
5M1G
 
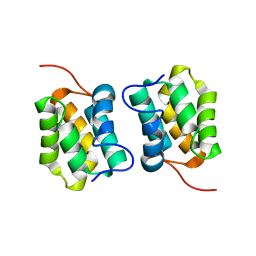 | |
5ILW
 
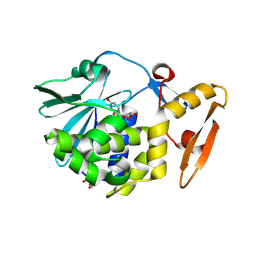 | | Crystal structure of the complex of type 1 Ribosome inactivating protein from Momordica balsamina with Uridine at 1.97 Angstrom resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, Ribosome inactivating protein, ... | | Authors: | Singh, P.K, Pandey, S, Kaur, P, Sharma, S, Singh, T.P. | | Deposit date: | 2016-03-05 | | Release date: | 2016-03-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Binding and structural studies of the complexes of type 1 ribosome inactivating protein from Momordica balsamina with uracil and uridine.
Proteins, 87, 2019
|
|
6VKV
 
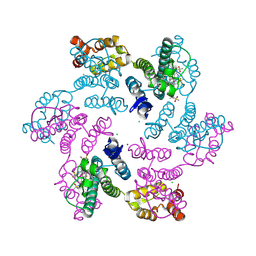 | |
7SNQ
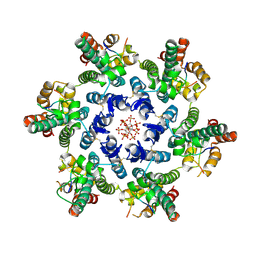 
 | |
5H7X
 
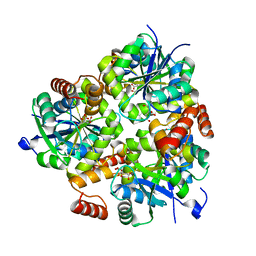 | | Crystal structure of the complex of Phosphopantetheine adenylyltransferase from Acinetobacter baumannii with 2-hydroxy-1,2,3-propane tricarboxylate at 1.76 A resolution | | Descriptor: | CITRIC ACID, Phosphopantetheine adenylyltransferase | | Authors: | Singh, P.K, Gupta, A, Kaur, P, Sharma, S, Singh, T.P. | | Deposit date: | 2016-11-21 | | Release date: | 2016-12-07 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Structural and binding studies of phosphopantetheine adenylyl transferase from Acinetobacter baumannii.
Biochim Biophys Acta Proteins Proteom, 1867, 2019
|
|
7NTU
 
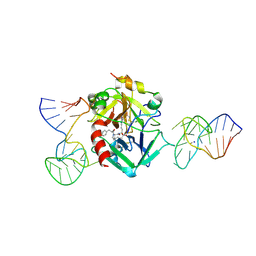 | | X-ray structure of the complex between human alpha thrombin and two duplex/quadruplex aptamers: NU172 and HD22_27mer | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, D-phenylalanyl-N-[(2S,3S)-6-{[amino(iminio)methyl]amino}-1-chloro-2-hydroxyhexan-3-yl]-L-prolinamide, HD22_27mer, ... | | Authors: | Troisi, R, Santamaria, A, Sica, F. | | Deposit date: | 2021-03-10 | | Release date: | 2021-04-28 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural and functional analysis of the simultaneous binding of two duplex/quadruplex aptamers to human alpha-thrombin.
Int.J.Biol.Macromol., 181, 2021
|
|
6PPW
 
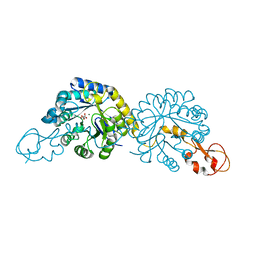 | | Crystal structure of NeuB, an N-acetylneuraminate synthase from Neisseria meningitidis, in complex with magnesium and malate | | Descriptor: | D-MALATE, MAGNESIUM ION, N-acetylneuraminate synthase | | Authors: | Rosanally, A.Z, Junop, M.S, Berti, P.J. | | Deposit date: | 2019-07-08 | | Release date: | 2019-10-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | NeuNAc Oxime: A Slow-Binding and Effectively Irreversible Inhibitor of the Sialic Acid Synthase NeuB.
Biochemistry, 58, 2019
|
|
6PPX
 
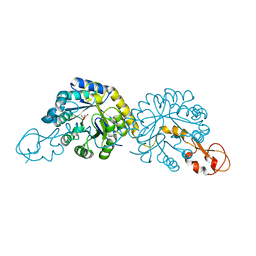 | |
6PPZ
 
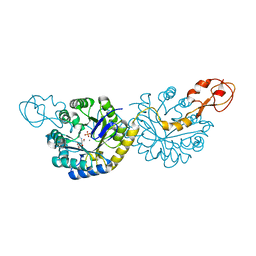 | | Crystal structure of NeuB, an N-acetylneuraminate synthase from Neisseria meningitidis, in complex with manganese, inorganic phosphate, and N-acetylmannosamine (NeuB.Mn2+.Pi.ManNAc) | | Descriptor: | 2-(ACETYLAMINO)-2-DEOXY-D-MANNOSE, MANGANESE (II) ION, N-acetylneuraminate synthase, ... | | Authors: | Berti, P.J, Junop, M.S. | | Deposit date: | 2019-07-08 | | Release date: | 2019-10-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | NeuNAc Oxime: A Slow-Binding and Effectively Irreversible Inhibitor of the Sialic Acid Synthase NeuB.
Biochemistry, 58, 2019
|
|
6PPY
 
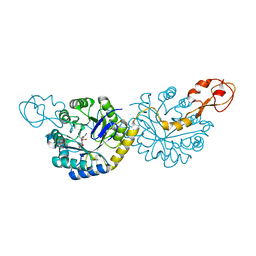 | | Crystal structure of NeuNAc oxime complexed with NeuB, an N-acetylneuraminate synthase from Neisseria meningitidis | | Descriptor: | (2E,4S,5R,6R,7S,8R)-5-(acetylamino)-4,6,7,8,9-pentahydroxy-2-(hydroxyimino)nonanoic acid (non-preferred name), N-acetylneuraminate synthase | | Authors: | Rosanally, A.Z, Junop, M.J, Berti, P.J. | | Deposit date: | 2019-07-08 | | Release date: | 2019-10-02 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | NeuNAc Oxime: A Slow-Binding and Effectively Irreversible Inhibitor of the Sialic Acid Synthase NeuB.
Biochemistry, 58, 2019
|
|
8DS5
 
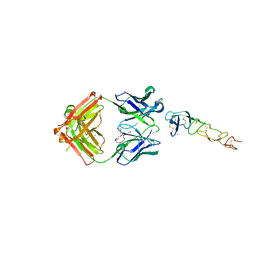 | | X-ray structure of the MK5890 Fab - CD27 antibody-antigen complex | | Descriptor: | CADMIUM ION, CD27 antigen, MK-5890 Fab heavy chain, ... | | Authors: | Fischmann, T.O. | | Deposit date: | 2022-07-21 | | Release date: | 2022-09-28 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.926 Å) | | Cite: | Preclinical characterization and clinical translation of pharmacodynamic markers for MK-5890: a human CD27 activating antibody for cancer immunotherapy.
J Immunother Cancer, 10, 2022
|
|
